Recent from talks
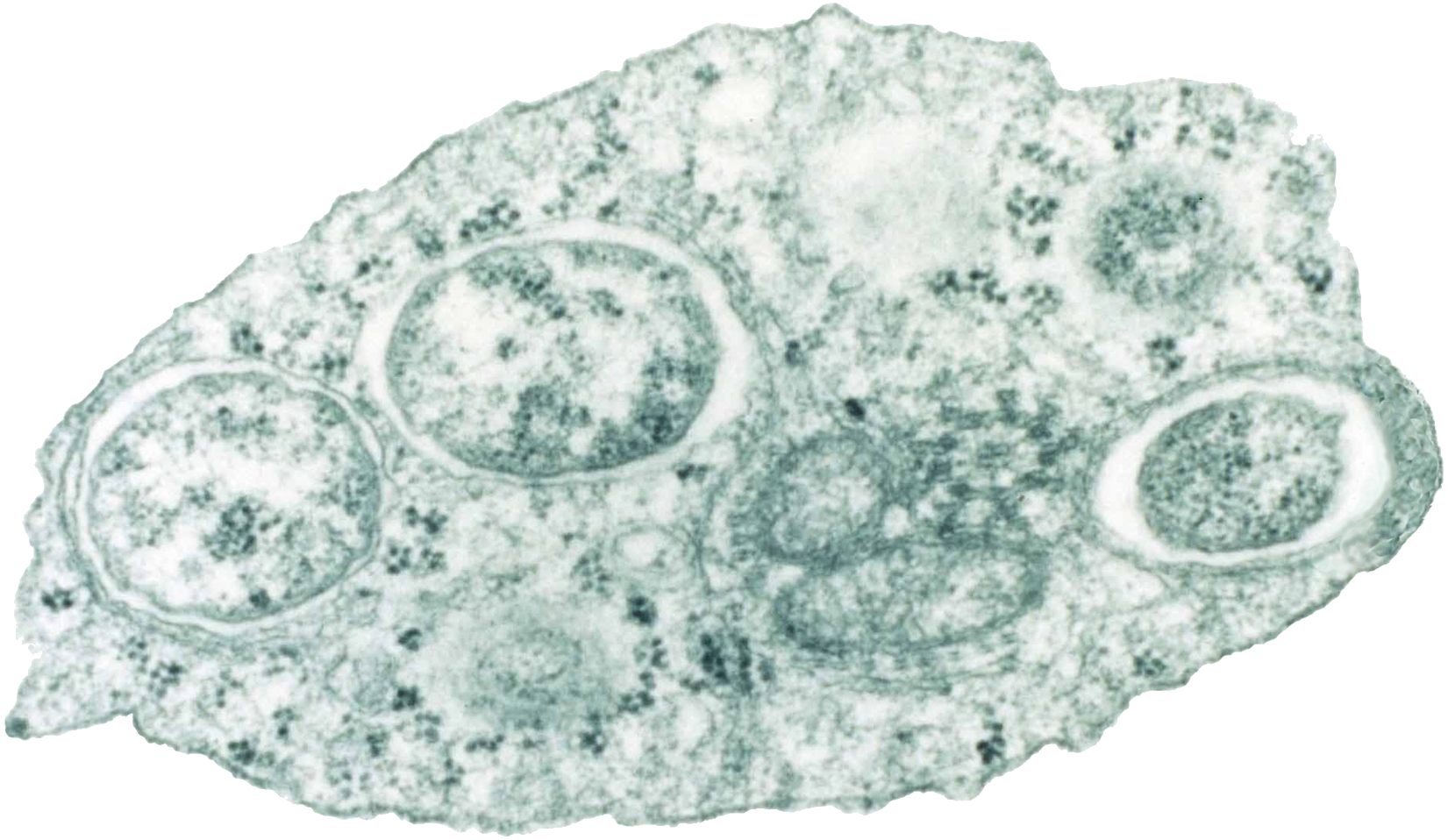
Alphaproteobacteria
Knowledge base stats:
Talk channels stats:
Members stats:
Alphaproteobacteria
Alphaproteobacteria or α-proteobacteria, also called α-Purple bacteria in earlier literature, is a class of bacteria in the phylum Pseudomonadota (also called "Proteobacteria"). The Magnetococcales and Mariprofundales are considered basal or sister to the Alphaproteobacteria. The Alphaproteobacteria are highly diverse and possess few commonalities, but nevertheless share a common ancestor. Like all proteobacteria, its members are gram-negative, although some of its intracellular parasitic members lack peptidoglycan and are consequently gram variable.
