Recent from talks
Knowledge base stats:
Talk channels stats:
Members stats:
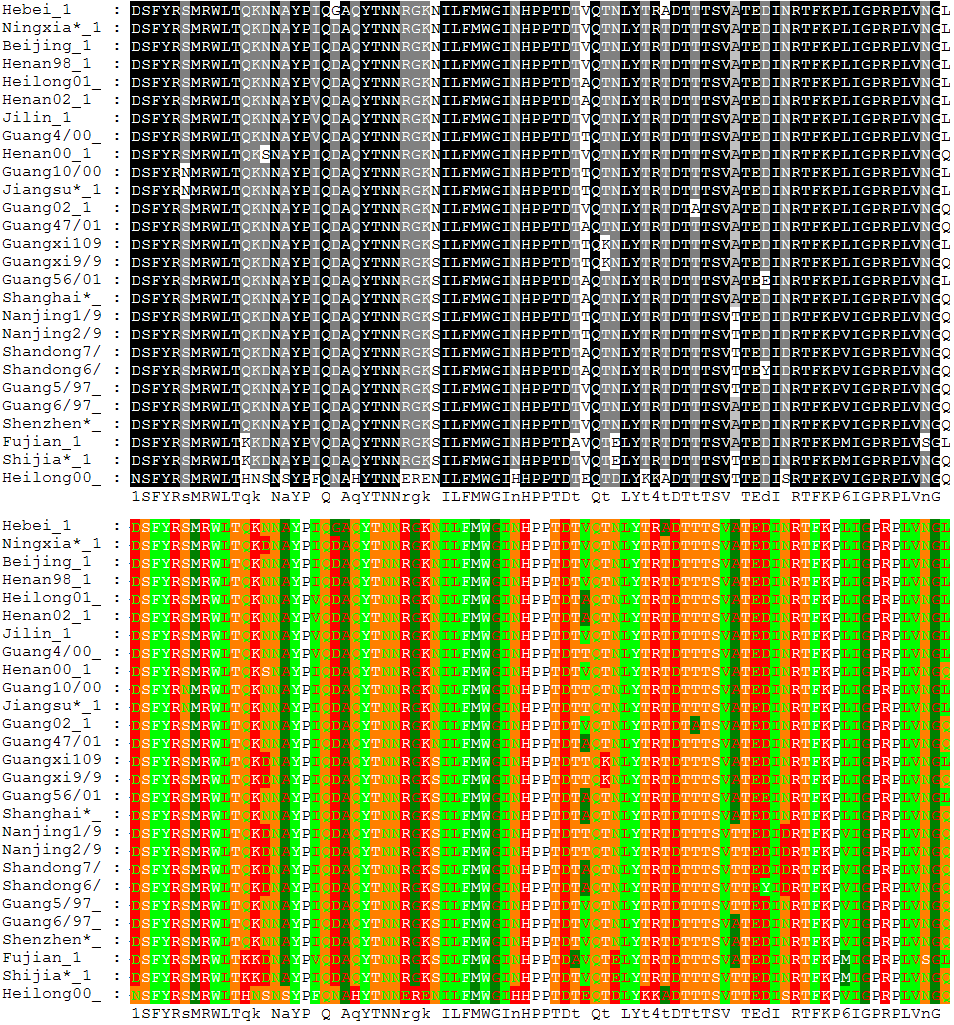
Clustal
Clustal is a computer program used for multiple sequence alignment in bioinformatics. It is one of the most widely cited bioinformatics software with two of its academic publications amongst the top 100 papers cited of all time, according to Nature in 2014.
Since its first publication in 1988, the software and its algorithms have gone through several iterations, with ClustalΩ (Omega) being the latest version as of 2011[update]. It is available as standalone software, via a web interface, and through a server hosted by the European Bioinformatics Institute.
The guide tree in the initial versions of Clustal was constructed via a UPGMA cluster analysis of the pairwise alignments, hence the name CLUSTAL.cf. The first four versions of Clustal were numbered using Arabic numerals (1 to 4), whereas the fifth version uses the Roman numeral V.cf. The next two versions proceed alphabetically using the Latin alphabet, with W standing for weighted and X for X Window to represent the changes introduced.cf. The name Omega was chosen to mark a change from the previous iterations.
Clustal aligns sequences using a heuristic that progressively builds a multiple sequence alignment from a set of pairwise alignments. This method works by analyzing the sequences as a whole and using the UPGMA/neighbor-joining method to generate a distance matrix. A guide tree is calculated from the scores of the sequences in the matrix, then subsequently used to build the multiple sequence alignment by progressively aligning the sequences in order of similarity.
Clustal creates multiple sequence alignments through three main steps:
These steps are carried out automatically by the function "Do Complete Alignment". Other options are "Do Alignment from guide tree and phylogeny" and "Produce guide tree only".
This program accepts a wide range of input formats, including NBRF/PIR, FASTA, EMBL/Swiss-Prot, Clustal, GCC/MSF, GCG9 RSF, and GDE.
The output format can be one or many of the following: Clustal, NBRF/PIR, GCG/MSF, PHYLIP, GDE, or NEXUS.
Hub AI
Clustal AI simulator
(@Clustal_simulator)
Clustal
Clustal is a computer program used for multiple sequence alignment in bioinformatics. It is one of the most widely cited bioinformatics software with two of its academic publications amongst the top 100 papers cited of all time, according to Nature in 2014.
Since its first publication in 1988, the software and its algorithms have gone through several iterations, with ClustalΩ (Omega) being the latest version as of 2011[update]. It is available as standalone software, via a web interface, and through a server hosted by the European Bioinformatics Institute.
The guide tree in the initial versions of Clustal was constructed via a UPGMA cluster analysis of the pairwise alignments, hence the name CLUSTAL.cf. The first four versions of Clustal were numbered using Arabic numerals (1 to 4), whereas the fifth version uses the Roman numeral V.cf. The next two versions proceed alphabetically using the Latin alphabet, with W standing for weighted and X for X Window to represent the changes introduced.cf. The name Omega was chosen to mark a change from the previous iterations.
Clustal aligns sequences using a heuristic that progressively builds a multiple sequence alignment from a set of pairwise alignments. This method works by analyzing the sequences as a whole and using the UPGMA/neighbor-joining method to generate a distance matrix. A guide tree is calculated from the scores of the sequences in the matrix, then subsequently used to build the multiple sequence alignment by progressively aligning the sequences in order of similarity.
Clustal creates multiple sequence alignments through three main steps:
These steps are carried out automatically by the function "Do Complete Alignment". Other options are "Do Alignment from guide tree and phylogeny" and "Produce guide tree only".
This program accepts a wide range of input formats, including NBRF/PIR, FASTA, EMBL/Swiss-Prot, Clustal, GCC/MSF, GCG9 RSF, and GDE.
The output format can be one or many of the following: Clustal, NBRF/PIR, GCG/MSF, PHYLIP, GDE, or NEXUS.