Recent from talks
Nothing was collected or created yet.
Haplogroup E-M215
View on WikipediaHaplogroup
| |
|---|---|
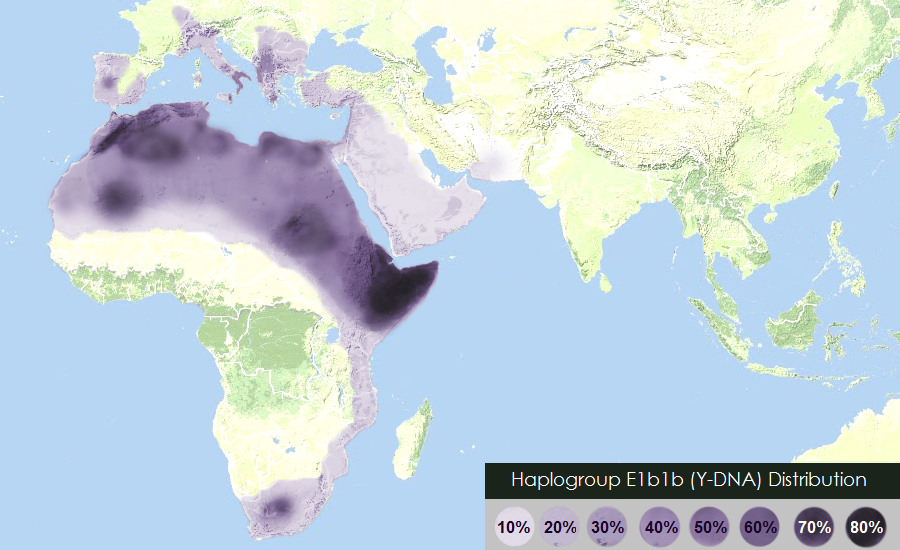
 Geographic distribution of the haplogroup E1b1b | |
| Possible time of origin | 47,500—22,400 BP[1][2][3] |
| Coalescence age | 34,800 BP[4] |
| Possible place of origin | East Africa[5][1] |
| Ancestor | E-P2 |
| Descendants |
|
| Defining mutations | M215 |
E-M215 or E1b1b, formerly known as E3b, is a major human Y-chromosome DNA haplogroup. E-M215 has two basal branches, E-M35 and E-M281. E-M35 is primarily distributed in North Africa and the Horn of Africa, and occurs at moderate frequencies in the Middle East, Europe, and Southern Africa. E-M281 occurs at a low frequency in Ethiopia.
Origins
[edit]The origins of E-M215 were dated by Cruciani in 2007 to about 22,400 years ago in East Africa.[3][Note 1]

Ancient DNA
[edit]According to Lazaridis et al. (2016), Natufian skeletal remains from the ancient Levant predominantly carried the Y-DNA haplogroup E1b1b. Of the five Natufian specimens analyzed for paternal lineages, three belonged to the E1b1b1b2(xE1b1b1b2a, E1b1b1b2b), E1b1(xE1b1a1, E1b1b1b1) and E1b1b1b2(xE1b1b1b2a, E1b1b1b2b) subclades (60%). Haplogroup E1b1b was also found at moderate frequencies among fossils from the ensuing Pre-Pottery Neolithic B culture, with the E1b1b1 and E1b1b1b2(xE1b1b1b2a, E1b1b1b2b) subclades observed in two of seven PPNB specimens (~29%). The scientists suggest that the Levantine early farmers may have spread southward into East Africa, bringing along Western Eurasian and Basal Eurasian ancestral components separate from that which would arrive later in North Africa.
Additionally, haplogroup E1b1b1 has been found in an ancient Egyptian mummy excavated at the Abusir el-Meleq archaeological site in Middle Egypt, which dates from a period between the late New Kingdom and the Roman era.[6] Fossils at the Iberomaurusian site of Ifri N'Amr Ou Moussa in Morocco, which have been dated to around 5,000 BCE, also carried haplotypes related to the E1b1b1b1a (E-M81) subclade. These ancient individuals bore an autochthonous Maghrebi genomic component that peaks among modern North Africans, indicating that they were ancestral to populations in the area.[7] The E1b1b haplogroup has likewise been observed in ancient Guanche fossils excavated in Gran Canaria and Tenerife on the Canary Islands, which have been radiocarbon-dated to between the 7th and 11th centuries CE. The clade-bearing individuals that were analysed for paternal DNA were inhumed at the Tenerife site, with all of these specimens found to belong to the E1b1b1b1a1 or E-M183 subclade (3/3; 100%).[8]
Loosdrecht et al. (2018) analysed genome-wide data from seven ancient Iberomaurusian individuals from the Grotte des Pigeons near Taforalt in eastern Morocco. The fossils were directly dated to between 15,100 and 13,900 calibrated years before present. The scientists found that five male specimens with sufficient nuclear DNA preservation belonged to the E1b1b1a1 (M78) subclade, with one skeleton bearing the E1b1b1a1b1 parent lineage to E-V13, another male specimen belonged to E1b1b (M215*).[9] Martiniano et al. (2022) later reassigned all the Taforalt samples to haplogroup E-M78 and none to E-L618, the predecessor to E-V13.[10]
Distribution
[edit]In Africa, E-M215 is distributed in highest frequencies in the Horn of Africa and North Africa, specifically in the countries Somalia and Morocco, whence it has in recent millennia expanded as far south as South Africa, and northwards into Western Asia and Europe (especially the Mediterranean and the Balkans).[11][12][13][14] E-M281 has been found in Ethiopia.[12]
Almost all E-M215 men are also in E-M35. In 2004, M215 was found to be older than M35 when individuals were found who have the M215 mutation, but do not have M35 mutation.[11] In 2013, Di Cristofaro et al. (2013) found one individual in Khorasan, North-East Iran to be positive for M215 but negative for M35.[15]
E-M215 and E-M35 are quite common among Afroasiatic speakers. The linguistic group and carriers of E-M35 lineage have a high probability to have arisen and dispersed together from the Afroasiatic Urheimat.[16] Amongst populations with an Afro-Asiatic speaking history, a significant proportion of Jewish male lineages are E-M35.[17] Haplogroup E-M35, which accounts for approximately 18%[12] to 20%[18][19] of Ashkenazi and 8.6%[20] to 30%[12] of Sephardi Y-chromosomes, appears to be one of the major founding lineages of the Jewish population.[21][Note 2]
E-M215 association with endurance
[edit]Moran et al. (2004) observed that among Y-DNA (paternal) clades borne by elite endurance athletes in Ethiopia, the haplogroup E3b1 was negatively correlated with elite athletic endurance performance,[22] whereas the haplogroups E*, E3*, K*(xP),[22] and J*(xJ2) were significantly more frequent among the elite endurance athletes.[22]
Subclades
[edit]E-M35
[edit]Haplogroup E-M35 is a subclade of E-M215.
E-M281
[edit]Haplogroup E-M281 is a subclade of E-M215.
Phylogenetics
[edit]Phylogenetic history
[edit]Prior to 2002, there were in academic literature at least seven naming systems for the Y-Chromosome phylogenetic tree. This led to considerable confusion. In 2002, the major research groups came together and formed the Y-Chromosome Consortium (YCC). They published a joint paper that created a single new tree that all agreed to use. Later, a group of citizen scientists with an interest in population genetics and genetic genealogy formed a working group to create an amateur tree aiming at being above all timely. The table below brings together all of these works at the point of the landmark 2002 YCC Tree. This allows a researcher reviewing older published literature to quickly move between nomenclatures.