The Alphaproteobacteria are a diverse taxon and comprise several phototrophic genera, several genera metabolising C1-compounds (e.g. Methylobacterium spp.), symbionts of plants (e.g. Rhizobium spp.), endosymbionts of arthropods (Wolbachia) and intracellular pathogens (e.g. Rickettsia). Moreover, the class is sister to the protomitochondrion, the bacterium that was engulfed by the eukaryotic ancestor and gave rise to the mitochondria, which are organelles in eukaryotic cells (see Endosymbiotic theory). A species of technological interest is Agrobacterium tumefaciens (also called Rhizobium radiobacter): scientists often use this species to transfer foreign DNA into plant genomes. Aerobic anoxygenic phototrophic bacteria, such as Pelagibacter ubique, are alphaproteobacteria that are a widely distributed and may constitute over 10% of the open ocean microbial community.
Several points of disagreement muddy the recovery of the phylogenetic relationships among the Alphaproteobacteria clades from the genomic data. One such point centers on the placement of the Pelagibacterales stemming from the large differences in gene content (e.g. genome streamlining in Pelagibacter ubique) and GC-content between members of several orders. Specifically, certain species within Pelagibacterales, Rickettsiales, and Holosporales possess AT-rich genomes, containing higher-assayed concentrations of adenine-thymine (AT) pairs than guanine-cytosine (GC) base pairs. While it could be a case of convergent evolution resulting in an artefactual clustering, several studies disagree and no consensus has been reached.
Furthermore, the GC-content of ribosomal RNA, the traditional phylogenetic marker for prokaryotes, does not correlate well with the GC-content of the genome. For example, members of the Holosporales have a much higher ribosomal GC-content than members of the Pelagibacterales and Rickettsiales, though they are more closely related to species with high genomic GC-contents than to members of the latter two orders.
Alphaproteobacteria are divided into three subclasses, Magnetococcidae, Rickettsidae, and Caulobacteridae. The basal group is Magnetococcidae, composed of a large diversity of magnetotactic bacteria only one of which, Magnetococcus marinus, is formally described. The Rickettsidae is composed of the intracellular Rickettsiales and the free-living Pelagibacterales. The Caulobacteridae is composed of the Holosporales, Rhodospirillales, Sphingomonadales, Rhodobacterales, Caulobacterales, Kiloniellales, Kordiimonadales, Parvularculales, and Sneathiellales.
Comparative analyses of the sequenced genomes have revealed many conserved insertion-deletions (indels) in widely distributed proteins and whole proteins (i.e. signature proteins) that are distinctive characteristics of either all Alphaproteobacteria, or their different main orders (viz. Rhizobiales, Rhodobacterales, Rhodospirillales, Rickettsiales, Sphingomonadales and Caulobacterales) and families (viz. Rickettsiaceae, Anaplasmataceae, Rhodospirillaceae, Acetobacteraceae, Bradyrhiozobiaceae, Brucellaceae and Bartonellaceae).
These molecular signatures provide a means to circumscribe the taxonomic groups and to identify and assign new species accurately. Phylogenetic analyses and conserved indels in large numbers of other proteins provide evidence that Alphaproteobacteria have branched off later than most other phyla and classes of Bacteria except Betaproteobacteria and Gammaproteobacteria.
Other phylogenetic debates turn on the placement of Magnetococcidae and the protomitochondrion. There are some debates for the inclusion of Magnetococcidae in Alphaproteobacteria. For example, an independent proteobacterial class ("Candidatus Etaproteobacteria") for Magnetococcidae has been proposed. A recent phylogenomic study suggests the placement of the protomitochondrial clade between Magnetococcidae and all other alphaproteobacterial taxa, which suggests an early divergence of the protomitochondrial lineage from the rest of alphaproteobacteria, except for Magnetococcidae. This phylogeny also suggests that the protomitochondrial lineage does not necessarily have a close relationship to Rickettsidae.
