Recent from talks
Knowledge base stats:
Talk channels stats:
Members stats:
PA clan of proteases
The PA clan (Proteases of mixed nucleophile, superfamily A) is the largest group of proteases with common ancestry as identified by structural homology. Members have a chymotrypsin-like fold and similar proteolysis mechanisms but can have identity of <10%. The clan contains both cysteine and serine proteases (different nucleophiles). PA clan proteases can be found in plants, animals, fungi, eubacteria, archaea and viruses.
The common use of the catalytic triad for hydrolysis by multiple clans of proteases, including the PA clan, represents an example of convergent evolution. The differences in the catalytic triad within the PA clan is also an example of divergent evolution of active sites in enzymes.
In the 1960s, the sequence similarity of several proteases indicated that they were evolutionarily related. These were grouped into the chymotrypsin-like serine proteases (now called the S1 family). As the structures of these, and other proteases were solved by X-ray crystallography in the 1970s and 80s, it was noticed that several viral proteases such as Tobacco Etch Virus protease showed structural homology despite no discernible sequence similarity and even a different nucleophile. Based on structural homology, a superfamily was defined and later named the PA clan (by the MEROPS classification system). As more structures are solved, more protease families have been added to the PA clan superfamily.
The P refers to Proteases of mixed nucleophile. The A indicates that it was the first such clan to be identified (there also exist the PB, PC, PD and PE clans).
Despite retaining as little as 10% sequence identity, PA clan members isolated from viruses, prokaryotes and eukaryotes show structural homology and can be aligned by structural similarity (e.g. with DALI).
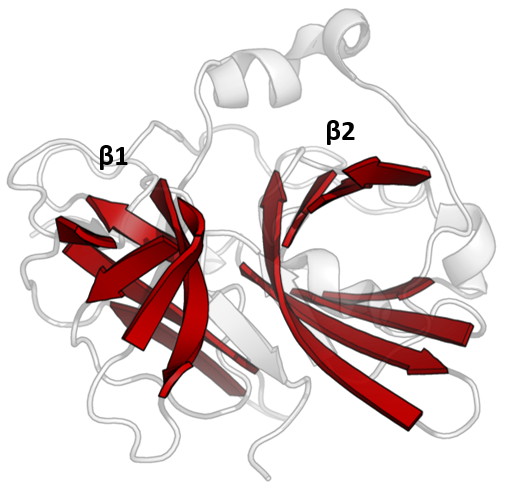
PA clan proteases all share a core motif of two β-barrels with covalent catalysis performed by an acid-histidine-nucleophile catalytic triad motif. The barrels are arranged perpendicularly beside each other with hydrophobic residues holding them together as the core scaffold for the enzyme. The triad residues are split between the two barrels so that catalysis takes place at their interface.
In addition to the double β-barrel core, some viral proteases (such as TEV protease) have a long, flexible C-terminal loop that forms a lid that completely covers the substrate and create a binding tunnel. This tunnel contains a set of tight binding pockets such that each side chain of the substrate peptide (P6 to P1’) is bound in a complementary site (S6 to S1’) and specificity is endowed by the large contact area between enzyme and substrate. Conversely, cellular proteases that lack this loop, such as trypsin have broader specificity.
Structural homology indicates that the PA clan members are descended from a common ancestor of the same fold. Although PA clan proteases use a catalytic triad perform 2-step nucleophilic catalysis, some families use serine as the nucleophile whereas others use cysteine. The superfamily is therefore an extreme example of divergent enzyme evolution since during evolutionary history, the core catalytic residue of the enzyme has switched in different families. In addition to their structural similarity, directed evolution has been shown to be able to convert a cysteine protease into an active serine protease. All cellular PA clan proteases are serine proteases, however there are both serine and cysteine protease families of viral proteases. The majority are endopeptidases, with the exception being the S46 family of exopeptidases.
Hub AI
PA clan of proteases AI simulator
(@PA clan of proteases_simulator)
PA clan of proteases
The PA clan (Proteases of mixed nucleophile, superfamily A) is the largest group of proteases with common ancestry as identified by structural homology. Members have a chymotrypsin-like fold and similar proteolysis mechanisms but can have identity of <10%. The clan contains both cysteine and serine proteases (different nucleophiles). PA clan proteases can be found in plants, animals, fungi, eubacteria, archaea and viruses.
The common use of the catalytic triad for hydrolysis by multiple clans of proteases, including the PA clan, represents an example of convergent evolution. The differences in the catalytic triad within the PA clan is also an example of divergent evolution of active sites in enzymes.
In the 1960s, the sequence similarity of several proteases indicated that they were evolutionarily related. These were grouped into the chymotrypsin-like serine proteases (now called the S1 family). As the structures of these, and other proteases were solved by X-ray crystallography in the 1970s and 80s, it was noticed that several viral proteases such as Tobacco Etch Virus protease showed structural homology despite no discernible sequence similarity and even a different nucleophile. Based on structural homology, a superfamily was defined and later named the PA clan (by the MEROPS classification system). As more structures are solved, more protease families have been added to the PA clan superfamily.
The P refers to Proteases of mixed nucleophile. The A indicates that it was the first such clan to be identified (there also exist the PB, PC, PD and PE clans).
Despite retaining as little as 10% sequence identity, PA clan members isolated from viruses, prokaryotes and eukaryotes show structural homology and can be aligned by structural similarity (e.g. with DALI).
PA clan proteases all share a core motif of two β-barrels with covalent catalysis performed by an acid-histidine-nucleophile catalytic triad motif. The barrels are arranged perpendicularly beside each other with hydrophobic residues holding them together as the core scaffold for the enzyme. The triad residues are split between the two barrels so that catalysis takes place at their interface.
In addition to the double β-barrel core, some viral proteases (such as TEV protease) have a long, flexible C-terminal loop that forms a lid that completely covers the substrate and create a binding tunnel. This tunnel contains a set of tight binding pockets such that each side chain of the substrate peptide (P6 to P1’) is bound in a complementary site (S6 to S1’) and specificity is endowed by the large contact area between enzyme and substrate. Conversely, cellular proteases that lack this loop, such as trypsin have broader specificity.
Structural homology indicates that the PA clan members are descended from a common ancestor of the same fold. Although PA clan proteases use a catalytic triad perform 2-step nucleophilic catalysis, some families use serine as the nucleophile whereas others use cysteine. The superfamily is therefore an extreme example of divergent enzyme evolution since during evolutionary history, the core catalytic residue of the enzyme has switched in different families. In addition to their structural similarity, directed evolution has been shown to be able to convert a cysteine protease into an active serine protease. All cellular PA clan proteases are serine proteases, however there are both serine and cysteine protease families of viral proteases. The majority are endopeptidases, with the exception being the S46 family of exopeptidases.