| YCC 2002/2008 (Shorthand) | (α) | (β) | (γ) | (δ) | (ε) | (ζ) | (η) | YCC 2002 (Longhand) | YCC 2005 (Longhand) | YCC 2008 (Longhand) | YCC 2010r (Longhand) | ISOGG 2006 | ISOGG 2007 | ISOGG 2008 | ISOGG 2009 | ISOGG 2010 | ISOGG 2011 | ISOGG 2012 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E-P29 | 21 | III | 3A | 13 | Eu3 | H2 | B | E* | E | E | E | E | E | E | E | E | E | E |
| E-M33 | 21 | III | 3A | 13 | Eu3 | H2 | B | E1* | E1 | E1a | E1a | E1 | E1 | E1a | E1a | E1a | E1a | E1a |
| E-M44 | 21 | III | 3A | 13 | Eu3 | H2 | B | E1a | E1a | E1a1 | E1a1 | E1a | E1a | E1a1 | E1a1 | E1a1 | E1a1 | E1a1 |
| E-M75 | 21 | III | 3A | 13 | Eu3 | H2 | B | E2a | E2 | E2 | E2 | E2 | E2 | E2 | E2 | E2 | E2 | E2 |
| E-M54 | 21 | III | 3A | 13 | Eu3 | H2 | B | E2b | E2b | E2b | E2b1 | - | - | - | - | - | - | - |
| E-P2 | 25 | III | 4 | 14 | Eu3 | H2 | B | E3* | E3 | E1b | E1b1 | E3 | E3 | E1b1 | E1b1 | E1b1 | E1b1 | E1b1 |
| E-M2 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a* | E3a | E1b1 | E1b1a | E3a | E3a | E1b1a | E1b1a | E1b1a | E1b1a1 | E1b1a1 |
| E-M58 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a1 | E3a1 | E1b1a1 | E1b1a1 | E3a1 | E3a1 | E1b1a1 | E1b1a1 | E1b1a1 | E1b1a1a1a | E1b1a1a1a |
| E-M116.2 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a2 | E3a2 | E1b1a2 | E1b1a2 | E3a2 | E3a2 | E1b1a2 | E1b1a2 | E1ba12 | removed | removed |
| E-M149 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a3 | E3a3 | E1b1a3 | E1b1a3 | E3a3 | E3a3 | E1b1a3 | E1b1a3 | E1b1a3 | E1b1a1a1c | E1b1a1a1c |
| E-M154 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a4 | E3a4 | E1b1a4 | E1b1a4 | E3a4 | E3a4 | E1b1a4 | E1b1a4 | E1b1a4 | E1b1a1a1g1c | E1b1a1a1g1c |
| E-M155 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a5 | E3a5 | E1b1a5 | E1b1a5 | E3a5 | E3a5 | E1b1a5 | E1b1a5 | E1b1a5 | E1b1a1a1d | E1b1a1a1d |
| E-M10 | 8 | III | 5 | 15 | Eu2 | H2 | B | E3a6 | E3a6 | E1b1a6 | E1b1a6 | E3a6 | E3a6 | E1b1a6 | E1b1a6 | E1b1a6 | E1b1a1a1e | E1b1a1a1e |
| E-M35 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b* | E3b | E1b1b1 | E1b1b1 | E3b1 | E3b1 | E1b1b1 | E1b1b1 | E1b1b1 | removed | removed |
| E-M78 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b1* | E3b1 | E1b1b1a | E1b1b1a1 | E3b1a | E3b1a | E1b1b1a | E1b1b1a | E1b1b1a | E1b1b1a1 | E1b1b1a1 |
| E-M148 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b1a | E3b1a | E1b1b1a3a | E1b1b1a1c1 | E3b1a3a | E3b1a3a | E1b1b1a3a | E1b1b1a3a | E1b1b1a3a | E1b1b1a1c1 | E1b1b1a1c1 |
| E-M81 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b2* | E3b2 | E1b1b1b | E1b1b1b1 | E3b1b | E3b1b | E1b1b1b | E1b1b1b | E1b1b1b | E1b1b1b1 | E1b1b1b1a |
| E-M107 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b2a | E3b2a | E1b1b1b1 | E1b1b1b1a | E3b1b1 | E3b1b1 | E1b1b1b1 | E1b1b1b1 | E1b1b1b1 | E1b1b1b1a | E1b1b1b1a1 |
| E-M165 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b2b | E3b2b | E1b1b1b2 | E1b1b1b1b1 | E3b1b2 | E3b1b2 | E1b1b1b2a | E1b1b1b2a | E1b1b1b2a | E1b1b1b2a | E1b1b1b1a2a |
| E-M123 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b3* | E3b3 | E1b1b1c | E1b1b1c | E3b1c | E3b1c | E1b1b1c | E1b1b1c | E1b1b1c | E1b1b1c | E1b1b1b2a |
| E-M34 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3b3a* | E3b3a | E1b1b1c1 | E1b1b1c1 | E3b1c1 | E3b1c1 | E1b1b1c1 | E1b1b1c1 | E1b1b1c1 | E1b1b1c1 | E1b1b1b2a1 |
| E-M136 | 25 | III | 4 | 14 | Eu4 | H2 | B | E3ba1 | E3b3a1 | E1b1b1c1a | E1b1b1c1a1 | E3b1c1a | E3b1c1a | E1b1b1c1a1 | E1b1b1c1a1 | E1b1b1c1a1 | E1b1b1c1a1 | E1b1b1b2a1a1 |
Research publications
[edit]The following research teams per their publications were represented in the creation of the YCC Tree.
Discussion
[edit]E-M215 and E1b1b1 are the currently accepted names found in the proposals of the Y Chromosome Consortium (YCC), for the clades defined by mutation M215 and M35 respectively, which can also be referred to as E-M215 and E-M35.[23] The nomenclature E3b (E-M215) and E3b1 (E-M35) respectively were the YCC defined names used to designate the same haplogroups in older literature with E-M35 branching as a separate subclade of E-M215 in 2004.[11] Prior to 2002 these haplogroups were not designated in a consistent way, and nor was their relationship to other related clades within haplogroup E and haplogroup DE. But in non-standard or older terminologies, E-M215 is for example approximately the same as "haplotype V", still used in publications such as Gérard et al. (2006).[24]
Phylogenetic trees
[edit]Cladogram with the main subclades:
| E1b1b (M215) |
| ||||||
The following phylogenetic tree is based on the YCC 2008 tree and subsequent published research as summarized by ISOGG. It includes all known subclades as of June 2015 (Trombetta et al. 2015)[25][23][24]
- E-M215 (E1b1b)
- E-M215*. Rare or non-existent.
- E-M35 (E1b1b1)
- E-V68 (E1b1b1a)
- E-V2009. Found in individuals in Sardinia and Morocco.
- E-M78 (E1b1b1a1). North Africa, Horn of Africa, West Asia, Sicily. (Formerly "E1b1b1a".)
- E-M78*
- E-V1477. Found in Tunisian Jews.
- E-V1083.
- E-V1083*. Found only in Eritrea (1.1%) and Sardinia (0.3%).
- E-V13
- E-V22
- E-V1129
- E-V12
- E-V12*
- E-V32
- E-V264
- E-V259. Found in North Cameroon.
- E-V65
- E-CTS194
- E-V12
- E-Z827 (E1b1b1b)[26]
- E-V257/L19 (L19, V257) – E1b1b1b1[26]
- E-PF2431
- E-M81 (M81)
- E-PF2546
- E-PF2546*
- E-CTS12227
- E-MZ11
- E-MZ12
- E-MZ11
- E-A929
- E-Z5009
- E-Z5009*
- E-Z5010
- E-Z5013
- E-Z5013*
- E-A1152
- E-A2227
- E-A428
- E-MZ16
- E-PF6794
- E-PF6794*
- E-PF6789
- E-MZ21
- E-MZ23
- E-MZ80
- E-A930
- E-Z2198/E-MZ46
- E-A601
- E-L351
- E-Z5009
- E-PF2546
- E-Z830 (Z830) – E1b1b1b2[26]
- E-M123 (M123)
- E-M34 (M34)
- E-M84 (M84)
- E-M136 (M136)
- E-M290 (M290)
- E-V23 (V23)
- E-L791 (L791,L792)
- E-M84 (M84)
- E-M34 (M34)
- E-V1515. E-V1515 and its subclades are mainly restricted to eastern Africa.
- E-V1515*
- E-V1486
- E-V1486*
- E-V2881
- E-V2881*
- E-V1792
- E-V92
- E-M293 (M293)
- E-M293*
- E-P72 (P72)
- E-V3065*
- E-V1700
- E-V42 (V42)
- E-V1785
- E-V1785*
- E-V6 (V6)
- E-M123 (M123)
- E-V257/L19 (L19, V257) – E1b1b1b1[26]
- E-V16/E-M281 (E1b1b2). Rare. Found in individuals in Ethiopia, Yemen and Saudi Arabia.
- E-V68 (E1b1b1a)
See also
[edit]Genetics
[edit]- African admixture in Europe
- genetic genealogy
- Haplogroup D
- Haplogroup DE
- Haplogroup
- Haplotype
- Human Y-chromosome DNA haplogroup
- Molecular phylogenetics
- Paragroup
- Subclade
- Y-chromosome haplogroups in populations of the world
- Y-DNA haplogroups by ethnic group
- Y-DNA haplogroups in populations of Sub-Saharan Africa
Y-DNA E subclades
[edit]- Haplogroup E-L485
- Haplogroup E-M123
- Haplogroup E-M180
- Haplogroup E-M215
- Haplogroup E-M33
- Haplogroup E-M521
- Haplogroup E-M75
- Haplogroup E-M96
- Haplogroup E-P147
- Haplogroup E-P177
- Haplogroup E-P2
- Haplogroup E-V12
- Haplogroup E-V13
- Haplogroup E-V22
- Haplogroup E-M2
- Haplogroup E-V65
- Haplogroup E-V68
- Haplogroup E-Z820
- Haplogroup E-Z827
Y-DNA backbone tree
[edit]Notes
[edit]- ^ Cruciani et al. (2004): "Several observations point to eastern Africa as the homeland for haplogroup E3b—that is, it had (1) the highest number of different E3b clades (table 1), (2) a high frequency of this haplogroup and a high microsatellite diversity, and, finally, (3) the exclusive presence of the undifferentiated E3b* paragroup." As mentioned above, "E3b" is the old name for E-M215. Semino et al. (2004): "This inference is further supported by the presence of additional Hg E lineal diversification and by the highest frequency of E-P2* and E-M35* in the same region. The distribution of E-P2* appears limited to eastern African peoples. The E-M35* lineage shows its highest frequency (19.2%) in the Ethiopian Oromo but with a wider distribution range than E-P2*." For E-M215 Cruciani et al. (2007) reduced their estimate to 22,400 from 25,600 in Cruciani et al. (2004), re-calibrating the same data.
- ^ "Paragroup E-M35 * and haplogroup J-12f2a* fit the criteria for major AJ founding lineages because they are widespread both in AJ populations and in Near Eastern populations, and occur at much lower frequencies in European non-Jewish populations." Behar et al. (2004)
References
[edit]- ^ a b Trombetta 2015.
- ^ Haber M, Jones AL, Connel BA, Asan, Arciero E, Huanming Y, Thomas MG, Xue Y, Tyler-Smith C (June 2019). "A Rare Deep-Rooting D0 African Y-chromosomal Haplogroup and its Implications for the Expansion of Modern Humans Out of Africa". Genetics. 212 (4): 1421–1428. doi:10.1534/genetics.119.302368. PMC 6707464. PMID 31196864.
- ^ a b Cruciani et al. (2007)
- ^ "E-M215 YTree".
- ^ Cruciani et al. (2004).
- ^ Schuenemann, Verena J.; et al. (2017). "Ancient Egyptian mummy genomes suggest an increase of Sub-Saharan African ancestry in post-Roman periods". Nature Communications. 8 15694. Bibcode:2017NatCo...815694S. doi:10.1038/ncomms15694. PMC 5459999. PMID 28556824.
- ^ Fregel; et al. (2018). "Ancient genomes from North Africa evidence prehistoric migrations to the Maghreb from both the Levant and Europe". bioRxiv 10.1101/191569.
