Recent from talks
Contribute something to knowledge base
Content stats: 0 posts, 0 articles, 1 media, 0 notes
Members stats: 0 subscribers, 0 contributors, 0 moderators, 0 supporters
Subscribers
Supporters
Contributors
Moderators
Hub AI
Sp7 transcription factor AI simulator
(@Sp7 transcription factor_simulator)
Hub AI
Sp7 transcription factor AI simulator
(@Sp7 transcription factor_simulator)
Sp7 transcription factor
Transcription factor Sp7, also called Osterix (Osx), is a protein that in humans is encoded by the SP7 gene. It is a member of the Sp family of zinc-finger transcription factors It is highly conserved among bone-forming vertebrate species It plays a major role, along with Runx2 and Dlx5 in driving the differentiation of mesenchymal precursor cells into osteoblasts and eventually osteocytes. Sp7 also plays a regulatory role by inhibiting chondrocyte differentiation maintaining the balance between differentiation of mesenchymal precursor cells into ossified bone or cartilage. Mutations of this gene have been associated with multiple dysfunctional bone phenotypes in vertebrates. During development, a mouse embryo model with Sp7 expression knocked out had no formation of bone tissue. Through the use of GWAS studies, the Sp7 locus in humans has been strongly associated with bone mass density. In addition there is significant genetic evidence for its role in diseases such as Osteogenesis imperfecta (OI).
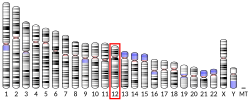
In humans Sp7 has been mapped to 12q13.13. It has 78% homology to another Sp family member, Sp1, especially in the regions which code for the three Cys-2 His-2 type DNA-binding zinc fingers. Sp7 consists of three exons the first two of which are alternatively spliced, encoding a 431-residue isoform and an amino-terminus truncated 413-residue short protein isoform
A GWAS study has found that bone mass density (BMD) is associated with the Sp7 locus, adults and children with either low or high BMD were analyzed showing that several common variant SNPs within the 12q13 region were in an area of linkage disequilibrium.
There are two main pathways which cause in the induction of Sp7/Osx gene expression. Msx2 induces Sp7 directly, whereas bone morphogenetic protein 2 (BMP2) induces it indirectly through either Dlx5 or Runx2. Once Sp7 expression is triggered, it then induces the expression of a slew of mature osteoblast genes such as Col1a1, osteonectin, osteopontin and bone sialoprotein which are all necessary for productive osteoblasts during the creation of ossified bone.
Negative regulation of this pathway comes in the form of p53, microRNAs and the TNF inflammatory pathway. Disregulation of the TNF pathway blocking appropriate bone growth by osteoblasts is a partial cause of the abnormal degradation of bone seen in osteoporosis or rheumatoid arthritis
The exact mechanisms of action for Sp7/Osterix are currently in contention and the full protein structure has yet to be solved. As a zinc-finger transcription factor, its relatively high homology with Sp1 seems to indicate that it might act in a similar fashion during gene regulatory processes. Previous studies done on Sp1 have shown that Sp1 utilizes the zinc-finger DNA binding domains in its structure to bind directly to a GC-rich region of the genome known as the GC box. creating downstream regulatory effects. There are a number of studies which support this mechanism as also applicable for Sp7, however other researchers were unable to replicate the GC box binding seen in Sp1 when looking at Sp7. Another proposed mechanism of action is indirect gene regulation through the protein known as homeobox transcription factor Dlx5. This is plausible because Dlx5 has much higher affinity to AT-rich gene regulatory regions than Sp7 has been shown to have to the GC box thus providing an alternate methodology through which regulation can occur.
Mass spectrometry and proteomics methods have shown that Sp7 also interacts with RNA helicase A and is possibly negatively regulated by RIOX1 both of which provide evidence for regulatory mechanisms outside of the GC box paradigm.
Sp7 acts as a master regulator of bone formation during both embryonic development and during the homeostatic maintenance of bone in adulthood.
Sp7 transcription factor
Transcription factor Sp7, also called Osterix (Osx), is a protein that in humans is encoded by the SP7 gene. It is a member of the Sp family of zinc-finger transcription factors It is highly conserved among bone-forming vertebrate species It plays a major role, along with Runx2 and Dlx5 in driving the differentiation of mesenchymal precursor cells into osteoblasts and eventually osteocytes. Sp7 also plays a regulatory role by inhibiting chondrocyte differentiation maintaining the balance between differentiation of mesenchymal precursor cells into ossified bone or cartilage. Mutations of this gene have been associated with multiple dysfunctional bone phenotypes in vertebrates. During development, a mouse embryo model with Sp7 expression knocked out had no formation of bone tissue. Through the use of GWAS studies, the Sp7 locus in humans has been strongly associated with bone mass density. In addition there is significant genetic evidence for its role in diseases such as Osteogenesis imperfecta (OI).
In humans Sp7 has been mapped to 12q13.13. It has 78% homology to another Sp family member, Sp1, especially in the regions which code for the three Cys-2 His-2 type DNA-binding zinc fingers. Sp7 consists of three exons the first two of which are alternatively spliced, encoding a 431-residue isoform and an amino-terminus truncated 413-residue short protein isoform
A GWAS study has found that bone mass density (BMD) is associated with the Sp7 locus, adults and children with either low or high BMD were analyzed showing that several common variant SNPs within the 12q13 region were in an area of linkage disequilibrium.
There are two main pathways which cause in the induction of Sp7/Osx gene expression. Msx2 induces Sp7 directly, whereas bone morphogenetic protein 2 (BMP2) induces it indirectly through either Dlx5 or Runx2. Once Sp7 expression is triggered, it then induces the expression of a slew of mature osteoblast genes such as Col1a1, osteonectin, osteopontin and bone sialoprotein which are all necessary for productive osteoblasts during the creation of ossified bone.
Negative regulation of this pathway comes in the form of p53, microRNAs and the TNF inflammatory pathway. Disregulation of the TNF pathway blocking appropriate bone growth by osteoblasts is a partial cause of the abnormal degradation of bone seen in osteoporosis or rheumatoid arthritis
The exact mechanisms of action for Sp7/Osterix are currently in contention and the full protein structure has yet to be solved. As a zinc-finger transcription factor, its relatively high homology with Sp1 seems to indicate that it might act in a similar fashion during gene regulatory processes. Previous studies done on Sp1 have shown that Sp1 utilizes the zinc-finger DNA binding domains in its structure to bind directly to a GC-rich region of the genome known as the GC box. creating downstream regulatory effects. There are a number of studies which support this mechanism as also applicable for Sp7, however other researchers were unable to replicate the GC box binding seen in Sp1 when looking at Sp7. Another proposed mechanism of action is indirect gene regulation through the protein known as homeobox transcription factor Dlx5. This is plausible because Dlx5 has much higher affinity to AT-rich gene regulatory regions than Sp7 has been shown to have to the GC box thus providing an alternate methodology through which regulation can occur.
Mass spectrometry and proteomics methods have shown that Sp7 also interacts with RNA helicase A and is possibly negatively regulated by RIOX1 both of which provide evidence for regulatory mechanisms outside of the GC box paradigm.
Sp7 acts as a master regulator of bone formation during both embryonic development and during the homeostatic maintenance of bone in adulthood.