Replisome
View on WikipediaThis article needs additional citations for verification. (July 2018) |

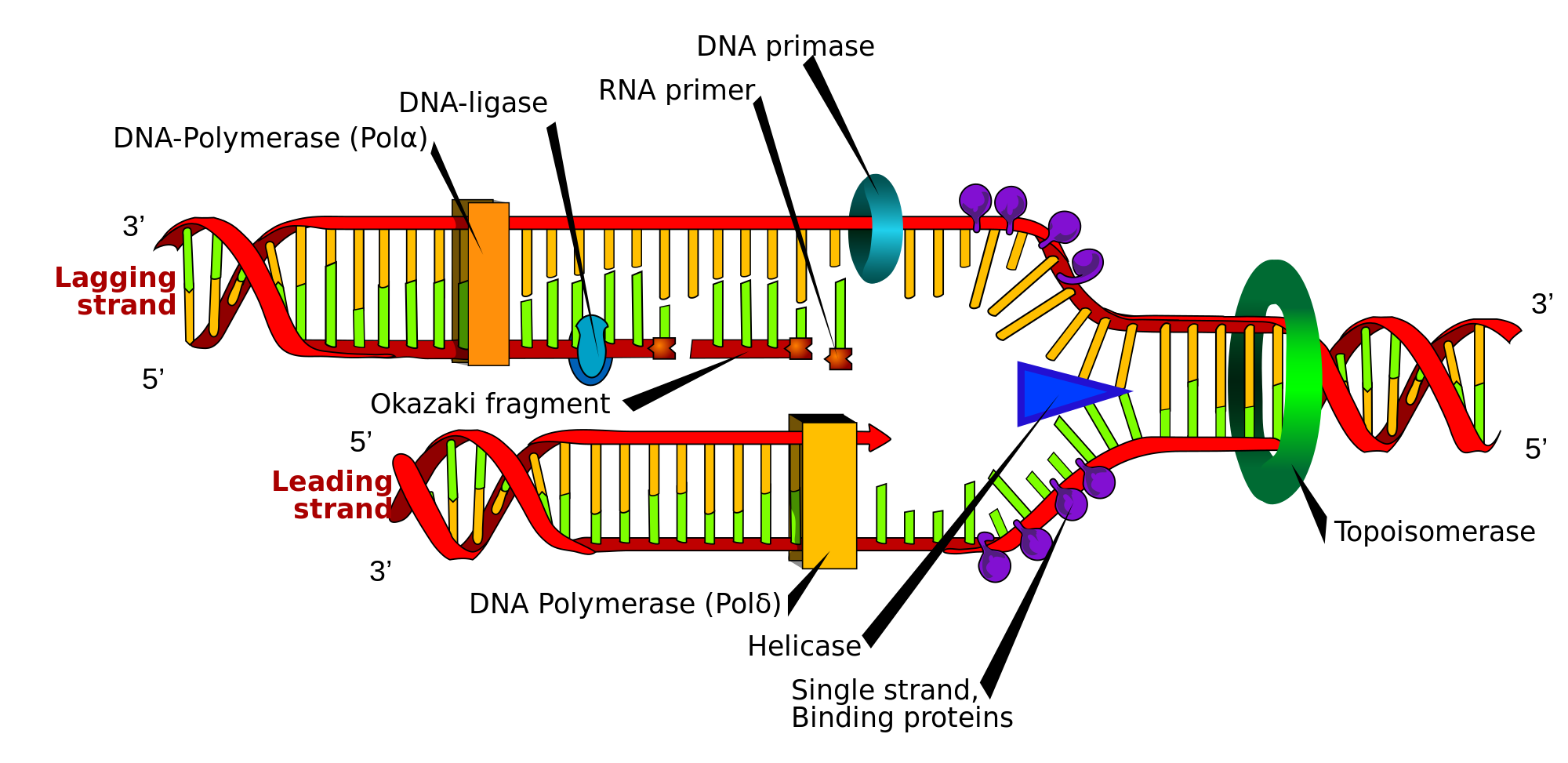
The replisome is a complex molecular machine that carries out replication of DNA. The replisome first unwinds double stranded DNA into two single strands. For each of the resulting single strands, a new complementary sequence of DNA is synthesized. The total result is formation of two new double stranded DNA sequences that are exact copies of the original double stranded DNA sequence.[1]
In terms of structure, the replisome is composed of two replicative polymerase complexes, one of which synthesizes the leading strand, while the other synthesizes the lagging strand. The replisome is composed of a number of proteins including helicase, RFC, PCNA, gyrase/topoisomerase, SSB/RPA, primase, DNA polymerase III, RNAse H, and DNA ligase.
Overview of prokaryotic DNA replication process
[edit]For prokaryotes, each dividing nucleoid (region containing genetic material which is not a nucleus) requires two replisomes for bidirectional replication. The two replisomes continue replication at both forks in the middle of the cell. Finally, as the termination site replicates, the two replisomes separate from the DNA. The replisome remains at a fixed, midcell location in the cell, attached to the membrane, and the template DNA threads through it. DNA is fed through the stationary pair of replisomes located at the cell membrane.
Overview of eukaryotic DNA replication process
[edit]For eukaryotes, numerous replication bubbles form at origins of replication throughout the chromosome. As with prokaryotes, two replisomes are required, one at each replication fork located at the terminus of the replication bubble. Because of significant differences in chromosome size, and the associated complexities of highly condensed chromosomes, various aspects of the DNA replication process in eukaryotes, including the terminal phases, are less well-characterised than for prokaryotes.
Challenges of DNA replication
[edit]The replisome is a system in which various factors work together to solve the structural and chemical challenges of DNA replication. Chromosome size and structure varies between organisms, but since DNA molecules are the reservoir of genetic information for all forms of life, many replication challenges and solutions are the same for different organisms. As a result, the replication factors that solve these problems are highly conserved in terms of structure, chemistry, functionality, or sequence. General structural and chemical challenges include the following:
- Efficient replisome assembly at origins of replication (origin recognition complexes or specific replication origin sequences in some organisms)
- Separating the duplex into the leading and lagging template strands (helicases)
- Protecting the leading and lagging strands from damage after duplex separation (SSB and RPA factors)
- Priming of the leading and lagging template strands (primase or DNA polymerase alpha)
- Ensuring processivity (clamp loading factors, ring-shaped clamp proteins, strand binding proteins)
- High-fidelity DNA replication (DNA polymerase III, DNA polymerase delta, DNA polymerase epsilon. All have intrinsically low error rates because of their structure and chemistry.)