- ^ Rodrı́guez-Varela; et al. (2017). "Genomic Analyses of Pre-European Conquest Human Remains from the Canary Islands Reveal Close Affinity to Modern North Africans". Current Biology. 27 (1–7): 3396–3402.e5. Bibcode:2017CBio...27E3396R. doi:10.1016/j.cub.2017.09.059. hdl:2164/13526. PMID 29107554.
- ^ Van De Loosdrecht, Marieke; Bouzouggar, Abdeljalil; Humphrey, Louise; Posth, Cosimo; Barton, Nick; Aximu-Petri, Ayinuer; Nickel, Birgit; Nagel, Sarah; Talbi, El Hassan; El Hajraoui, Mohammed Abdeljalil; Amzazi, Saaïd; Hublin, Jean-Jacques; Pääbo, Svante; Schiffels, Stephan; Meyer, Matthias; Haak, Wolfgang; Jeong, Choongwon; Krause, Johannes (4 May 2018). "Pleistocene North African genomes link Near Eastern and sub-Saharan African human populations". Science. 360 (6388): 548–552. Bibcode:2018Sci...360..548V. doi:10.1126/science.aar8380. PMID 29545507. S2CID 206666517.
- ^ Martiniano, Rui; De Sanctis, Bianca; Hallast, Pille; Durbin, Richard (February 2022). "Placing Ancient DNA Sequences into Reference Phylogenies". Molecular Biology and Evolution. 39 (2) msac017. doi:10.1093/molbev/msac017. PMC 8857924. PMID 35084493.
- ^ a b c Cruciani et al. (2004)
- ^ a b c d Semino et al. (2004)
- ^ Rosser et al. (2000)
- ^ Firasat et al. (2006)
- ^ Di Cristofaro, Julie; et al. (October 18, 2013). "Afghan Hindu Kush: Where Eurasian Sub-Continent Gene Flows Converge". PLOS ONE. 8 (10) e76748. Bibcode:2013PLoSO...876748D. doi:10.1371/journal.pone.0076748. ISSN 1932-6203. OCLC 5534533323. PMC 3799995. PMID 24204668. S2CID 16455960.
- ^ Ehret, Keita & Newman (2004); Keita & Boyce (2005); Keita (2008).
- ^ Behar et al. (2003)
- ^ Behar et al. (2004)
- ^ Shen et al. (2004)
- ^ Adams et al. (2008)
- ^ Nebel et al. (2001)
- ^ a b c Moran, Colin N.; et al. (2004). "Y chromosome haplogroups of elite Ethiopian endurance runners". Human Genetics. 115 (6): 492–7. doi:10.1007/s00439-004-1202-y. PMID 15503146. S2CID 13960753. Retrieved 6 February 2017.
- ^ a b Karafet et al. (2008)
- ^ a b Y Chromosome Consortium "YCC" (2002)
- ^ ISOGG (2011)
- ^ a b c ISOGG 2015
Bibliography
[edit]- Adams, Susan M; Bosch, Elena; Balaresque, Patricia L.; Ballereau, Stéphane J.; Lee, Andrew C.; Arroyo, Eduardo; López-Parra, Ana M.; Aler, Mercedes; et al. (2008), "The Genetic Legacy of Religious Diversity and Intolerance: Paternal Lineages of Christians, Jews, and Muslims in the Iberian Peninsula", The American Journal of Human Genetics, 83 (6): 725–36, doi:10.1016/j.ajhg.2008.11.007, PMC 2668061, PMID 19061982
- Alvarez; Santos, Cristina; Montiel, Rafael; Caeiro, Blazquez; Baali, Abdellatif; Dugoujon, Jean-Michel; Aluja, Maria Pilar (2009), "Y-chromosome variation in South Iberia: Insights into the North African contribution", American Journal of Human Biology, 21 (3): 407–409, doi:10.1002/ajhb.20888, PMID 19213004, S2CID 7041905
- Arredi, B; Poloni, E; Paracchini, S; Zerjal, T; Fathallah, D; Makrelouf, M; Pascali, V; Novelletto, A; Tylersmith, C (2004), "A Predominantly Neolithic Origin for Y-Chromosomal DNA Variation in North Africa", American Journal of Human Genetics, 75 (2): 338–345, doi:10.1086/423147, PMC 1216069, PMID 15202071
- Badro, Danielle A.; Douaihy, Bouchra; Haber, Marc; Youhanna, Sonia C.; Salloum, Angélique; Ghassibe-Sabbagh, Michella; Johnsrud, Brian; Khazen, Georges; Matisoo-Smith, Elizabeth; Soria-Hernanz, David F.; Wells, R. Spencer; Tyler-Smith, Chris; Platt, Daniel E.; Zalloua, Pierre A. (2013), "Y-Chromosome and mtDNA Genetics Reveal Significant Contrasts in Affinities of Modern Middle Eastern Populations with European and African Populations", PLOS ONE, 8 (1: e54616) e54616, Bibcode:2013PLoSO...854616B, doi:10.1371/journal.pone.0054616, PMC 3559847, PMID 23382925
- Battaglia, Vincenza; Fornarino, Simona; Al-Zahery, Nadia; Olivieri, Anna; Pala, Maria; Myres, Natalie M; King, Roy J; Rootsi, Siiri; et al. (2008), "Y-chromosomal evidence of the cultural diffusion of agriculture in southeast Europe", European Journal of Human Genetics, 17 (6): 820–830, doi:10.1038/ejhg.2008.249, PMC 2947100, PMID 19107149
- Behar, Doron M.; Thomas, Mark G.; Skorecki, Karl; Hammer, Michael F.; Bulygina, Ekaterina; Rosengarten, Dror; Jones, Abigail L.; Held, Karen; et al. (October 2003), "Multiple Origins of Ashkenazi Levites: Y Chromosome Evidence for Both Near Eastern and European Ancestries", Am. J. Hum. Genet., vol. 73, no. 4, pp. 768–779, doi:10.1086/378506, PMC 1180600, PMID 13680527. Also at http://www.ucl.ac.uk/tcga/tcgapdf/Behar-AJHG-03.pdf and https://web.archive.org/web/20090304100321/http://www.familytreedna.com/pdf/400971.pdf
- Behar; Garrigan; Kaplan; Mobasher; Rosengarten (November 2004), "Contrasting patterns of Y chromosome variation in Ashkenazi Jewish and host non-Jewish European populations" (PDF), Hum. Genet., vol. 114, no. 4, pp. 354–365, doi:10.1007/s00439-003-1073-7, PMID 14740294, S2CID 10310338, archived from the original (PDF) on 2011-11-10, retrieved 2009-09-11
- Beleza, Sandra; Gusmao, Leonor; Lopes, Alexandra; Alves, Cintia; Gomes, Iva; Giouzeli, Maria; Calafell, Francesc; Carracedo, Angel; Amorim, Antonio (2006), "Micro-Phylogeographic and Demographic History of Portuguese Male Lineages", Annals of Human Genetics, 70 (2): 181–194, doi:10.1111/j.1529-8817.2005.00221.x, PMID 16626329, S2CID 4652154
- Bird, Steven (2007), "Haplogroup E3b1a2 as a Possible Indicator of Settlement in Roman Britain by Soldiers of Balkan Origin", Journal of Genetic Genealogy, 3 (2), archived from the original on 2016-04-22, retrieved 2008-11-07
- Bortolini; Thomas, Mark G.; Chikhi, Lourdes; Aguilar, Juan A.; Castro-De-Guerra, Dinorah; Salzano, Francisco M.; Ruiz-Linares, Andres (2004), "Ribeiro's typology, genomes, and Spanish colonialism, as viewed from Gran Canaria and Colombia" (PDF), Genetics and Molecular Biology, 27 (1): 1–8, doi:10.1590/S1415-47572004000100001
- Bosch, Elena; Calafell, Francesc; Comas, David; Oefner, Peter J.; Underhill, Peter A.; Bertranpetit, Jaume (2001), "High-resolution analysis of human Y-chromosome variation shows a sharp discontinuity and limited gene flow between north-western Africa and the Iberian Peninsula", Am J Hum Genet, 68 (4): 1019–1029, doi:10.1086/319521, PMC 1275654, PMID 11254456
- Bosch, E.; Calafell, F.; Gonzalez-Neira, A.; Flaiz, C.; Mateu, E.; Scheil, H.-G.; Huckenbeck, W.; Efremovska, L.; et al. (2006), "Paternal and maternal lineages in the Balkans show a homogeneous landscape over linguistic barriers, except for the isolated Aromuns", Annals of Human Genetics, 70 (4): 459–487, doi:10.1111/j.1469-1809.2005.00251.x, PMID 16759179, S2CID 23156886, archived from the original on 2012-12-10
- Cadenas; Zhivotovsky, Lev A; Cavalli-Sforza, Luca L; Underhill, Peter A; Herrera, Rene J (2007), "Y-chromosome diversity characterizes the Gulf of Oman", European Journal of Human Genetics, 16 (3): 1–13, doi:10.1038/sj.ejhg.5201934, PMID 17928816
- Capelli, Cristian; Redhead, Nicola; Abernethy, Julia K.; Gratrix, Fiona; Wilson, James F.; Moen, Torolf; Hervig, Tor; Richards, Martin; et al. (2003), "A Y Chromosome Census of the British Isles", Current Biology, 13 (11): 979–84, Bibcode:2003CBio...13..979C, doi:10.1016/S0960-9822(03)00373-7, hdl:20.500.11820/8acb01f3-a7c1-45f5-89de-b796266d651e, PMID 12781138 also at [1]
- Caratti; Gino, S.; Torre, C.; Robino, C. (2009), "Subtyping of Y-chromosomal haplogroup E-M78 (E1b1b1a) by SNP assay and its forensic application", International Journal of Legal Medicine, 123 (4): 357–360, doi:10.1007/s00414-009-0350-y, PMID 19430804, S2CID 5657112