Hub AI
Alphaproteobacteria AI simulator
(@Alphaproteobacteria_simulator)
Alphaproteobacteria
Alphaproteobacteria or α-proteobacteria, also called α-Purple bacteria in earlier literature, is a class of bacteria in the phylum Pseudomonadota (also called "Proteobacteria"). The Magnetococcales and Mariprofundales are considered basal or sister to the Alphaproteobacteria. The Alphaproteobacteria are highly diverse and possess few commonalities, but nevertheless share a common ancestor. Like all proteobacteria, its members are gram-negative, although some of its intracellular parasitic members lack peptidoglycan and are consequently gram variable.
The Alphaproteobacteria are a diverse taxon and comprise several phototrophic genera, several genera metabolising C1-compounds (e.g. Methylobacterium spp.), symbionts of plants (e.g. Rhizobium spp.), endosymbionts of arthropods (Wolbachia) and intracellular pathogens (e.g. Rickettsia). Moreover, the class is sister to the protomitochondrion, the bacterium that was engulfed by the eukaryotic ancestor and gave rise to the mitochondria, which are organelles in eukaryotic cells (see Endosymbiotic theory). A species of technological interest is Agrobacterium tumefaciens (also called Rhizobium radiobacter): scientists often use this species to transfer foreign DNA into plant genomes. Aerobic anoxygenic phototrophic bacteria, such as Pelagibacter ubique, are alphaproteobacteria that are a widely distributed and may constitute over 10% of the open ocean microbial community.
Several points of disagreement muddy the recovery of the phylogenetic relationships among the Alphaproteobacteria clades from the genomic data. One such point centers on the placement of the Pelagibacterales stemming from the large differences in gene content (e.g. genome streamlining in Pelagibacter ubique) and GC-content between members of several orders. Specifically, certain species within Pelagibacterales, Rickettsiales, and Holosporales possess AT-rich genomes, containing higher-assayed concentrations of adenine-thymine (AT) pairs than guanine-cytosine (GC) base pairs. While it could be a case of convergent evolution resulting in an artefactual clustering, several studies disagree and no consensus has been reached.
Furthermore, the GC-content of ribosomal RNA, the traditional phylogenetic marker for prokaryotes, does not correlate well with the GC-content of the genome. For example, members of the Holosporales have a much higher ribosomal GC-content than members of the Pelagibacterales and Rickettsiales, though they are more closely related to species with high genomic GC-contents than to members of the latter two orders.
Alphaproteobacteria are divided into three subclasses, Magnetococcidae, Rickettsidae, and Caulobacteridae. The basal group is Magnetococcidae, composed of a large diversity of magnetotactic bacteria only one of which, Magnetococcus marinus, is formally described. The Rickettsidae is composed of the intracellular Rickettsiales and the free-living Pelagibacterales. The Caulobacteridae is composed of the Holosporales, Rhodospirillales, Sphingomonadales, Rhodobacterales, Caulobacterales, Kiloniellales, Kordiimonadales, Parvularculales, and Sneathiellales.
Comparative analyses of the sequenced genomes have revealed many conserved insertion-deletions (indels) in widely distributed proteins and whole proteins (i.e. signature proteins) that are distinctive characteristics of either all Alphaproteobacteria, or their different main orders (viz. Rhizobiales, Rhodobacterales, Rhodospirillales, Rickettsiales, Sphingomonadales and Caulobacterales) and families (viz. Rickettsiaceae, Anaplasmataceae, Rhodospirillaceae, Acetobacteraceae, Bradyrhiozobiaceae, Brucellaceae and Bartonellaceae).
These molecular signatures provide a means to circumscribe the taxonomic groups and to identify and assign new species accurately. Phylogenetic analyses and conserved indels in large numbers of other proteins provide evidence that Alphaproteobacteria have branched off later than most other phyla and classes of Bacteria except Betaproteobacteria and Gammaproteobacteria.
Other phylogenetic debates turn on the placement of Magnetococcidae and the protomitochondrion. There are some debates for the inclusion of Magnetococcidae in Alphaproteobacteria. For example, an independent proteobacterial class ("Candidatus Etaproteobacteria") for Magnetococcidae has been proposed. A recent phylogenomic study suggests the placement of the protomitochondrial clade between Magnetococcidae and all other alphaproteobacterial taxa, which suggests an early divergence of the protomitochondrial lineage from the rest of alphaproteobacteria, except for Magnetococcidae. This phylogeny also suggests that the protomitochondrial lineage does not necessarily have a close relationship to Rickettsidae.