- Error correction (replicative polymerase active sites sense errors; 3' to 5' exonuclease domains of replicative polymerases fix errors)
- Synchronised polymerisation of leading and lagging strands despite anti-parallel structure (replication fork structure, dimerisation of replicative polymerases)
- Primer removal (DNA polymerase I, RNAse H, flap endonucleases such as FEN1, or other DNA repair factors)
- Formation of phosphodiester bonds at gaps between Okazaki fragments (ligase)
In general, the challenges of DNA replication involve the structure of the molecules, the chemistry of the molecules, and, from a systems perspective, the underlying relationships between the structure and the chemistry.
Solving the challenges of DNA replication
[edit]Many of the structural and chemical problems associated with DNA replication are managed by molecular machinery that is highly conserved across organisms. This section discusses how replisome factors solve the structural and chemical challenges of DNA replication.
Replisome assembly
[edit]DNA replication begins at sites called origins of replication. In organisms with small genomes and simple chromosome structure, such as bacteria, there may be only a few origins of replication on each chromosome. Organisms with large genomes and complex chromosome structure, such as humans, may have hundreds, or even thousands, of origins of replication spread across multiple chromosomes.
DNA structure varies with time, space, and sequence, and it is thought that these variations, in addition to their role in gene expression, also play active roles in replisome assembly during DNA synthesis. Replisome assembly at an origin of replication is roughly divided into three phases.
For bacteria:
- Formation of pre-replication complex. DnaA binds to the origin recognition complex and separates the duplex. This attracts DnaB helicase and DnaC, which maintain the replication bubble.
- Formation of pre-initiation complex. SSB binds to the single strand and then gamma (clamp loading factor) binds to SSB.
- Formation of initiation complex. Gamma deposits the sliding clamp (beta) and attracts DNA polymerase III.
For eukaryotes:
- Formation of pre-replication complex. MCM factors bind to the origin recognition complex and separate the duplex, forming a replication bubble.
- Formation of pre-initiation complex. Replication protein A (RPA) binds to the single stranded DNA and then RFC (clamp loading factor) binds to RPA.
- Formation of initiation complex. RFC deposits the sliding clamp (PCNA) and attracts DNA polymerases such as alpha (α), delta (δ), epsilon (ε).
For both bacteria and eukaryotes, the next stage is generally referred to as 'elongation', and it is during this phase that the majority of DNA synthesis occurs.
Separating the duplex
[edit]DNA is a duplex formed by two anti-parallel strands. Following Meselson-Stahl, the process of DNA replication is semi-conservative, whereby during replication the original DNA duplex is separated into two daughter strands (referred to as the leading and lagging strand templates). Each daughter strand becomes part of a new DNA duplex. Factors generically referred to as helicases unwind the duplex.
Helicases
[edit]Helicase is an enzyme which breaks hydrogen bonds between the base pairs in the middle of the DNA duplex. Its doughnut like structure wraps around DNA and separates the strands ahead of DNA synthesis. In eukaryotes, the Mcm2-7 complex acts as a helicase, though which subunits are required for helicase activity is not entirely clear.[2] This helicase translocates in the same direction as the DNA polymerase (3' to 5' with respect to the template strand). In prokaryotic organisms, the helicases are better identified and include dnaB, which moves 5' to 3' on the strand opposite the DNA polymerase.
Unwinding supercoils and decatenation
[edit]
As helicase unwinds the double helix, topological changes induced by the rotational motion of the helicase lead to supercoil formation ahead of the helicase (similar to what happens when you twist a piece of thread).
Gyrase and topoisomerases
[edit]Gyrase (a form of topoisomerase) relaxes and undoes the supercoiling caused by helicase. It does this by cutting the DNA strands, allowing it to rotate and release the supercoil, and then rejoining the strands. Gyrase is most commonly found upstream of the replication fork, where the supercoils form.
Protecting the leading and lagging strands
[edit]
Single-stranded DNA is highly unstable and can form hydrogen bonds with itself that are referred to as 'hairpins' (or the single strand can improperly bond to the other single strand). To counteract this instability, single-strand binding proteins (SSB in prokaryotes and Replication protein A in eukaryotes) bind to the exposed bases to prevent improper ligation.
If you consider each strand as a "dynamic, stretchy string", the structural potential for improper ligation should be obvious.
| The lagging strand without binding proteins. |
|---|
|
An expanded schematic reveals the underlying chemistry of the problem: the potential for hydrogen bond formation between unrelated base pairs.
| Schematic view of newly separated DNA strands without strand binding proteins. |
|---|
|
Binding proteins stabilise the single strand and protected the strand from damage caused by unlicensed chemical reactions.
| The lagging strand coated with binding proteins (*) that prevent improper ligation. |
|---|
|
The combination of a single strand and its binding proteins serves as a better substrate for replicative polymerases than a naked single strand (binding proteins provide extra thermodynamic driving force for the polymerisation reaction). Strand binding proteins are removed by replicative polymerases.
Priming the leading and lagging strands
[edit]From both a structural and chemical perspective, a single strand of DNA by itself (and the associated single strand binding proteins) is not suitable for polymerisation. This is because the chemical reactions catalysed by replicative polymerases require a free 3' OH in order to initiate nucleotide chain elongation. In terms of structure, the conformation of replicative polymerase active sites (which is highly related to the inherent accuracy of replicative polymerases) means these factors cannot start chain elongation without a pre-existing chain of nucleotides, because no known replicative polymerase can start chain elongation de novo.
Priming enzymes, (which are DNA-dependent RNA polymerases), solve this problem by creating an RNA primer on the leading and lagging strands. The leading strand is primed once, and the lagging strand is primed approximately every 1000 (+/- 200) base pairs (one primer for each Okazaki fragment on the lagging strand). Each RNA primer is approximately 10 bases long.
| Single strand of DNA with strand binding proteins (*) and RNA primer added by priming enzymes (UAGCUAUAUAUA). |
|---|
|
The interface at (A*) contains a free 3' OH that is chemically suitable for the reaction catalysed by replicative polymerases, and the "overhang" configuration is structurally suitable for chain elongation by a replicative polymerase. Thus, replicative polymerases can begin chain elongation at (A*).