- Capelli, Cristian; Onofri, Valerio; Brisighelli, Francesca; Boschi, Ilaria; Scarnicci, Francesca; Masullo, Mara; Ferri, Gianmarco; Tofanelli, Sergio; et al. (2009), "Moors and Saracens in Europe: estimating the medieval North African male legacy in southern Europe", European Journal of Human Genetics, 17 (6): 848–852, doi:10.1038/ejhg.2008.258, PMC 2947089, PMID 19156170
- Cinnioğlu, Cengiz; King, Roy; Kivisild, Toomas; Kalfoglu, Ersi; Atasoy, Sevil; Cavalleri, Gianpiero L.; Lillie, Anita S.; Roseman, Charles C.; et al. (2004), "Excavating Y-chromosome haplotype strata in Anatolia", Hum Genet, 114 (2): 127–48, doi:10.1007/s00439-003-1031-4, PMID 14586639, S2CID 10763736
- Contu, Daniela; Morelli, Daniela; Santoni, Federico; Foster, Jamie W.; Francalacci, Paolo; Cucca, Francesco (2008), "Y-Chromosome Based Evidence for Pre-Neolithic Origin of the Genetically Homogeneous but Diverse Sardinian Population: Inference for Association Scans", PLOS ONE, 3 (1) e1430, Bibcode:2008PLoSO...3.1430C, doi:10.1371/journal.pone.0001430, PMC 2174525, PMID 18183308
- Cruciani, Fulvio; Santolamazza, Piero; Shen, Peidong; MacAulay, Vincent; Moral, Pedro; Olckers, Antonel; Modiano, David; Holmes, Susan (2002), "A Back Migration from Asia to Sub-Saharan Africa Is Supported by High-Resolution Analysis of Human Y-Chromosome Haplotypes", American Journal of Human Genetics, 70 (5): 1197–1214, doi:10.1086/340257, PMC 447595, PMID 11910562
- Cruciani; La Fratta; Santolamazza; Sellitto (May 2004), "Phylogeographic Analysis of Haplogroup E3b (E-M215) Y Chromosomes Reveals Multiple Migratory Events Within and Out Of Africa" (PDF), American Journal of Human Genetics, 74 (5): 1014–1022, doi:10.1086/386294, PMC 1181964, PMID 15042509, archived from the original (PDF) on 2008-06-26, retrieved 2008-05-17
- Cruciani; La Fratta; Torroni; Underhill; Scozzari (2006), "Molecular Dissection of the Y Chromosome Haplogroup E-M78 (E3b1a): A Posteriori Evaluation of a Microsatellite-Network-Based Approach Through Six New Biallelic Markers", Human Mutation, 27 (8): 831–2, doi:10.1002/humu.9445, PMID 16835895, S2CID 26886757
- Cruciani, F.; La Fratta, R.; Trombetta, B.; Santolamazza, P.; Sellitto, D.; Colomb, E. B.; Dugoujon, J.-M.; Crivellaro, F.; et al. (2007), "Tracing Past Human Male Movements in Northern/Eastern Africa and Western Eurasia: New Clues from Y-Chromosomal Haplogroups E-M78 and J-M12", Molecular Biology and Evolution, 24 (6): 1300–1311, doi:10.1093/molbev/msm049, PMID 17351267[dead link] Also see Supplementary Data.
- Di Gaetano; Cerutti, Francesca; Crobu, Carlo; Robino (2009), "Differential Greek and northern African migrations to Sicily are supported by genetic evidence from the Y chromosome", European Journal of Human Genetics, 17 (1): 91–99, doi:10.1038/ejhg.2008.120, PMC 2985948, PMID 18685561
- Dugoujon; Coudray; Torroni; Cruciani; Scozzari; Moral; Louali; Kossmann (2009), d'Errico; Hombert (eds.), "The Berber and the Berbers: Genetic and linguistic diversities" (PDF), Becoming Eloquent Advances in the Emergence of Language, Human Cognition, and Modern Cultures: 123–146, doi:10.1075/z.152.05ch4, ISBN 978-90-272-3269-4, archived from the original (PDF) on 2016-10-19, retrieved 2013-01-26
- Ehret, C.; Keita, SO; Newman, P (2004), "The Origins of Afroasiatic", Science, 306 (5702): 1680, doi:10.1126/science.306.5702.1680c, PMID 15576591, S2CID 8057990
- El-Sibai, Mirvat; Platt, Daniel E.; Haber, Marc; Xue, Yali; Youhanna, Sonia C.; Wells, R. Spencer; Izaabel, Hassan; Sanyoura, May F.; et al. (2009), "Geographical Structure of the Y-chromosomal Genetic Landscape of the Levant: A coastal-inland contrast", Annals of Human Genetics, 73 (6): 568–581, doi:10.1111/j.1469-1809.2009.00538.x, PMC 3312577, PMID 19686289
- Firasat; Khaliq, Shagufta; Mohyuddin, Aisha; Papaioannou, Myrto; Tyler-Smith, Chris; Underhill, Peter A; Ayub, Qasim (2006), "Y-chromosomal evidence for a limited Greek contribution to the Pathan population of Pakistan", European Journal of Human Genetics, 15 (1): 121–126, doi:10.1038/sj.ejhg.5201726, PMC 2588664, PMID 17047675
- Flores, Carlos; Maca-Meyer, Nicole; González, Ana M; Oefner, Peter J; Shen, Peidong; Pérez, Jose A; Rojas, Antonio; Larruga, Jose M; Underhill, Peter A (2004), "Reduced genetic structure of the Iberian peninsula revealed by Y-chromosome analysis: implications for population demography", European Journal of Human Genetics, 12 (10): 855–863, doi:10.1038/sj.ejhg.5201225, PMID 15280900, S2CID 16765118
- Flores; Maca-Meyer, Nicole; Larruga, Jose M.; Cabrera, Vicente M.; Karadsheh, Naif; Gonzalez, Ana M. (2005), "Isolates in a corridor of migrations: a high-resolution analysis of Y-chromosome variation in Jordan", J Hum Genet, 50 (9): 435–441, doi:10.1007/s10038-005-0274-4, PMID 16142507
- Francalacci, P.; Morelli, L.; Underhill, P.A.; Lillie, A.S.; Passarino, G.; Useli, A.; Madeddu, R.; Paoli, G.; et al. (2003), "Peopling of Three Mediterranean Islands (Corsica, Sardinia, and Sicily) Inferred by Y-Chromosome Biallelic Variability", American Journal of Physical Anthropology, 121 (3): 270–279, doi:10.1002/ajpa.10265, PMID 12772214
- Fregel, Rosa; Gomes, Verónica; Gusmão, Leonor; González, Ana M; Cabrera, Vicente M; Amorim, António; Larruga, Jose M (2009), "Demographic history of Canary Islands male gene-pool: replacement of native lineages by European", BMC Evolutionary Biology, 9 (1): 181, Bibcode:2009BMCEE...9..181F, doi:10.1186/1471-2148-9-181, PMC 2728732, PMID 19650893
- Gérard; Berriche, S; Aouizérate, A; Diéterlen, F; Lucotte, G (2006), "North African Berber and Arab influences in the western Mediterranean revealed by Y-chromosome DNA haplotypes", Human Biology, 78 (3): 307–316, doi:10.1353/hub.2006.0045, PMID 17216803, S2CID 13347549
- Gonçalves, R; Freitas, A; Branco, M; Rosa, A; Fernandes, AT; Zhivotovsky, LA; Underhill, PA; Kivisild, T; Brehm, A (2005), "Y-chromosome Lineages from Portugal, Madeira and Açores Record Elements of Sephardim and Berber Ancestry", Annals of Human Genetics, 69 (Pt 4): 443–454, doi:10.1111/j.1529-8817.2005.00161.x, hdl:10400.13/3018, PMID 15996172, S2CID 3229760[dead link]
- Hammer (2003), "Human population structure and its effects on sampling Y chromosome sequence variation", Genetics, 164 (4): 1495–1509, doi:10.1093/genetics/164.4.1495, PMC 1462677, PMID 12930755
- Hassan, Hisham Y.; Underhill, Peter A.; Cavalli-Sforza, Luca L.; Ibrahim, Muntaser E. (2008), "Y-Chromosome Variation Among Sudanese: Restricted Gene Flow, Concordance With Language, Geography, and History" (PDF), American Journal of Physical Anthropology, 137 (3): 316–23, doi:10.1002/ajpa.20876, PMID 18618658, archived from the original (PDF) on 2009-03-04
- Henn, B. M.; Gignoux, C.; Lin, Alice A; Oefner, Peter J.; Shen, P.; Scozzari, R.; Cruciani, F.; Tishkoff, S. A.; Mountain, J. L.; Underhill, P. A. (2008), "Y-chromosomal evidence of a pastoralist migration through Tanzania to southern Africa", PNAS, 105 (31): 10693–8, Bibcode:2008PNAS..10510693H, doi:10.1073/pnas.0801184105, PMC 2504844, PMID 18678889. See comment on Dienekes blog, comment on the Spitoon blog and public release Archived 2021-05-08 at the Wayback Machine.
- Y-DNA Haplogroup E and its Subclades – 2011, International Society of Genetic Genealogy (ISOGG), 2011[clarification needed]
- Jobling, M.A.; Tyler-Smith, C. (2000), "New uses for new haplotypes the human Y chromosome, disease and selection", Trends Genet., 16 (8): 356–362, doi:10.1016/S0168-9525(00)02057-6, PMID 10904265
- Karafet, T. M.; Mendez, F. L.; Meilerman, M. B.; Underhill, P. A.; Zegura, S. L.; Hammer, M. F. (May 2008), "New Binary Polymorphisms Reshape and Increase Resolution of the Human Y-Chromosomal Haplogroup Tree", Genome Research, 18 (5): 830–8, doi:10.1101/gr.7172008, PMC 2336805, PMID 18385274. Published online April 2, 2008. See also Supplementary Material.
