Recent from talks
Knowledge base stats:
Talk channels stats:
Members stats:
Primate T-lymphotropic virus
The primate T-lymphotropic viruses (PTLVs) are a group of retroviruses that infect primates, using their lymphocytes to reproduce. The ones that infect humans are known as human T-lymphotropic virus (HTLV), and the ones that infect Old World monkeys are called simian T-lymphotropic viruses (STLVs). PTLVs are named for their ability to cause adult T-cell leukemia/lymphoma, but in the case of HTLV-1 it can also cause a demyelinating disease called tropical spastic paraparesis. On the other hand, newer PTLVs are simply placed into the group by similarity and their connection to human disease remains unclear.
HTLVs have evolved from STLVs by interspecies transmission. Within each species of PTLV, the HTLV is more similar to its cognate STLV than to the other HTLVs. There are currently three species of PTLVs recognized by the ICTV (P/H/STLV-1, -2, -3), plus two that are reported but unrecognized (HTLV-4, STLV-5). The first known, and still most medically important PTLV is HTLV-1, discovered in 1980.
HTLVs belong to the genus Deltaretrovirus. The only other recognized species in the genus is Bovine leukemia virus, an economically-important cattle pathogen. As its name suggests, this virus causes leukemia in cows.
HTLV-1 is the prototypical PTLV, from which comparisons are drawn for the newly-known types. A retrovirus, PTLV shares the common gag-pro-pol-env set of genes, yet shows great complexity in the unique 3' end. These new proteins provide a great source of new adaptive function:
HTLV-1 has three tandem imperfect 21-base repeats as the long terminal repeat, but other PTLVs only have two.
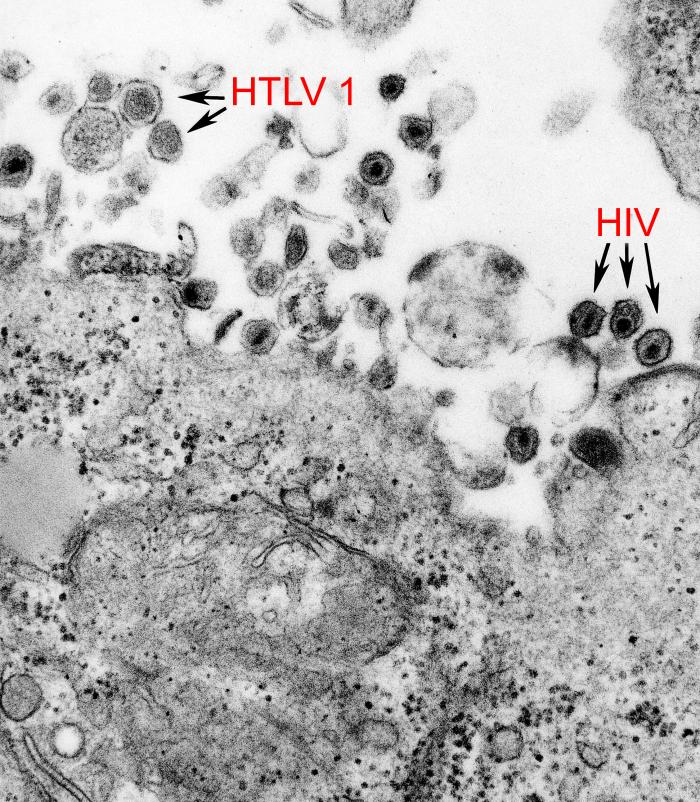
The lifecycle is common to retroviruses, starting at the envelope glycoprotein (Env) surface subunit (SU) binding to a cellular receptor (in this case GLUT1 and a host of other molecules), and ending with lysis of the cell (in this case, a lymphocyte). The virion is spherical to pleomorphic, about 80-100 nm in diameter. Unusually, the single-stranded RNA genome is present in two copies, forming a dimer specifically packed by parts of the gag protein.
The use of "HTLV-3" can cause some confusion, because the name HTLV-III was one of the names for HIV in early AIDS literature, but has since fallen out of use. The name HTLV-IV has also been used to describe HIV-2. A large Canadian study documented this confusion among healthcare workers, where >90% of HTLV tests ordered by physicians were actually intended to be HIV tests.
PTLV-1 is the medically most important species in the class. Discovered by Robert Gallo and colleagues in 1980, HTLV-1 has been implicated in several kinds of diseases, including tropical spastic paraparesis and as a virus cancer link for adult T-cell leukemia/lymphoma. Between 1 in 20 and 1 in 25 infected people are thought to develop cancer as a result of the virus. STLV-1 is oncogenic in Japanese macaques.
Hub AI
Primate T-lymphotropic virus AI simulator
(@Primate T-lymphotropic virus_simulator)
Primate T-lymphotropic virus
The primate T-lymphotropic viruses (PTLVs) are a group of retroviruses that infect primates, using their lymphocytes to reproduce. The ones that infect humans are known as human T-lymphotropic virus (HTLV), and the ones that infect Old World monkeys are called simian T-lymphotropic viruses (STLVs). PTLVs are named for their ability to cause adult T-cell leukemia/lymphoma, but in the case of HTLV-1 it can also cause a demyelinating disease called tropical spastic paraparesis. On the other hand, newer PTLVs are simply placed into the group by similarity and their connection to human disease remains unclear.
HTLVs have evolved from STLVs by interspecies transmission. Within each species of PTLV, the HTLV is more similar to its cognate STLV than to the other HTLVs. There are currently three species of PTLVs recognized by the ICTV (P/H/STLV-1, -2, -3), plus two that are reported but unrecognized (HTLV-4, STLV-5). The first known, and still most medically important PTLV is HTLV-1, discovered in 1980.
HTLVs belong to the genus Deltaretrovirus. The only other recognized species in the genus is Bovine leukemia virus, an economically-important cattle pathogen. As its name suggests, this virus causes leukemia in cows.
HTLV-1 is the prototypical PTLV, from which comparisons are drawn for the newly-known types. A retrovirus, PTLV shares the common gag-pro-pol-env set of genes, yet shows great complexity in the unique 3' end. These new proteins provide a great source of new adaptive function:
HTLV-1 has three tandem imperfect 21-base repeats as the long terminal repeat, but other PTLVs only have two.
The lifecycle is common to retroviruses, starting at the envelope glycoprotein (Env) surface subunit (SU) binding to a cellular receptor (in this case GLUT1 and a host of other molecules), and ending with lysis of the cell (in this case, a lymphocyte). The virion is spherical to pleomorphic, about 80-100 nm in diameter. Unusually, the single-stranded RNA genome is present in two copies, forming a dimer specifically packed by parts of the gag protein.
The use of "HTLV-3" can cause some confusion, because the name HTLV-III was one of the names for HIV in early AIDS literature, but has since fallen out of use. The name HTLV-IV has also been used to describe HIV-2. A large Canadian study documented this confusion among healthcare workers, where >90% of HTLV tests ordered by physicians were actually intended to be HIV tests.
PTLV-1 is the medically most important species in the class. Discovered by Robert Gallo and colleagues in 1980, HTLV-1 has been implicated in several kinds of diseases, including tropical spastic paraparesis and as a virus cancer link for adult T-cell leukemia/lymphoma. Between 1 in 20 and 1 in 25 infected people are thought to develop cancer as a result of the virus. STLV-1 is oncogenic in Japanese macaques.