Primase
[edit]In prokaryotes, the primase creates an RNA primer at the beginning of the newly separated leading and lagging strands.
DNA polymerase alpha
[edit]In eukaryotes, DNA polymerase alpha creates an RNA primer at the beginning of the newly separated leading and lagging strands, and, unlike primase, DNA polymerase alpha also synthesizes a short chain of deoxynucleotides after creating the primer.
Ensuring processivity and synchronisation
[edit]Processivity refers to both speed and continuity of DNA replication, and high processivity is a requirement for timely replication. High processivity is in part ensured by ring-shaped proteins referred to as 'clamps' that help replicative polymerases stay associated with the leading and lagging strands. There are other variables as well: from a chemical perspective, strand binding proteins stimulate polymerisation and provide extra thermodynamic energy for the reaction. From a systems perspective, the structure and chemistry of many replisome factors (such as the AAA+ ATPase features of the individual clamp loading sub-units, along with the helical conformation they adopt), and the associations between clamp loading factors and other accessory factors, also increases processivity.
To this point, according to research by Kuriyan et al.,[3] due to their role in recruiting and binding other factors such as priming enzymes and replicative polymerases, clamp loaders and sliding clamps are at the heart of the replisome machinery. Research has found that clamp loading and sliding clamp factors are absolutely essential to replication, which explains the high degree of structural conservation observed for clamp loading and sliding clamp factors. This architectural and structural conservation is seen in organisms as diverse as bacteria, phages, yeast, and humans. That such a significant degree of structural conservation is observed without sequence homology further underpins the significance of these structural solutions to replication challenges.
Clamp loader
[edit]Clamp loader is a generic term that refers to replication factors called gamma (bacteria) or RFC (eukaryotes). The combination of template DNA and primer RNA is referred to as 'A-form DNA' and it is thought that clamp loading replication proteins (helical heteropentamers) want to associate with A-form DNA because of its shape (the structure of the major/minor groove) and chemistry (patterns of hydrogen bond donors and acceptors).[3][4] Thus, clamp loading proteins associate with the primed region of the strand which causes hydrolysis of ATP and provides energy to open the clamp and attach it to the strand.[3][4]
Sliding clamp
[edit]
Sliding clamp is a generic term that refers to ring-shaped replication factors called beta (bacteria) or PCNA (eukaryotes and archaea). Clamp proteins attract and tether replicative polymerases, such as DNA polymerase III, in order to extend the amount of time that a replicative polymerase stays associated with the strand. From a chemical perspective, the clamp has a slightly positive charge at its centre that is a near perfect match for the slightly negative charge of the DNA strand.
In some organisms, the clamp is a dimer, and in other organisms the clamp is a trimer. Regardless, the conserved ring architecture allows the clamp to enclose the strand.
Dimerisation of replicative polymerases
[edit]Replicative polymerases form an asymmetric dimer at the replication fork by binding to sub-units of the clamp loading factor. This asymmetric conformation is capable of simultaneously replicating the leading and lagging strands, and the collection of factors that includes the replicative polymerases is generally referred to as a holoenzyme. However, significant challenges remain: the leading and lagging strands are anti-parallel. This means that nucleotide synthesis on the leading strand naturally occurs in the 5' to 3' direction. However, the lagging strand runs in the opposite direction and this presents quite a challenge since no known replicative polymerases can synthesise DNA in the 3' to 5' direction.
The dimerisation of the replicative polymerases solves the problems related to efficient synchronisation of leading and lagging strand synthesis at the replication fork, but the tight spatial-structural coupling of the replicative polymerases, while solving the difficult issue of synchronisation, creates another challenge: dimerisation of the replicative polymerases at the replication fork means that nucleotide synthesis for both strands must take place at the same spatial location, despite the fact that the lagging strand must be synthesised backwards relative to the leading strand. Lagging strand synthesis takes place after the helicase has unwound a sufficient quantity of the lagging strand, and this "sufficient quantity of the lagging strand" is polymerised in discrete nucleotide chains called Okazaki fragments.
Consider the following: the helicase continuously unwinds the parental duplex, but the lagging strand must be polymerised in the opposite direction. This means that, while polymerisation of the leading strand proceeds, polymerisation of the lagging strand only occurs after enough of the lagging strand has been unwound by the helicase. At this point, the lagging strand replicative polymerase associates with the clamp and primer in order to start polymerisation. During lagging strand synthesis, the replicative polymerase sends the lagging strand back toward the replication fork. The replicative polymerase disassociates when it reaches an RNA primer. Helicase continues to unwind the parental duplex, the priming enzyme affixes another primer, and the replicative polymerase reassociates with the clamp and primer when a sufficient quantity of the lagging strand has unwound.
Collectively, leading and lagging strand synthesis is referred to as being 'semidiscontinuous'.
High-fidelity DNA replication
[edit]This article is missing information about multifunctional polD of archaea. (July 2022) |
Prokaryotic and eukaryotic organisms use a variety of replicative polymerases, some of which are well-characterised:
- DNA polymerase III
- DNA polymerase delta
- DNA polymerase epsilon
DNA polymerase III
[edit]This polymerase synthesizes leading and lagging strand DNA in bacteria.
DNA polymerase delta
[edit]This polymerase synthesizes lagging strand DNA in eukaryotes.[5] (Thought to form an asymmetric dimer with DNA polymerase epsilon.)[6]
DNA polymerase epsilon
[edit]This polymerase synthesizes leading strand DNA in eukaryotes.[7] (Thought to form an asymmetric dimer with DNA polymerase delta.)[5]
Proof-reading and error correction
[edit]Although rare, incorrect base pairing polymerisation does occur during chain elongation. (The structure and chemistry of replicative polymerases mean that errors are unlikely, but they do occur.) Many replicative polymerases contain an "error correction" mechanism in the form of a 3' to 5' exonuclease domain that is capable of removing base pairs from the exposed 3' end of the growing chain. Error correction is possible because base pair errors distort the position of the magnesium ions in the polymerisation sub-unit, and the structural-chemical distortion of the polymerisation unit effectively stalls the polymerisation process by slowing the reaction.[8] Subsequently, the chemical reaction in the exonuclease unit takes over and removes nucleotides from the exposed 3' end of the growing chain.[9] Once an error is removed, the structure and chemistry of the polymerisation unit returns to normal and DNA replication continues. Working collectively in this fashion, the polymerisation active site can be thought of as the "proof-reader", since it senses mismatches, and the exonuclease is the "editor", since it corrects the errors.