- Keita, Shomarka (2008), "Geography, selected Afro-Asiatic families, and Y chromosome lineage variation", In Hot Pursuit of Language in Prehistory: Essays in the Four Fields of Anthropology: In Honor of Harold Crane Fleming, John Benjamins, ISBN 978-90-272-3252-6
- Keita, S. O. Y.; Boyce, A. J. (Anthony J.) (2005), "Genetics, Egypt, and History: Interpreting Geographical Patterns of Y Chromosome Variation", History in Africa, 32: 221–246, doi:10.1353/hia.2005.0013, S2CID 163020672
- King, R. J.; Özcan, S. S.; Carter, T.; Kalfoğlu, E.; Atasoy, S.; Triantaphyllidis, C.; Kouvatsi, A.; Lin, A. A.; et al. (2008), "Differential Y-chromosome Anatolian Influences on the Greek and Cretan Neolithic" (PDF), Annals of Human Genetics, 72 (2): 205–214, doi:10.1111/j.1469-1809.2007.00414.x, PMID 18269686, S2CID 22406638, archived from the original (PDF) on 2009-03-05
- King; Underhill (2002), "Congruent distribution of Neolithic painted pottery and ceramic figurines with Y-chromosome lineages", Antiquity, 76 (293): 707–14, doi:10.1017/S0003598X00091158, S2CID 160359661
- Kujanova; Pereira; Fernandes; Pereira; Cerný (2009), "Near Eastern Neolithic Genetic Input in a Small Oasis of the Egyptian Western Desert", American Journal of Physical Anthropology, 140 (2): 336–346, doi:10.1002/ajpa.21078, PMID 19425100
- Lacan, Marie; Keyser, Christine; Ricaut, François-Xavier; Brucato, Nicolas; Tarrús, Josep; Bosch, Angel; Guilaine, Jean; Crubézy, Eric; Ludes, Bertrand (2011), "Ancient DNA suggests the leading role played by men in the Neolithic dissemination", PNAS, 108 (45): 18255–9, Bibcode:2011PNAS..10818255L, doi:10.1073/pnas.1113061108, PMC 3215063, PMID 22042855
- Lancaster, Andrew (2009), "Y Haplogroups, Archaeological Cultures and Language Families: a Review of the Multidisciplinary Comparisons using the case of E-M35" (PDF), Journal of Genetic Genealogy, 5 (1), archived from the original (PDF) on 2016-05-06, retrieved 2009-06-26
- Luis, J; Rowold, D; Regueiro, M; Caeiro, B; Cinnioglu, C; Roseman, C; Underhill, P; Cavallisforza, L; Herrera, R (2004), "The Levant versus the Horn of Africa: Evidence for Bidirectional Corridors of Human Migrations" (PDF), American Journal of Human Genetics, 74 (3): 532–544, doi:10.1086/382286, PMC 1182266, PMID 14973781, archived from the original (PDF) on 2012-02-16. (Also see Errata)
- Maca-Meyer N, Sánchez-Velasco P, Flores C, Larruga JM, González AM, Oterino A, Leyva-Cobián F, et al. (2003), "Y Chromosome and Mitochondrial DNA Characterization of Pasiegos, a Human Isolate from Cantabria (Spain)", Annals of Human Genetics, 67 (Pt 4): 329–339, CiteSeerX 10.1.1.584.4253, doi:10.1046/j.1469-1809.2003.00045.x, PMID 12914567, S2CID 40355653.
- Martinez, Laisel; Underhill, Peter A; Zhivotovsky, Lev A; Gayden, Tenzin; Moschonas, Nicholas K; Chow, Cheryl-Emiliane T; Conti, Simon; Mamolini, Elisabetta; Cavalli-Sforza, L Luca; Herrera, Rene (April 1, 2007), "Paleolithic Y-haplogroup heritage predominates in a Cretan highland plateau", European Journal of Human Genetics, 15 (4): 485–493, doi:10.1038/sj.ejhg.5201769, ISSN 1018-4813, PMID 17264870
- Mendizabal, Isabel; Sandoval, Karla; Berniell-Lee, Gemma; Calafell, Francesc; Salas, Antonio; Martinez-Fuentes, Antonio; Comas, David (2008), "Genetic origin, admixture, and asymmetry in maternal and paternal human lineages in Cuba", BMC Evol. Biol., 8 (1): 213, Bibcode:2008BMCEE...8..213M, doi:10.1186/1471-2148-8-213, PMC 2492877, PMID 18644108
- Nebel, Almut; Filon, D; Brinkmann, B; Majumder, P; Faerman, M; Oppenheim, A (2001), "The Y Chromosome Pool of Jews as Part of the Genetic Landscape of the Middle East", American Journal of Human Genetics, 69 (5): 1095–1112, doi:10.1086/324070, PMC 1274378, PMID 11573163
- Onofri, Valerio; Alessandrini, Federica; Turchi, Chiara; Pesaresi, Mauro; Buscemi, Loredana; Tagliabracci, Adriano (2006), "Development of multiplex PCRs for evolutionary and forensic applications of 37 human Y chromosome SNPs" (PDF), Forensic Science International, 157 (1): 23–35, doi:10.1016/j.forsciint.2005.03.014, PMID 15896936[permanent dead link]
- Paracchini; Pearce, CL; Kolonel, LN; Altshuler, D; Henderson, BE; Tyler-Smith, C (2003), "A Y chromosomal influence on prostate cancer risk: the multi-ethnic cohort study", J Med Genet, 40 (11): 815–819, doi:10.1136/jmg.40.11.815, PMC 1735314, PMID 14627670
- Pelotti; Ceccardi, S; Lugaresi, F; Trane, R; Falconi, M; Bini, C; Willuweit, S; Roewer, L (2007), "Microgeographic genetic variation of Y chromosome in a population sample of Ravenna's area in the Emilia-Romagna region (North of Italy)", Forensic Science International: Genetics Supplement Series, 1 (1): 242–243, doi:10.1016/j.fsigss.2007.10.025
- Pereira, Luísa; Černý, Viktor; Cerezo, María; Silva, Nuno M; Hájek, Martin; Vašíková, Alžběta; Kujanová, Martina; Brdička, Radim; Salas, Antonio (2010), "Linking the sub-Saharan and West Eurasian gene pools: maternal and paternal heritage of the Tuareg nomads from the African Sahel" (PDF), European Journal of Human Genetics, 18 (8): 915–923, doi:10.1038/ejhg.2010.21, PMC 2987384, PMID 20234393, archived from the original (PDF) on 2013-05-28
- Peričic, M.; Lauc, LB; Klarić, IM; Rootsi, S; Janićijevic, B; Rudan, I; Terzić, R; Colak, I; et al. (2005), "High-resolution phylogenetic analysis of southeastern Europe traces major episodes of paternal gene flow among Slavic populations", Mol. Biol. Evol., vol. 22, no. 10, pp. 1964–75, doi:10.1093/molbev/msi185, PMID 15944443, archived from the original on June 30, 2012.
- Pontikos D. "Phylogeographic refinement of haplogroup E" http://dienekes.blogspot.ru/2015/07/phylogeographic-refinement-of.html
- Ramos-Luisa, E.; Blanco-Verea, A.; Brión, M.; Van Huffel, V.; Carracedo, A.; Sánchez-Diz, P. (2009), "Phylogeography of French male lineages (unpublished data 23rd International ISFG Congress)", Forensic Science International, 2: 439–441, doi:10.1016/j.fsigss.2009.09.026, S2CID 85134429[dead link]
- Regueiro, M.; Cadenas, A.M.; Gayden, T.; Underhill, P.A.; Herrera, R.J. (2006), "Iran: Tricontinental Nexus for Y-Chromosome Driven Migration", Hum Hered, 61 (3): 132–143, doi:10.1159/000093774, PMID 16770078, S2CID 7017701
- Robino, C.; Crobu, F.; Gaetano, C.; Bekada, A.; Benhamamouch, S.; Cerutti, N.; Piazza, A.; Inturri, S.; Torre, C. (2008), "Analysis of Y-chromosomal SNP haplogroups and STR haplotypes in an Algerian population sample", Journal International Journal of Legal Medicine, 122 (3): 251–5, doi:10.1007/s00414-007-0203-5, PMID 17909833, S2CID 11556974
- Rosa, Alexandra; Ornelas, Carolina; Jobling, Mark A; Brehm, António; Villems, Richard (2007), "Y-chromosomal diversity in the population of Guinea-Bissau: a multiethnic perspective", BMC Evolutionary Biology, 7 (1): 124, Bibcode:2007BMCEE...7..124R, doi:10.1186/1471-2148-7-124, PMC 1976131, PMID 17662131
- Rosser, Z; Zerjal, T; Hurles, M; Adojaan, M; Alavantic, D; Amorim, A; Amos, W; Armenteros, M; et al. (2000), "Y-Chromosomal Diversity in Europe Is Clinal and Influenced Primarily by Geography, Rather than by Language", American Journal of Human Genetics, 67 (6): 1526–1543, doi:10.1086/316890, PMC 1287948, PMID 11078479, archived from the original on 2008-05-06
- Sanchez, Juan J; Hallenberg, Charlotte; Børsting, Claus; Hernandez, Alexis; Gorlin, RJ (2005), "High frequencies of Y chromosome lineages characterized by E3b1, DYS19-11, DYS392-12 in Somali males", European Journal of Human Genetics, 13 (7): 856–866, doi:10.1038/sj.ejhg.5201390, PMID 15756297. Published online 9 March 2005
- Scozzari, Rosaria; Cruciani, F; Pangrazio, A; Santolamazza, P; Vona, G; Moral, P; Latini, V; Varesi, L; et al. (2001), "Human Y-Chromosome Variation in the Western Mediterranean Area: Implications for the Peopling of the Region" (PDF), Human Immunology, 62 (9): 871–884, CiteSeerX 10.1.1.408.4857, doi:10.1016/S0198-8859(01)00286-5, PMID 11543889, archived from the original (PDF) on 2012-12-17, retrieved 2009-03-01
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA; Arbuzova, S; Beckman, LE; De Benedictis, G; Francalacci, P; et al. (2000), "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective" (PDF), Science, vol. 290, no. 5494, pp. 1155–59, Bibcode:2000Sci...290.1155S, doi:10.1126/science.290.5494.1155, PMID 11073453, archived from the original (PDF) on 2003-11-25.
