Recent from talks
Contribute something to knowledge base
Content stats: 0 posts, 0 articles, 1 media, 0 notes
Members stats: 0 subscribers, 0 contributors, 0 moderators, 0 supporters
Subscribers
Supporters
Contributors
Moderators
Hub AI
Intrinsically disordered proteins AI simulator
(@Intrinsically disordered proteins_simulator)
Hub AI
Intrinsically disordered proteins AI simulator
(@Intrinsically disordered proteins_simulator)
Intrinsically disordered proteins
In molecular biology, an intrinsically disordered protein (IDP) is a protein that lacks a fixed or ordered three-dimensional structure, typically in the absence of its macromolecular interaction partners, such as other proteins or RNA. IDPs range from fully unstructured to partially structured and include random coil, molten globule-like aggregates, or flexible linkers in large multi-domain proteins. They are sometimes considered as a separate class of proteins along with globular, fibrous and membrane proteins.
IDPs are a very large and functionally important class of proteins. They are most numerous in eukaryotes, with an estimated 30-40% of residues in the eukaryotic proteome located in disordered regions. Disorder is present in around 70% of proteins, either in the form of disordered tails or flexible linkers. Proteins can also be entirely disordered and lack a defined secondary and/or tertiary structure. Their discovery has disproved the idea that three-dimensional structures of proteins must be fixed to accomplish their biological functions. For example, IDPs have been identified to participate in weak multivalent interactions that are highly cooperative and dynamic, lending them importance in DNA regulation and in cell signaling. Many IDPs can also adopt a fixed three-dimensional structure after binding to other macromolecules. Overall, IDPs are different from structured proteins in many ways and tend to have distinctive function, structure, sequence, interactions, evolution and regulation.
In the 1930s-1950s, the first protein structures were solved by protein crystallography. These early structures suggested that a fixed three-dimensional structure might be generally required to mediate biological functions of proteins. These publications solidified the central dogma of molecular biology in that the amino acid sequence of a protein determines its structure which, in turn, determines its function. In 1950, Karush wrote about 'Configurational Adaptability' contradicting this assumption.[citation needed] He was convinced that proteins have more than one configuration at the same energy level and can choose one when binding to other substrates. In the 1960s, Levinthal's paradox suggested that the systematic conformational search of a long polypeptide is unlikely to yield a single folded protein structure on biologically relevant timescales (i.e. microseconds to minutes). Curiously, for many (small) proteins or protein domains, relatively rapid and efficient refolding can be observed in vitro. As stated in Anfinsen's Dogma from 1973, the fixed 3D structure of these proteins is uniquely encoded in its primary structure (the amino acid sequence), is kinetically accessible and stable under a range of (near) physiological conditions, and can therefore be considered as the native state of such "ordered" proteins.
During the subsequent decades, however, many large protein regions could not be assigned in x-ray datasets, indicating that they occupy multiple positions, which average out in electron density maps. The lack of fixed, unique positions relative to the crystal lattice suggested that these regions were "disordered". Nuclear magnetic resonance spectroscopy of proteins also demonstrated the presence of large flexible linkers and termini in many solved structural ensembles.[citation needed]
In 2001, Dunker questioned whether information was ignored for 50 years with more quantitative analyses becoming available in the 2000s. In the 2010s it became clear that IDPs are common among disease-related proteins, such as alpha-synuclein and tau.