Base pair errors distort the polymerase active site for between 4 and 6 nucleotides, which means, depending on the type of mismatch, there are up to six chances for error correction.[8] The error sensing and error correction features, combined with the inherent accuracy that arises from the structure and chemistry of replicative polymerases, contribute to an error rate of approximately 1 base pair mismatch in 108 to 1010 base pairs.
| Schematic view of correct base pairs followed by 8 possible base pair mismatches.[10] |
|---|
|
Errors can be classified in three categories: purine-purine mismatches, pyrimidine-pyrimidine mismatches, and pyrimidine-purine mismatches. The chemistry of each mismatch varies, and so does the behaviour of the replicative polymerase with respect to its mismatch sensing activity.
The replication of bacteriophage T4 DNA upon infection of E. coli is a well-studied DNA replication system. During the period of exponential DNA increase at 37°C, the rate of elongation is 749 nucleotides per second.[11] The mutation rate during replication is 1.7 mutations per 108 base pairs.[12] Thus DNA replication in this system is both very rapid and highly accurate.
Primer removal and nick ligation
[edit]There are two problems after leading and lagging strand synthesis: RNA remains in the duplex and there are nicks between each Okazaki fragment in the lagging duplex. These problems are solved by a variety of DNA repair enzymes that vary by organism, including: DNA polymerase I, DNA polymerase beta, RNAse H, ligase, and DNA2. This process is well-characterised in bacteria and much less well-characterised in many eukaryotes.
In general, DNA repair enzymes complete the Okazaki fragments through a variety of means, including: base pair excision and 5' to 3' exonuclease activity that removes the chemically unstable ribonucleotides from the lagging duplex and replaces them with stable deoxynucleotides. This process is referred to as 'maturation of Okazaki fragments', and ligase (see below) completes the final step in the maturation process.
| RNA-DNA duplex with ribonucleotides added by a priming enzyme (-) and deoxynucleotides added by a replicative polymerase (+). |
|---|
|
Primer removal and nick ligation can be thought of as DNA repair processes that produce a chemically-stable, error-free duplex. To this point, with respect to the chemistry of an RNA-DNA duplex, in addition to the presence of uracil in the duplex, the presence of ribose (which has a reactive 2' OH) tends to make the duplex much less chemically-stable than a duplex containing only deoxyribose (which has a non-reactive 2' H).
DNA polymerase I
[edit]DNA polymerase I is an enzyme that repairs DNA.
RNAse H
[edit]RNAse H is an enzyme that removes RNA from an RNA-DNA duplex.
Ligase
[edit]After DNA repair factors replace the ribonucleotides of the primer with deoxynucleotides, a single gap remains in the sugar-phosphate backbone between each Okazaki fragment in the lagging duplex. An enzyme called DNA ligase connects the gap in the backbone by forming a phosphodiester bond between each gap that separates the Okazaki fragments. The structural and chemical aspects of this process, generally referred to as 'nick translation', exceed the scope of this article.
| A schematic view of the new, lagging strand daughter DNA duplex is shown below, along with the sugar-phosphate backbone. |
|---|
|
| The finished duplex: |
|---|
|
Replication stress
[edit]Replication stress can result in a stalled replication fork. One type of replicative stress results from DNA damage such as inter-strand cross-links (ICLs). An ICL can block replicative fork progression due to failure of DNA strand separation. In vertebrate cells, replication of an ICL-containing chromatin template triggers recruitment of more than 90 DNA repair and genome maintenance factors.[13] These factors include proteins that perform sequential incisions and homologous recombination.
History
[edit]Katherine Lemon and Alan Grossman showed using Bacillus subtilis that replisomes do not move like trains along a track but DNA is actually fed through a stationary pair of replisomes located at the cell membrane. In their experiment, the replisomes in B. subtilis were each tagged with green fluorescent protein, and the location of the complex was monitored in replicating cells using fluorescence microscopy. If the replisomes moved like a train on a track, the polymerase-GFP protein would be found at different positions in each cell. Instead, however, in every replicating cell, replisomes were observed as distinct fluorescent foci located at or near midcell. Cellular DNA stained with a blue fluorescent dye (DAPI) clearly occupied most of the cytoplasmic space.[14]
References
[edit]- ^ Yao, Nina Y.; O'Donnell, Mike (2010). "SnapShot: The Replisome". Cell. 141 (6). Elsevier BV: 1088–1088.e1. doi:10.1016/j.cell.2010.05.042. ISSN 0092-8674. PMC 4007198. PMID 20550941.
- ^ Bochman ML, Schwacha A (July 2008). "The Mcm2-7 complex has in vitro helicase activity". Mol. Cell. 31 (2): 287–93. doi:10.1016/j.molcel.2008.05.020. PMID 18657510.
- ^ a b c Kelch BA, Makino DL, O'Donnell M, Kuriyan J (2012). "Clamp loader ATPases and the evolution of DNA replication machinery". BMC Biol. 10: 34. doi:10.1186/1741-7007-10-34. PMC 3331839. PMID 22520345.
- ^ a b Bowman GD, O'Donnell M, Kuriyan J (June 2004). "Structural analysis of a eukaryotic sliding DNA clamp-clamp loader complex". Nature. 429 (6993): 724–30. doi:10.1038/nature02585. PMID 15201901. S2CID 4346799.
- ^ a b Swan MK, Johnson RE, Prakash L, Prakash S, Aggarwal AK (September 2009). "Structural basis of high-fidelity DNA synthesis by yeast DNA polymerase delta". Nat. Struct. Mol. Biol. 16 (9): 979–86. doi:10.1038/nsmb.1663. PMC 3055789. PMID 19718023.