- Semino; Santachiara-Benerecetti, A. Silvana; Falaschi, Francesco; Cavalli-Sforza, L. Luca; Underhill, Peter A. (2002), "Ethiopians and Khoisan share the deepest clades of the human Y-chromosome phylogeny" (PDF), Am J Hum Genet, vol. 70, no. 1, pp. 265–268, doi:10.1086/338306, PMC 384897, PMID 11719903, archived from the original (PDF) on 2006-03-15
- Semino, Ornella; Magri, Chiara; Benuzzi, Giorgia; Lin, Alice A.; Al-Zahery, Nadia; Battaglia, Vincenza; MacCioni, Liliana; Triantaphyllidis, Costas; et al. (2004), "Origin, Diffusion, and Differentiation of Y-Chromosome Haplogroups E and J: Inferences on the Neolithization of Europe and Later Migratory Events in the Mediterranean Area", American Journal of Human Genetics, vol. 74, no. 5, pp. 1023–1034, doi:10.1086/386295, PMC 1181965, PMID 15069642
- Shen, Peidong; Lavi, Tal; Kivisild, Toomas; Chou, Vivian; Sengun, Deniz; Gefel, Dov; Shpirer, Issac; Woolf, Eilon; et al. (2004), "Reconstruction of Patrilineages and Matrilineages of Samaritans and Other Israeli Populations From Y-Chromosome and Mitochondrial DNA Sequence Variation" (PDF), Human Mutation, vol. 24, no. 3, pp. 248–60, doi:10.1002/humu.20077, PMID 15300852, S2CID 1571356, archived from the original (PDF) on 2009-03-05, retrieved 2008-11-07
- Silva; Carvalho, Elizeu; Costa, Guilherme; Tavares, Lígia; Amorim, António; Gusmão, Leonor (2006), "Y-chromosome genetic variation in Rio de Janeiro population", American Journal of Human Biology, 18 (6): 829–837, doi:10.1002/ajhb.20567, PMID 17039481, S2CID 23778828, archived from the original on 2012-10-18
- Shlush; Behar, Doron M.; Yudkovsky, Guennady; Templeton, Alan; Hadid, Yarin; Basis, Fuad; Hammer, Michael; Itzkovitz, Shalev; Skorecki, Karl (2008), Gemmell, Neil John (ed.), "The Druze: A Population Genetic Refugium of the Near East", PLOS ONE, 3 (5) e2105, Bibcode:2008PLoSO...3.2105S, doi:10.1371/journal.pone.0002105, PMC 2324201, PMID 18461126
- Solé-Morata, Neus; García-Fernández, Carla; Urasin, Vadim; Bekada, Asmahan; Fadhlaoui-Zid, Karima; Zalloua, Pierre; Comas, David; Calafell, Francesc (21 November 2017), "Whole Y-chromosome sequences reveal an extremely recent origin of the most common North African paternal lineage E-M183 (M81)", Scientific Reports, 7 (1): 15941, Bibcode:2017NatSR...715941S, doi:10.1038/s41598-017-16271-y, ISSN 2045-2322, PMC 5698413, PMID 29162904
- Sykes, Bryan (2006), Blood of the Isles: Exploring the Genetic Roots of Our Tribal History, Bantam, ISBN 978-0-593-05652-3
- Tishkoff; Gonder; Henn; Mortensen; Knight (2007), "History of Click-Speaking Populations of Africa Inferred from mtDNA and Y Chromosome Genetic Variation", Mol Biol Evol, 24 (10): 2180–2195, doi:10.1093/molbev/msm155, PMID 17656633, archived from the original on 2011-10-10
- Thomas; Stumpf, M. P.H; Harke, H. (2006), "Evidence for an apartheid-like social structure in early Anglo-Saxon England" (PDF), Proceedings of the Royal Society B, 273 (1601): 2651–2657, doi:10.1098/rspb.2006.3627, PMC 1635457, PMID 17002951, archived from the original (PDF) on 2009-03-05
- Trombetta, Beniamino; Cruciani, Fulvio; Sellitto, Daniele; Scozzari, Rosaria (2011), MacAulay, Vincent (ed.), "A New Topology of the Human Y Chromosome Haplogroup E1b1 (E-P2) Revealed through the Use of Newly Characterized Binary Polymorphisms", PLOS ONE, 6 (1) e16073, Bibcode:2011PLoSO...616073T, doi:10.1371/journal.pone.0016073, PMC 3017091, PMID 21253605
- Trombetta, Beniamino; et al. (July 2015). "Phylogeographic Refinement and Large Scale Genotyping of Human Y Chromosome Haplogroup E Provide New Insights into the Dispersal of Early Pastoralists in the African Continent". Genome Biology and Evolution. 7 (7): 1940–1950. doi:10.1093/gbe/evv118. ISSN 1759-6653. OCLC 5854174538. PMC 4524485. PMID 26108492. S2CID 16352575.
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li; Passarino, Giuseppe; Yang, Wei H.; Kauffman, Erin; Bonné-Tamir, Batsheva; et al. (2000), "Y chromosome sequence variation and the history of human populations", Nat Genet, vol. 26, no. 3, pp. 358–361, doi:10.1038/81685, PMID 11062480, S2CID 12893406
- Underhill; Passarino, G.; Lin, A. A.; Shen, P.; Mirazon Lahr, M.; Foley, R. A.; Oefner, P. J.; Cavalli-Sforza, L. L. (2001), "The phylogeography of Y chromosome binary haplotypes and the origins of modern human populations", Ann Hum Genet, vol. 65, no. Pt 1, pp. 43–62, doi:10.1046/j.1469-1809.2001.6510043.x, PMID 11415522, S2CID 9441236
- Underhill (2002), "Inference of Neolithic Population Histories using Y-chromosome Haplotypes", in Bellwood, Peter; Renfrew, A. Colin (eds.), Examining the farming/language dispersal hypothesis, Cambridge: McDonald Institute for Archaeological Research, ISBN 978-1-902937-20-5
- Underhill, Peter A.; Kivisild, Toomas (2007), "Use of Y Chromosome and Mitochondrial DNA Population Structure in Tracing Human Migrations", Annu. Rev. Genet., 41: 539–64, doi:10.1146/annurev.genet.41.110306.130407, PMID 18076332
- Weale, M. E.; Weiss, D. A.; Jager, R. F.; Bradman, N.; Thomas, M. G. (2002), "Y Chromosome Evidence for Anglo-Saxon Mass Migration" (PDF), Mol. Biol. Evol., vol. 19, no. 7, pp. 1008–1021, doi:10.1093/oxfordjournals.molbev.a004160, PMID 12082121, archived from the original (PDF) on 2005-10-29.
- Weale; Shah, T; Jones, AL; Greenhalgh, J; Wilson, JF; Nymadawa, P; Zeitlin, D; Connell, BA; et al. (September 1, 2003), "Rare Deep-Rooting Y Chromosome Lineages in Humans: Lessons for Phylogeography", Genetics, vol. 165, no. 1, pp. 229–234, doi:10.1093/genetics/165.1.229, PMC 1462739, PMID 14504230
- Wood; Stover; Ehret; Destro-Bisol (2005), "Contrasting patterns of Y chromosome and mtDNA variation in Africa: evidence for sex-biased demographic processes", European Journal of Human Genetics, 13 (7): 867–876, doi:10.1038/sj.ejhg.5201408, PMID 15856073
- Y Chromosome Consortium "YCC" (2002), "A Nomenclature System for the Tree of Human Y-Chromosomal Binary Haplogroups", Genome Research, vol. 12, no. 2, pp. 339–348, doi:10.1101/gr.217602, PMC 155271, PMID 11827954
- Zalloua, Pierre A.; Xue, Yali; Khalife, Jade; Makhoul, Nadine; Debiane, Labib; Platt, Daniel E.; Royyuru, Ajay K.; Herrera, Rene J.; et al. (2008), "Y-Chromosomal Diversity in Lebanon Is Structured by Recent Historical Events", American Journal of Human Genetics, 82 (4): 873–882, doi:10.1016/j.ajhg.2008.01.020, PMC 2427286, PMID 18374297
- Zalloua, Pierre A.; Platt, Daniel E.; El Sibai, Mirvat; Khalife, Jade; Makhoul, Nadine; Haber, Marc; Xue, Yali; Izaabel, Hassan; et al. (2008), "Identifying Genetic Traces of Historical Expansions: Phoenician Footprints in the Mediterranean", The American Journal of Human Genetics, 83 (5): 633–642, doi:10.1016/j.ajhg.2008.10.012, PMC 2668035, PMID 18976729
- Zerjal (1999), The use of Y-chromosomal DNA variation to investigate population history; in Papiha SS, Deka R, Chakraborty R (eds): Genomic diversity: applications in human population genetics, Kluwer Academic/Plenum Publishers, pp. 91–101
- Zhao; Khan, Faisal; Borkar, Minal; Herrera, Rene; Agrawal, Suraksha (2009), "Presence of three different paternal lineages among North Indians: A study of 560 Y chromosomes", Annals of Human Biology, 36 (1): 1–14, doi:10.1080/03014460802558522, PMC 2755252, PMID 19058044
Sources for conversion tables
[edit]- Capelli, Cristian; Wilson, James F.; Richards, Martin; Stumpf, Michael P.H.; et al. (February 2001). "A Predominantly Indigenous Paternal Heritage for the Austronesian-Speaking Peoples of Insular Southeast Asia and Oceania". The American Journal of Human Genetics. 68 (2): 432–443. doi:10.1086/318205. PMC 1235276. PMID 11170891.