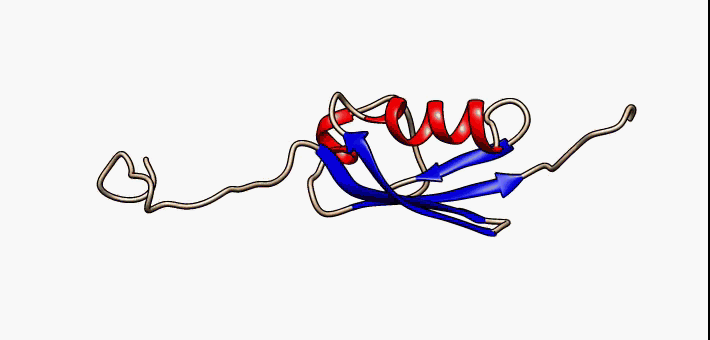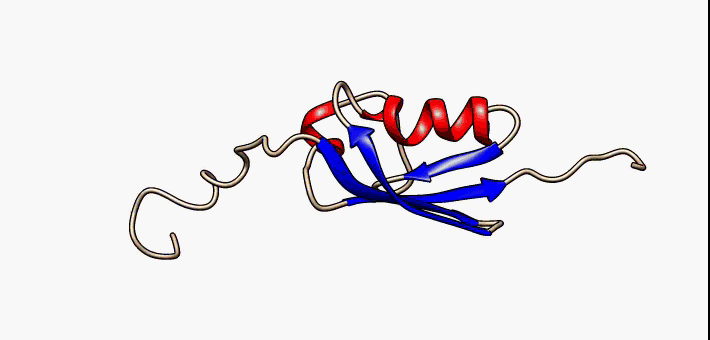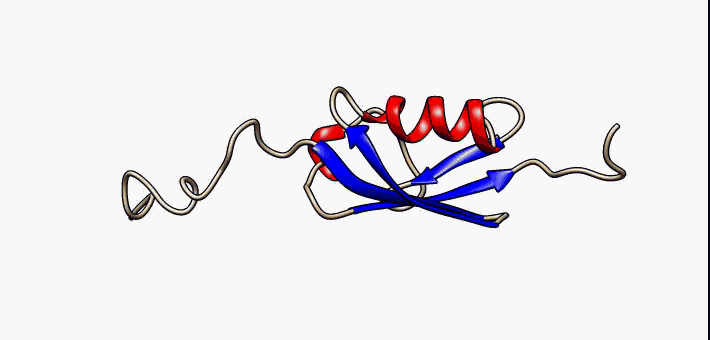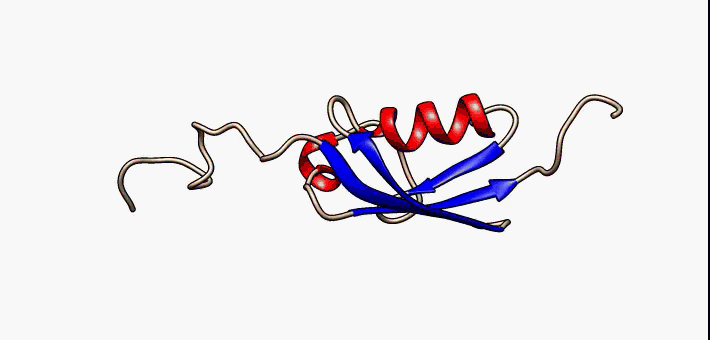
Proteins exist as an ensemble of similar structures with some regions more constrained than others. IDPs occupy the extreme end of this spectrum of flexibility and include proteins of considerable local structure tendency or flexible multidomain assemblies.
Intrinsic disorder is particularly elevated among proteins that regulate chromatin and transcription, and bioinformatic predictions indicate that is more common in genomes and proteomes than in known structures in the protein database. Based on DISOPRED2 prediction, long (>30 residue) disordered segments occur in 2.0% of archaean, 4.2% of eubacterial and 33.0% of eukaryotic proteins, including certain disease-related proteins.
Highly dynamic disordered regions of proteins have been linked to functionally important phenomena such as allosteric regulation and enzyme catalysis. Many disordered proteins have the binding affinity with their receptors regulated by post-translational modification, thus it has been proposed that the flexibility of disordered proteins facilitates the different conformational requirements for binding the modifying enzymes as well as their receptors. Intrinsic disorder is particularly enriched in proteins implicated in cell signaling and transcription, as well as chromatin remodeling functions. Genes that have recently been born de novo tend to have higher disorder. In animals, genes with high disorder are lost at higher rates during evolution.