- ^ Miyabe I, Kunkel TA, Carr AM (December 2011). "The major roles of DNA polymerases epsilon and delta at the eukaryotic replication fork are evolutionarily conserved". PLOS Genet. 7 (12) e1002407. doi:10.1371/journal.pgen.1002407. PMC 3228825. PMID 22144917.
- ^ Pursell ZF, Isoz I, Lundström EB, Johansson E, Kunkel TA (July 2007). "Yeast DNA polymerase epsilon participates in leading-strand DNA replication". Science. 317 (5834): 127–30. doi:10.1126/science.1144067. PMC 2233713. PMID 17615360.
- ^ a b Johnson SJ, Beese LS (March 2004). "Structures of mismatch replication errors observed in a DNA polymerase". Cell. 116 (6): 803–16. doi:10.1016/S0092-8674(04)00252-1. PMID 15035983.
- ^ Jiricny J (March 2004). "Unfaithful DNA polymerase caught in the act" (PDF). Mol. Cell. 13 (6): 768–9. doi:10.1016/S1097-2765(04)00149-2. PMID 15053870.
- ^ "Mutagenesis and DNA repair". ATDBio Ltd.
- ^ McCarthy D, Minner C, Bernstein H, Bernstein C (1976). "DNA elongation rates and growing point distributions of wild-type phage T4 and a DNA-delay amber mutant". J. Mol. Biol. 106 (4): 963–81. doi:10.1016/0022-2836(76)90346-6. PMID 789903.
- ^ Drake JW (1970) The Molecular Basis of Mutation. Holden-Day, San Francisco ISBN 0816224501 ISBN 978-0816224500
- ^ Räschle M, Smeenk G, Hansen RK, Temu T, Oka Y, Hein MY, Nagaraj N, Long DT, Walter JC, Hofmann K, Storchova Z, Cox J, Bekker-Jensen S, Mailand N, Mann M (2015). "DNA repair. Proteomics reveals dynamic assembly of repair complexes during bypass of DNA cross-links". Science. 348 (6234) 1253671. doi:10.1126/science.1253671. PMC 5331883. PMID 25931565.
- ^ Foster JB, Slonczewski J (2010). Microbiology: An Evolving Science (Second ed.). New York: W. W. Norton & Company. ISBN 978-0-393-93447-2.
Further reading
[edit]- Pomerantz RT, O'Donnell M (April 2007). "Replisome mechanics: insights into a twin DNA polymerase machine". Trends Microbiol. 15 (4): 156–64. doi:10.1016/j.tim.2007.02.007. PMID 17350265.
External links
[edit]- DNA+replisome at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
Replisome
View on GrokipediaIntroduction to DNA Replication
The Replication Fork and Semi-Conservative Mechanism
The replication fork is the Y-shaped region formed during DNA replication where the double-stranded parental DNA helix unwinds, separating the two antiparallel strands to expose single-stranded templates for the synthesis of new complementary strands.[4] This structure arises as replication proceeds bidirectionally from an origin, creating two forks that move away from each other, facilitating the duplication of the entire genome.[5] DNA replication follows a semi-conservative mechanism, in which each parental strand serves as a template for the synthesis of a new daughter strand, resulting in two hybrid double helices each containing one original and one newly synthesized strand.[6] This process relies on Watson-Crick base pairing, where adenine pairs with thymine and guanine with cytosine, ensuring the faithful copying of genetic information through complementary hydrogen bonding between nucleotide bases.[7] The semi-conservative nature was experimentally confirmed using density-labeled DNA in Escherichia coli, demonstrating that after one round of replication, all DNA molecules have intermediate density, shifting to half hybrid and half light after a second round.[6] At the replication fork, synthesis occurs asymmetrically due to the antiparallel orientation of DNA strands and the 5' to 3' polarity of nucleotide addition. The leading strand is synthesized continuously in the 5' to 3' direction toward the advancing fork, while the lagging strand is synthesized discontinuously away from the fork as short segments known as Okazaki fragments, each initiated from an RNA primer and later joined.[4] This can be visualized as a dynamic Y-shaped structure progressing along the DNA, with the leading strand forming a straight extension and the lagging strand looping back to allow repeated priming and extension in the required direction. The rate of fork progression varies by organism, typically around 1000 nucleotides per second in E. coli at 37°C, enabling rapid genome duplication.[8] In eukaryotes, the rate is slower, approximately 50 nucleotides per second, reflecting the larger genome size and additional regulatory complexity.[9] The replisome, a multi-protein complex, coordinates this fork movement to ensure efficient and accurate replication.[10]Essential Challenges in Replicating DNA
One of the primary biophysical challenges in DNA replication is the unwinding of the stable double helix, which generates positive supercoils ahead of the replication fork due to the accumulation of torsional stress as the strands separate. This positive linking number difference (ΔLk) increases by approximately one turn per 10 base pairs unwound, creating mechanical strain that resists further fork progression and can halt replication if unresolved.[11] The process demands significant energy, primarily supplied through ATP hydrolysis, to overcome the torsional barriers inherent in separating the intertwined strands.[12] Following unwinding, the separated single-stranded DNA faces biochemical vulnerabilities, including a propensity to reanneal spontaneously due to base-pairing affinities and susceptibility to degradation by cellular nucleases, which could compromise template integrity and replication accuracy. Additionally, the inherent 5' to 3' directionality of DNA polymerases imposes a fundamental asymmetry: while the leading strand can be synthesized continuously in the direction of fork movement, the antiparallel lagging strand requires discontinuous synthesis in short segments, complicating coordinated progression and increasing the risk of gaps or errors at junctions.[13] This polarity limitation necessitates primers for initiation, as polymerases cannot start de novo; short RNA primers are thus required to provide the necessary 3' hydroxyl group, but their subsequent removal and replacement with DNA introduce further challenges to ensure seamless strand completion without discontinuities.[14] To replicate entire genomes faithfully, DNA synthesis must achieve high processivity, enabling polymerases to traverse long stretches—often millions of base pairs—without dissociating, while maintaining exceptional fidelity with an overall error rate below 1 in 10^9 base pairs incorporated. These demands arise from the need to copy vast genetic information accurately across cell divisions, where even minor lapses could lead to mutations accumulating over generations. Multi-enzyme complexes like the replisome address these obstacles through coordinated mechanisms.[15]Prokaryotic Replisome