- Hammer, Michael F.; Karafet, Tatiana M.; Redd, Alan J.; Jarjanazi, Hamdi; et al. (1 July 2001). "Hierarchical Patterns of Global Human Y-Chromosome Diversity". Molecular Biology and Evolution. 18 (7): 1189–1203. doi:10.1093/oxfordjournals.molbev.a003906. PMID 11420360.
- Kaladjieva, Luba; Calafell, Francesc; Jobling, Mark A; Angelicheva, Dora; et al. (February 2001). "Patterns of inter- and intra-group genetic diversity in the Vlax Roma as revealed by Y chromosome and mitochondrial DNA lineages". European Journal of Human Genetics. 9 (2): 97–104. doi:10.1038/sj.ejhg.5200597. PMID 11313742. S2CID 21432405.
- Karafet, Tatiana; Xu, Liping; Du, Ruofu; Wang, William; et al. (September 2001). "Paternal Population History of East Asia: Sources, Patterns, and Microevolutionary Processes". The American Journal of Human Genetics. 69 (3): 615–628. doi:10.1086/323299. PMC 1235490. PMID 11481588.
- Semino, O.; Passarino, G; Oefner, PJ; Lin, AA; et al. (2000), "The Genetic Legacy of Paleolithic Homo sapiens sapiens in Extant Europeans: A Y Chromosome Perspective", Science, 290 (5494): 1155–9, Bibcode:2000Sci...290.1155S, doi:10.1126/science.290.5494.1155, PMID 11073453
- Su, Bing; Xiao, Junhua; Underhill, Peter; Deka, Ranjan; et al. (December 1999). "Y-Chromosome Evidence for a Northward Migration of Modern Humans into Eastern Asia during the Last Ice Age". The American Journal of Human Genetics. 65 (6): 1718–1724. doi:10.1086/302680. PMC 1288383. PMID 10577926.
- Underhill, Peter A.; Shen, Peidong; Lin, Alice A.; Jin, Li; et al. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–361. doi:10.1038/81685. PMID 11062480. S2CID 12893406.
Haplogroup E-M215
View on GrokipediaNomenclature and Overview
Definition and Characteristics
Haplogroup E-M215, also known as E1b1b and formerly designated as E3b, is a major human Y-chromosome DNA haplogroup defined by the single-nucleotide polymorphism (SNP) M215.[6][7] This haplogroup represents a primary branch of the larger Haplogroup E (defined by mutations such as M96), specifically under the E1b1 (E-P2) subclade.[6][8] E-M215 diverges into two basal subclades: E-M35 and the rarer E-M281.[6] Within the broader human Y-DNA phylogenetic tree, Haplogroup E-M215 falls under the macrohaplogroup DE (defined by M145), which encompasses the major African and Asian Y-chromosome lineages. As a Y-chromosome haplogroup, E-M215 is characterized by its basis in SNPs located on the non-recombining region of the Y chromosome, which undergoes no genetic recombination during meiosis and is transmitted exclusively from father to son in a patrilineal manner.[7] This inheritance pattern enables E-M215 and other Y-DNA haplogroups to serve as reliable markers for tracing deep paternal ancestry and reconstructing historical male-mediated population migrations across continents.[6][7]Historical Classification
The historical classification of Haplogroup E-M215 traces back to the early development of Y-chromosome nomenclature in the mid-1990s. In their seminal review, Jobling and Tyler-Smith (1995) proposed a letter-based system for major Y-haplogroups, designating the lineage now known as E-M215 as part of haplogroup E3, based on initial binary polymorphisms like the 12f2 marker associated with African populations.[9][10] This nomenclature reflected the limited resolution of early genetic markers, which grouped diverse African Y-lineages under broad categories without distinguishing finer subclades.[10] Subsequent studies refined this classification amid growing phylogenetic detail. Hammer et al. (1998) provided one of the first identifications of key markers within haplogroup E through nested cladistic analysis, highlighting its deep African roots but still using the E3 designation for the broader clade encompassing what would later be split. By 2001, Underhill et al. introduced the subclade E3b to specifically denote the branch defined by the M215 mutation, distinguishing it from E3a (now E1b1a or E-M2), which predominates in sub-Saharan Africa; this separation addressed early confusions in distribution patterns, as E3b was linked more to North Africa and the Near East. Cruciani et al. (2002) further solidified this by confirming M215 as the defining mutation for E3b through high-resolution haplotype analysis, resolving ambiguities in marker stability and supporting its role in Eurasian back-migrations.[11] The nomenclature underwent a major shift in the late 2000s as phylogenetic trees became more hierarchical. Following the Y Chromosome Consortium's 2002 standardization, which emphasized mutation-based naming, the International Society of Genetic Genealogy (ISOGG) updated its tree in 2008 to reclassify E3b as E1b1b, placing it under the E1b1 paragroup to better reflect its position relative to E1b1a (E-M2) and other basal E branches; this change incorporated refined SNP data and avoided the alphabetical limitations of earlier systems.[12] The ISOGG nomenclature for E-M215 as E1b1b has remained consistent in subsequent updates through 2020, with focus shifting to finer subclade resolutions.[13] Early controversies, such as overlaps in geographic signals between E3a and E3b leading to debates on single versus multiple African origins, were largely clarified by these updates, emphasizing distinct evolutionary trajectories within the broader haplogroup E.[10]Origins and Age Estimates
Proposed Geographic Origin
Haplogroup E-M215, a major subclade of the African macrohaplogroup E-P2, is widely accepted to have originated in East Africa, with the highest probability placing its emergence in the Horn of Africa or Ethiopia based on phylogeographic analyses of Y-chromosome diversity.[14] This consensus arises from Bayesian modeling of SNP data across thousands of samples, which assigns a posterior probability of 0.97 to an eastern African origin for the clade's diversity.[14] Supporting this, the greatest basal diversity of E-M215 lineages, including rare paragroups like E-M215*, is observed in Ethiopian and Eritrean populations, where frequencies of deep-rooting branches exceed those in North Africa or the Levant.[1] These patterns link E-M215 to the broader diversification of modern human Y-chromosomes in Africa during early population expansions across the continent.[3] Earlier studies, such as Semino et al. (2004), debated possible North African or Levantine origins for certain E-M215 subclades like E-M78 or E-M123, inferring back-migrations from Eurasia based on limited microsatellite data and distributions in Mediterranean populations.[15] However, subsequent high-resolution SNP genotyping and diversity metrics have refuted these hypotheses, demonstrating that the root of E-M215 lies in East Africa rather than a secondary Eurasian center, with North African clades like E-M81 representing later regional adaptations.[1][14] Cruciani et al. (2004) further solidified the East African homeland through network analyses showing elevated variance and ancient lineages in sub-Saharan contexts.[1] The evolutionary context of E-M215 aligns with Pleistocene environmental shifts in East Africa, including fluctuating climates that facilitated human adaptations to diverse habitats from savannas to highlands.[3] This period coincides with the haplogroup's emergence around the late Pleistocene, potentially tying it to early pastoralist innovations, as evidenced by correlations between E-M215 subclades and Afro-Asiatic-speaking groups practicing animal herding in the Horn of Africa.[3] Such associations suggest that carriers of E-M215 contributed to cultural dispersals linked to Neolithic transitions in the region.[14]Temporal Estimates and Methods
The time to the most recent common ancestor (TMRCA) for Haplogroup E-M215 has been estimated at approximately 34,600 years before present (ybp) based on single nucleotide polymorphism (SNP) counting calibrated against ancient DNA and modern genomes in the YFull YTree database.[2] Similarly, FamilyTreeDNA's Discover tool, drawing from extensive Big Y testing data, places the TMRCA around 35,000 ybp (33,000 BCE, with a 95% confidence interval of 37,934–28,704 BCE).[16] An earlier estimate from Cruciani et al. (2004), using microsatellite (STR) variance in 509 E-M215 chromosomes, yielded a TMRCA of 25,600 ybp (95% CI: 24,300–27,400 ybp).[1] These age estimates are derived primarily through Y-chromosome short tandem repeat (STR) mutation rates, which measure genetic distance via the average squared difference (ASD) or rho statistic from the haplogroup founder haplotype, and SNP-based methods that count substitutions along phylogenetic branches calibrated to a molecular clock.[17] Whole-genome sequencing enhances precision by integrating SNP data with STRs, while Bayesian coalescent models, such as those implemented in BATWING software, account for population size fluctuations and generate posterior distributions of coalescence times using Markov chain Monte Carlo sampling on SNP and STR datasets.[18] These approaches prioritize evolutionary mutation rates over pedigree-based ones to better reflect long-term Y-chromosome dynamics. Recent refinements have incorporated ancient DNA for clock calibration, reducing uncertainties by anchoring phylogenies to dated archaeological samples; for instance, Lazaridis et al. (2016) provided genomic data from ancient Near Eastern individuals, including E-M215 carriers, that helps validate Y-haplogroup divergence times against historical timelines.[19] Discrepancies persist between SNP-based clocks, which tend to yield older estimates due to their stability, and STR-based clocks, which can inflate ages from higher apparent mutation rates influenced by multiple hits.[20] Key factors affecting these estimates include back-mutation rates in STR loci, where reversals to ancestral states can underestimate coalescence times if not modeled stepwise, and sampling biases in African populations, where underrepresentation of diverse lineages may skew variance calculations toward younger apparent ages.[21][22]Phylogenetic Structure
Basal Branches