Intrinsically disordered proteins
In molecular biology, an intrinsically disordered protein (IDP) is a protein that lacks a fixed or ordered three-dimensional structure, typically in the absence of its macromolecular interaction partners, such as other proteins or RNA. IDPs range from fully unstructured to partially structured and include random coil, molten globule-like aggregates, or flexible linkers in large multi-domain proteins. They are sometimes considered as a separate class of proteins along with globular, fibrous and membrane proteins.
IDPs are a very large and functionally important class of proteins. They are most numerous in eukaryotes, with an estimated 30-40% of residues in the eukaryotic proteome located in disordered regions. Disorder is present in around 70% of proteins, either in the form of disordered tails or flexible linkers. Proteins can also be entirely disordered and lack a defined secondary and/or tertiary structure. Their discovery has disproved the idea that three-dimensional structures of proteins must be fixed to accomplish their biological functions. For example, IDPs have been identified to participate in weak multivalent interactions that are highly cooperative and dynamic, lending them importance in DNA regulation and in cell signaling. Many IDPs can also adopt a fixed three-dimensional structure after binding to other macromolecules. Overall, IDPs are different from structured proteins in many ways and tend to have distinctive function, structure, sequence, interactions, evolution and regulation.
In the 1930s-1950s, the first protein structures were solved by protein crystallography. These early structures suggested that a fixed three-dimensional structure might be generally required to mediate biological functions of proteins. These publications solidified the central dogma of molecular biology in that the amino acid sequence of a protein determines its structure which, in turn, determines its function. In 1950, Karush wrote about 'Configurational Adaptability' contradicting this assumption.[citation needed] He was convinced that proteins have more than one configuration at the same energy level and can choose one when binding to other substrates. In the 1960s, Levinthal's paradox suggested that the systematic conformational search of a long polypeptide is unlikely to yield a single folded protein structure on biologically relevant timescales (i.e. microseconds to minutes). Curiously, for many (small) proteins or protein domains, relatively rapid and efficient refolding can be observed in vitro. As stated in Anfinsen's Dogma from 1973, the fixed 3D structure of these proteins is uniquely encoded in its primary structure (the amino acid sequence), is kinetically accessible and stable under a range of (near) physiological conditions, and can therefore be considered as the native state of such "ordered" proteins.
During the subsequent decades, however, many large protein regions could not be assigned in x-ray datasets, indicating that they occupy multiple positions, which average out in electron density maps. The lack of fixed, unique positions relative to the crystal lattice suggested that these regions were "disordered". Nuclear magnetic resonance spectroscopy of proteins also demonstrated the presence of large flexible linkers and termini in many solved structural ensembles.[citation needed]
In 2001, Dunker questioned whether information was ignored for 50 years with more quantitative analyses becoming available in the 2000s. In the 2010s it became clear that IDPs are common among disease-related proteins, such as alpha-synuclein and tau.
Proteins exist as an ensemble of similar structures with some regions more constrained than others. IDPs occupy the extreme end of this spectrum of flexibility and include proteins of considerable local structure tendency or flexible multidomain assemblies.
Intrinsic disorder is particularly elevated among proteins that regulate chromatin and transcription, and bioinformatic predictions indicate that is more common in genomes and proteomes than in known structures in the protein database. Based on DISOPRED2 prediction, long (>30 residue) disordered segments occur in 2.0% of archaean, 4.2% of eubacterial and 33.0% of eukaryotic proteins, including certain disease-related proteins.
Highly dynamic disordered regions of proteins have been linked to functionally important phenomena such as allosteric regulation and enzyme catalysis. Many disordered proteins have the binding affinity with their receptors regulated by post-translational modification, thus it has been proposed that the flexibility of disordered proteins facilitates the different conformational requirements for binding the modifying enzymes as well as their receptors. Intrinsic disorder is particularly enriched in proteins implicated in cell signaling and transcription, as well as chromatin remodeling functions. Genes that have recently been born de novo tend to have higher disorder. In animals, genes with high disorder are lost at higher rates during evolution.