Core Components and Assembly
In Escherichia coli, replication initiation begins at the chromosomal origin oriC, where the initiator protein DnaA binds to multiple DnaA boxes within the ~250 base pair sequence, forming a nucleoprotein complex that promotes localized unwinding of an AT-rich region known as the DNA unwinding element (DUE).[16] This unwinding exposes single-stranded DNA, allowing the helicase loader DnaC to deliver two hexameric DnaB helicases in an ATP-dependent manner: one onto each separated strand, with the helicases encircling the DNA in opposite orientations to facilitate bidirectional replication fork progression.[17] Single-strand binding (SSB) proteins then rapidly coat the exposed single-stranded DNA, preventing reannealing and protecting it from nucleases while stabilizing the structure for subsequent replisome assembly.[16] The core components of the prokaryotic replisome include the DnaB helicase, which unwinds double-stranded DNA at the replication fork in a 5' to 3' direction; DnaG primase, which synthesizes short RNA primers essential for DNA polymerase initiation; and the DNA polymerase III (Pol III) holoenzyme, comprising the core polymerase with α (catalytic subunit), ε (3'→5' exonuclease for proofreading), and θ (stabilizing subunit) subunits.[18] The holoenzyme is completed by the β sliding clamp (a dimeric ring that tethers the polymerase to DNA for processivity) and the γ complex clamp loader (composed of γ, δ, δ', χ, and ψ subunits), which assembles the β clamps onto DNA.[19] SSB proteins, forming tetramers that bind cooperatively to single-stranded DNA, further support replisome function by coordinating protein interactions at the fork.[20] Assembly of the replisome proceeds with DnaG primase associating directly with the DnaB helicase via protein-protein interactions, enabling priming on the single-stranded templates generated by helicase activity; this interaction is critical for the synthesis of RNA primers (typically 10-12 nucleotides long) on both leading and lagging strands.[21] The γ complex then recruits the Pol III core to the primed sites by loading β clamps in an ATP-dependent process, forming a stable holoenzyme that initiates DNA synthesis.[22] This assembly occurs bidirectionally from oriC, establishing two replisomes that migrate in opposite directions to replicate the circular chromosome.[16] The loading of the β clamp exemplifies the energy coupling in replisome assembly, where the γ complex harnesses ATP hydrolysis to open and close the ring-shaped clamp. ATP binding to the AAA+ domains of the γ complex induces a conformational change that pries open the β dimer at its dimer interface, allowing the clamp to encircle double-stranded DNA at a primer-template junction:Enzymatic Functions and Coordination
The DnaB helicase forms a hexameric ring that encircles and translocates along the single-stranded lagging strand template in the 5' to 3' direction, unwinding the parental DNA duplex to expose single-stranded templates for replication.[24] Within the assembled prokaryotic replisome, this unwinding activity proceeds at a coordinated rate of approximately 1000 base pairs per second, facilitated by direct interactions with other replisome components that enhance its velocity beyond the slower intrinsic rate observed in isolation.[25] The single-stranded DNA-binding protein (SSB) plays a crucial supporting role by coating the newly exposed single-stranded regions, thereby preventing the formation of inhibitory secondary structures and protecting against nuclease degradation to maintain template accessibility.[26] The DnaG primase associates transiently with the DnaB helicase through direct protein-protein interactions, enabling it to recognize specific sites on the lagging strand template and synthesize short RNA primers of 10-12 nucleotides in length.[21] These primers are generated periodically, approximately every 1000-2000 nucleotides, to initiate the synthesis of each Okazaki fragment on the discontinuous lagging strand.[27] This priming action is tightly coupled to helicase progression, ensuring that primer synthesis occurs in synchrony with DNA unwinding without excessive delays that could stall the fork. The DNA polymerase III (Pol III) holoenzyme functions as a dimeric complex, with two core polymerases tethered by the τ subunits of the DnaX complex, allowing simultaneous and coordinated extension of the leading strand continuously and the lagging strand discontinuously from each new primer.[28] The τ subunit not only links the polymerases but also directly contacts the DnaB helicase, promoting rapid fork progression by coupling polymerase activity to helicase unwinding.[29] Topoisomerases, such as DNA gyrase, relieve the positive supercoils that accumulate ahead of the advancing fork to prevent torsional stress from impeding replisome movement.[30] This dynamic interplay among helicase, primase, polymerase, and accessory proteins ensures efficient, bidirectional DNA synthesis during the elongation phase of replication in prokaryotes.Achieving Processivity and Fidelity
The processivity of DNA polymerase III (Pol III) in prokaryotes, particularly in Escherichia coli, is dramatically enhanced by the β-dimer sliding clamp, a ring-shaped homodimer that encircles the DNA double helix and tethers the polymerase to the template, allowing continuous synthesis over distances exceeding 500 kb without dissociation.[31] This topological linkage prevents the core polymerase from sliding off the DNA during replication, enabling the holoenzyme to replicate the entire bacterial chromosome in a single binding event.[32] The β-clamp is loaded onto primed DNA by the γ-complex, an ATP-driven pentameric clamp loader composed of subunits δ, δ', three γ (or τ) units, χ, and ψ, which assembles the closed ring around the DNA in an ATP-dependent manner.00463-9) The complex recognizes primer-template junctions, uses ATP hydrolysis to open the clamp and encircle the DNA, and then closes it, ensuring efficient and repeated loading at multiple sites along the lagging strand.00381-X) This mechanism is conserved across domains of life, with the eukaryotic homolog proliferating cell nuclear antigen (PCNA) serving a similar role in enhancing polymerase processivity.00463-9) Fidelity during synthesis is maintained by the ε subunit of Pol III, which possesses 3'→5' exonuclease activity that proofreads and excises mismatched nucleotides immediately after incorporation, reducing the base substitution error rate by approximately 100- to 1,000-fold, from ~10^{-5} to ~10^{-7} per nucleotide.