Haplogroup E-M215 is primarily divided into two basal branches: E-M35, which accounts for nearly all (>99%) of E-M215 lineages, and E-M281, an extremely rare subclade comprising less than 1% of the total.[23] These branches represent the immediate phylogenetic structure downstream of the E-M215 defining mutation, with E-M35 dominating modern distributions and E-M281 showing more restricted occurrence.[4] E-M35 is defined by the M35 (also known as P64) single nucleotide polymorphism and further subdivides into major lineages such as E-V68 (including E-M78) and E-Z827 (including E-M123 and E-M81), among others.[23][4] This branch is strongly associated with historical expansions originating in East Africa, followed by significant dispersals into North Africa and across the Mediterranean region, likely linked to Neolithic migrations and the spread of Afro-Asiatic languages.[23][1] In contrast, E-M281 (also known as E-V16) is characterized by the M281 mutation and observed at low frequencies primarily in the Arabian Peninsula and the Horn of Africa. As of 2025, YFull estimates the TMRCA of E-V16 at 4,100 years before present, with documented carriers mainly from Saudi Arabia, Yemen, Qatar, and Kuwait.[24][25] This lineage shows evidence of broader dispersal beyond East Africa, including across the Red Sea region.[26][4] The paragroup E-M215*, consisting of lineages that do not carry the defining mutations of E-M35 or E-M281, is exceedingly rare and may represent unsampled or extinct basal branches within the haplogroup.[23][1] Modern surveys have identified few or no such chromosomes, underscoring the completeness of resolution for the primary divisions.[23]Phylogenetic Trees and Updates
The phylogenetic structure of Haplogroup E-M215 is commonly represented in standardized trees maintained by genetic genealogy organizations. In the International Society of Genetic Genealogy (ISOGG) Y-DNA haplogroup tree (version 2023), E-M215 is positioned as a primary subclade under the broader E-P2 lineage, with its two main basal branches being E-M35 (further ramifying into E-V68, E-M78, and E-M81, among others) and the less common E-M281 (also known as E-V16). This tree structure relies on a curated set of single nucleotide polymorphisms (SNPs) to delineate evolutionary relationships, emphasizing E-M35 as the dominant branch leading to widespread subclades such as E-V13 (under E-M78). Recent updates have refined this phylogeny through high-throughput sequencing data. The YFull YTree database (as of early 2025) estimates E-M215 formation at approximately 41,200 years before present (ybp) and its time to most recent common ancestor (TMRCA) at 34,600 ybp, incorporating over 90 defining SNPs and expanding subclades with novel variants such as E-Y338012 and E-FTC43334 added in late 2024.[27] Similarly, FamilyTreeDNA's Big Y-700 testing in 2024 confirmed an ancient split upstream of E-M215, between E-P177 and E-P2 at around 49,000 ybp, based on analysis of a Yemenite lineage sample that revealed 8 upstream and 113 downstream SNPs, including the private variant P75.[28] The historical development of the E-M215 phylogeny has progressed from initial binary marker-based classifications in the early 2000s, which used a limited set of SNPs like M215 to define broad haplogroups, to a highly resolved framework today enabled by next-generation sequencing (NGS). Seminal studies, such as the 2008 analysis of new binary polymorphisms, expanded the human Y-chromosome tree from 153 to over 300 haplogroups by integrating hundreds of SNPs, with specific refinements to E-M215 topology emerging in 2010 through short tandem repeat (STR) and SNP correlations that clarified subclade equivalences like V43 and V95 under E-M2.[4] By 2013, unbiased SNP discovery via NGS further deepened the African structure of E lineages, adding chronological depth and resolving basal dichotomies within E-M215.[29] Interactive tools for exploring these trees include the ISOGG online Y-DNA haplogroup browser, which provides downloadable SNP lists and branch diagrams; YFull's dynamic YTree viewer, updated quarterly with user-submitted NGS data; and FamilyTreeDNA's Discover platform, which integrates Big Y results to display personalized placements and ancient correlations without embedding static images.[13][27][16]Major Subclades
E-M35
Haplogroup E-M35 is defined by the single nucleotide polymorphism (SNP) mutation M35 on the Y-chromosome and represents the primary and most widespread subclade of the parent haplogroup E-M215.[14] This lineage encompasses the majority of E-M215 diversity, accounting for its extensive distribution across Africa, Europe, and the Middle East.[14] Age estimates for the origin of E-M35, derived from Bayesian phylogenetic analysis of high-coverage sequencing data, place its emergence at approximately 25,000 years before present (ybp), with a 95% confidence interval of 20,000–30,000 ybp.[14] This timing positions E-M35 as a relatively ancient branch within E-M215, predating significant post-glacial human expansions.[30] The phylogenetic structure of E-M35 features two main basal branches: E-V68 and E-Z827.[14] Under E-V68 lies E-M78, which is prevalent in North Africa and Europe and further branches into E-V13, a subclade associated with Balkan populations.[14] E-Z827 includes E-M81, dominant among Berber groups in North Africa, and E-M123, found primarily in the Middle East.[14] These downstream subclades highlight E-M35's role in shaping regional genetic profiles through subsequent diversifications.[14] Historically, E-M35 has been associated with Neolithic expansions, including migrations that facilitated the spread of agriculture and pastoralism from North Africa into Europe and surrounding regions.[30] Genetic diversity within E-M35 is notably high in Berber populations of North Africa, where multiple subclades such as E-M81 and refined paragroups contribute to elevated haplotype variation.[14] Recent studies, including large-scale genotyping of over 5,000 samples, have refined the internal structure of E-M35 by resolving polytomies and identifying new clades, such as those under E-V1515, thereby enhancing understanding of its dispersal patterns among pastoralist groups.[14]E-M281
E-M281 is a rare subclade of human Y-chromosome haplogroup E-M215, defined by the single nucleotide polymorphism (SNP) M281, a G-to-A transition at position 280 in the Y-chromosome sequence.[31] This lineage was first identified in a study of Ethiopian populations, where it was observed in two individuals from the Oromo ethnic group.[31] Subsequent analyses, including a large-scale survey of over 3,400 males across multiple continents, tested for the M281 marker but detected no carriers, underscoring its extreme rarity outside specific African contexts.[1] The distribution of E-M281 is highly restricted, occurring almost exclusively in Ethiopia among Cushitic-speaking Oromo populations.[31] Its low frequency—limited to just a few documented cases—suggests either a recent population expansion from a small founding group or a genetic bottleneck that reduced diversity within the clade.[1] Age estimates suggest E-M281 diverged from E-M35 approximately 34,000 years ago.[28] This pattern aligns with its association with indigenous East African groups, potentially reflecting localized demographic histories rather than widespread dispersal. As one of the basal branches of E-M215 alongside E-M35, E-M281 represents a distinct lineage that likely remained confined to Africa, contrasting with the broader migratory patterns observed in E-M35.[31] Its presence in Ethiopian samples highlights the deep-rooted diversity of haplogroup E in the Horn of Africa, offering insights into pre-Neolithic population structures in the region.[1]Modern Distribution
Geographic Spread
Haplogroup E-M215 is most prominently distributed in North Africa, where it reaches peak frequencies among Berber-speaking populations, such as 80% in the Mozabite Berbers of Algeria (based on a small sample of N=20).[1] In the Horn of Africa, it maintains high prevalence, with representative frequencies of approximately 50% among Somalis and 20-30% in Ethiopian groups like the Oromo and Amhara.[1] These regions represent the core of its modern range, reflecting deep-rooted associations with indigenous African populations. The haplogroup's secondary spread extends to the Middle East, particularly the Levant, where frequencies range from 10-20% in populations such as Palestinians.[1] In Southern Europe, including areas like Greece and Italy, it occurs at moderate levels of 10-20%, often linked to historical gene flow across the Mediterranean, with peaks up to 20-32% in the Balkans.[1] Traces of E-M215 also appear in the Americas, primarily among populations of African descent, resulting from colonial-era transatlantic migrations.[32] This distribution pattern arises from post-Last Glacial Maximum expansions, with migrations facilitating its dispersal via Mediterranean routes from eastern Africa toward Eurasia.[1] Subclades such as E-M35 have notably driven much of this broader dissemination.[3] Genetic mapping studies reveal a characteristic cline, with frequencies gradually declining from high levels in Africa to lower incidences in Europe, underscoring these historical movements.[1]Population Frequencies and Associations
Haplogroup E-M215 displays marked variation in frequency across global populations, with the highest concentrations observed in North Africa and the Horn of Africa, reflecting its deep roots in these regions. In Algerian Mozabites, a Berber-speaking group, E-M215 reaches 80% (N=20), predominantly through the E-M81 subclade, underscoring its association with indigenous North African lineages.[1] Similar elevated levels are reported in Libya at approximately 42%, where it contributes significantly to the paternal gene pool amid diverse historical migrations.[33] In Egypt, the frequency stands at about 20%, primarily driven by subclades like E-M78, linking local populations to broader Northeast African patterns.[1] Among Jewish populations, the subclade E-M34 (under E-M35) occurs in Cohanim, suggesting ties to ancient Near Eastern dispersals and the Jewish diaspora.[34] E-M215 shows strong correlations with Afro-Asiatic language speakers, including Berber, Semitic, and Cushitic groups, where it often exceeds 30-50% and aligns with pastoralist expansions.[3] In contrast, frequencies drop outside Africa, but remain 10-20% in southern European populations due to historical gene flow.[1] Recent research highlights ethnic-specific variations, such as higher E-M215 prevalence in Cushitic-speaking groups (e.g., Oromo at ~32%) compared to Nilotic ones (often <10%), reflecting differential Neolithic influences in East Africa.[1] These patterns emphasize E-M215's role as a marker of Afro-Asiatic demographic history without direct ties to specific phenotypic traits.| Population Group | E-M215 Frequency (%) | Key Notes |
|---|---|---|
| Algerian Mozabites | 80 (N=20) | Predominantly E-M81; Berber isolate.[1] |
| Libyans | ~42 | Reflects North African diversity; includes E-M81 (33.7%) and E-M78 (8%).[33] |
| Egyptians | 20 | Mainly E-M78 in northern samples.[1] |
| Ethiopians (general) | 40 | Higher in Cushitic speakers like Oromo.[1] |
| Southern Europeans | 10-20 | Elevated in Greece, Italy, and Balkans.[1] |
| Jewish Cohanim | Present (E-M34 subclade) | Linked to diaspora migrations.[34] |