[33][34] This proofreading occurs via a direct interaction between ε and the β-clamp, stabilizing the replisome and allowing rapid correction without halting replication.[35] On the lagging strand, RNA primers synthesized by primase must be removed and replaced with DNA to complete Okazaki fragments; this is achieved by DNA polymerase I (Pol I), which uses its 5'→3' exonuclease activity to degrade the RNA primers while simultaneously filling the resulting gaps via its polymerase domain. The nicks left after gap filling are then sealed by DNA ligase, which catalyzes the formation of phosphodiester bonds in an ATP- or NAD+-dependent reaction, ensuring a continuous DNA backbone. Even with these mechanisms, residual errors require post-replication correction by the mismatch repair (MMR) system, involving proteins like MutS, MutL, and MutH, which recognize and excise mismatched bases on the newly synthesized strand, achieving an overall replication fidelity of 10^{-9} to 10^{-10} errors per base pair.[36][37]Eukaryotic Replisome
Initiation at Replication Origins
In eukaryotic cells, initiation of DNA replication begins with the recognition of specific genomic sites known as origins of replication. The origin recognition complex (ORC), a heterohexameric protein composed of subunits Orc1–6, binds to autonomously replicating sequence (ARS) elements, which are AT-rich DNA motifs that serve as origins.[38] This binding occurs throughout the cell cycle but is particularly stable during G1 phase, where ORC recruits the AAA+ ATPase Cdc6 and the licensing factor Cdt1 to the origin DNA.00182-3) Cdc6 associates with ORC to form a platform that facilitates the delivery of Cdt1-bound MCM2-7 complexes, ensuring precise origin selection.[39] During G1 phase, the pre-replicative complex (pre-RC) assembles through the loading of the MCM2-7 helicase onto double-stranded DNA. ORC, Cdc6, and Cdt1 cooperatively load two MCM2-7 hexamers in a head-to-head orientation, forming a double hexamer that encircles the DNA without unwinding it.01303-8) This process, termed replication licensing, inactivates the helicase and prepares multiple origins for activation, contrasting with the single oriC origin used in prokaryotes for assembly.00423-7) The double hexamer structure provides a bidirectional platform for subsequent replisome formation, with each MCM2-7 ring poised to unwind DNA strands upon activation. Eukaryotic genomes utilize thousands of such origins—estimated at around 50,000 in humans—to enable parallel replication of large chromosomes during S phase.[40] Activation of the pre-RC occurs at the onset of S phase, triggered by phosphorylation events that convert the inactive MCM2-7 double hexamer into active helicases. The Dbf4-dependent kinase (DDK), consisting of Cdc7 and Dbf4, first phosphorylates specific MCM subunits (notably MCM2, 4, and 6), promoting the recruitment of Cdc45 and the GINS complex.00654-4) Subsequently, S-phase cyclin-dependent kinase (S-CDK) phosphorylates additional sites on MCM and associated factors, stabilizing the assembly of two Cdc45–MCM–GINS (CMG) helicases that encircle single-stranded DNA and initiate bidirectional unwinding.[41] This coordinated phosphorylation ensures timely origin firing and replisome activation. To maintain genomic stability, eukaryotic cells enforce strict regulation of replication licensing to prevent re-replication within a single cell cycle. Licensing factors such as Cdc6, Cdt1, and ORC are tightly controlled: Cdc6 and Cdt1 are degraded or inhibited by geminin (in higher eukaryotes) and CDK-mediated phosphorylation during S, G2, and M phases, while ORC activity is modulated by cyclin-dependent kinases.[42] These mechanisms ensure that pre-RC formation is restricted to late mitosis and G1, blocking new MCM2-7 loading after S phase entry and averting catastrophic DNA over-replication.01566-7)Structural Organization and Key Proteins
The eukaryotic replisome functions as a massive macromolecular assembly of approximately 2 MDa in mass and incorporating more than 20 distinct subunits, enabling coordinated unwinding and polymerization of DNA during replication.[43] This modular architecture positions key enzymatic components in a spatially organized manner around the replication fork, with the central CMG helicase serving as the core motor that drives fork progression.[44] At the heart of the replisome lies the CMG complex, an 11-subunit replicative helicase composed of the heterohexameric MCM2-7 ATPase ring as the primary motor, the regulatory subunit Cdc45, and the GINS heterotetramer (consisting of Psf1, Psf2, Psf3, and Sld5).[45] The MCM2-7 ring encircles and translocates along the leading-strand template in a 3' to 5' direction, powered by ATP hydrolysis, while Cdc45 and GINS stabilize the complex and facilitate DNA unwinding at rates of approximately 25-50 base pairs per second in vivo.[46] This helicase activity generates single-stranded DNA templates for polymerization, with the eukaryotic CMG sharing mechanistic homology to prokaryotic DnaB helicase in its ring-shaped, ATP-dependent unwinding.[44] Primer synthesis is initiated by the Pol α-primase holoenzyme, a four-subunit complex (Pol1, Pol12, Pri1, and Pri2) that generates short RNA-DNA primers of about 8-12 nucleotides RNA followed by 20-30 nucleotides DNA, primarily on the lagging strand to initiate Okazaki fragments.[47] Following primer formation, a handover occurs: the leading strand is extended continuously by DNA polymerase ε (Pol ε), a four-subunit enzyme (Pol2, Dpb2, Dpb3, Dpb4) that directly associates with the trailing face of the CMG complex to form a stable 15-subunit CMGE holoenzyme, ensuring coupled helicase-polymerase activity.[45] In contrast, the lagging strand is synthesized discontinuously by DNA polymerase δ (Pol δ), a three-subunit core (Pol3, Pol31, Pol32) that relies on the PCNA trimeric sliding clamp for high processivity.[47] Several accessory proteins support the core replisome's efficiency and fidelity. Replication protein A (RPA), a heterotrimeric complex, coats exposed single-stranded DNA to prevent reannealing and secondary structure formation, thereby facilitating template access for polymerases.[44] The RFC clamp loader, a five-subunit AAA+ ATPase, recognizes primed DNA sites and loads the PCNA trimer onto the lagging-strand template, encircling the DNA to tether Pol δ and enable rapid, processive synthesis.[47] Additionally, Flap endonuclease 1 (Fen1) processes the RNA primers and flaps generated during Okazaki fragment maturation on the lagging strand, creating ligatable nicks for final sealing by DNA ligase I.[48]Coordination of Leading and Lagging Strand Synthesis
In eukaryotic DNA replication, the replisome coordinates continuous leading-strand synthesis by DNA polymerase ε (Pol ε) with discontinuous lagging-strand synthesis by DNA polymerase δ (Pol δ), ensuring balanced progression despite the antiparallel nature of DNA strands. This synchronization is achieved through dynamic interactions within the Cdc45-MCM-GINS (CMG) helicase complex, which unwinds DNA at the fork while facilitating polymerase activities on both strands. The process maintains replication fidelity and efficiency, adapting to eukaryotic-specific challenges like chromatin packaging. On the lagging strand, Okazaki fragments are typically 100-200 nucleotides long, synthesized by Pol δ in a processive manner facilitated by loop formation, akin to the trombone model observed in prokaryotes where polymerase dimerization aids coordination. As the replisome advances, the lagging-strand template loops back, allowing Pol δ, bound to proliferating cell nuclear antigen (PCNA), to recycle and initiate new fragments upon priming without dissociating from the fork. This looping mechanism, supported by interactions with the CMG helicase, enables rapid cycling between fragment synthesis and release, preventing uncoupled fork progression. Polymerase switching is critical for efficient elongation, involving a timely handover from the primase-polymerase complex Pol α, which synthesizes short RNA-DNA primers (∼10 nt RNA + ∼20-30 nt DNA), to the replicative polymerases Pol δ and Pol ε. This transition is mediated by direct interactions between Pol α and the MCM helicase subunits within the CMG complex, which positions Pol ε for leading-strand takeover and displaces Pol α to allow Pol δ access on the lagging strand. In reconstituted systems, this switching suppresses primer degradation and ensures seamless extension, with Pol δ initially contributing to leading-strand priming before Pol ε dominance. Replication protein A (RPA) coats single-stranded DNA (ssDNA) exposed at the fork, stabilizing templates and preventing secondary structures, but must be displaced during polymerase traversal of primer-template junctions. Pol δ and Pol ε coordinate RPA eviction through direct binding and steric exclusion as they extend nascent strands, with RPA's winged-helix domains facilitating handoff from Pol α-primase to Pol δ on the lagging strand. This dynamic displacement, observed in biochemical assays, maintains ssDNA protection while allowing continuous synthesis without replication stalling. Topoisomerase II (Topo II) plays an essential role in decatenation during elongation, resolving intertwinings (catenanes) that accumulate between newly replicated sister chromatids as the fork progresses. By introducing transient double-strand breaks and strand passage, Topo II alleviates topological stress ahead of the replisome, promoting fork convergence and completion of replication. Inhibition of Topo II leads to persistent catenanes and replication defects, underscoring its coordination with the CMG-polymerase machinery. Eukaryotic replication forks progress at slower speeds, typically 25-50 base pairs per second in vivo, compared to prokaryotic rates, primarily due to chromatin barriers that impede helicase and polymerase movement. Nucleosomes cause transient pausing, requiring histone chaperones and remodelers for disassembly and reassembly, which temporarily halts fork advancement to preserve chromatin integrity.Comparative Aspects and Structural Insights
Similarities and Differences Between Prokaryotes and Eukaryotes
The replisomes of prokaryotes and eukaryotes share fundamental architectural and mechanistic principles that enable the coordinated unwinding and synthesis of DNA. Both utilize ring-shaped helicases to unwind the parental duplex: prokaryotes employ the hexameric DnaB helicase, which translocates 5'→3' along the lagging strand, while eukaryotes use the 11-subunit CMG (Cdc45-Mcm2-7-GINS) complex, a hexameric ring that encircles and translocates 3'→5' along the leading strand. These helicases are powered by ATP hydrolysis, consuming one ATP per 1-2 nucleotides unwound in both systems. Processivity is enhanced by sliding clamps that tether polymerases to DNA: the dimeric β-clamp in prokaryotes and the trimeric PCNA in eukaryotes, both of which are loaded onto primed DNA by pentameric AAA+ ATPases—the γ-complex in prokaryotes and replication factor C (RFC) in eukaryotes—that hydrolyze ATP to open and close the clamps. DNA polymerization proceeds exclusively in the 5'→3' direction in both domains, with continuous synthesis on the leading strand and discontinuous Okazaki fragment production on the lagging strand, ensuring semi-conservative replication. Despite these conserved elements, prokaryotic and eukaryotic replisomes exhibit domain-specific adaptations reflecting their genomic contexts. Prokaryotic replisomes, exemplified by the Escherichia coli system, are structurally simpler, comprising fewer core components dominated by DNA polymerase III (Pol III) holoenzyme, which includes three polymerase cores (one for the leading strand and two for the lagging strand). They operate rapidly from a single chromosomal origin, unhindered by chromatin, achieving high processivity over distances exceeding 100 kb without dissociation. In contrast, eukaryotic replisomes are larger and more complex, assembling at thousands of origins per cell cycle and requiring extensive regulation by cyclin-dependent kinases for initiation and progression. Eukaryotes divide labor between specialized polymerases—Pol ε primarily for the leading strand and Pol δ for the lagging strand—while navigating chromatin barriers through dedicated remodeling factors and histone chaperones, such as FACT (facilitates chromatin transcription), which redeposits histones behind the fork; the Mcm2 subunit of CMG also directly binds core histones to aid this process. Unlike prokaryotes, eukaryotic lagging-strand Okazaki fragments are shorter (100-200 nucleotides versus 1-2 kb), reflecting slower primer extension and coordinated polymerase handoffs. These differences manifest in distinct replication dynamics, as summarized below:| Feature | Prokaryotes (e.g., E. coli) | Eukaryotes (e.g., yeast/human) | Source |
|---|---|---|---|
| Replication speed | 600–1,000 nt/s | 25–100 nt/s | PNAS 2005; Mol Cell 2016 |
| Processivity | >100 kb | 10–50 kb | JSM Biochem Mol Biol 2016; Nature 2017 |
| Fidelity (error rate with proofreading, pre-MMR) | ~10^{-7} per base | ~10^{-7} per base | Mol Cell 2016 |