Recent from talks
Nothing was collected or created yet.
Nucleic acid structure
View on Wikipedia


Nucleic acid structure refers to the structure of nucleic acids such as DNA and RNA. Chemically speaking, DNA and RNA are very similar. Nucleic acid structure is often divided into four different levels: primary, secondary, tertiary, and quaternary.
Primary structure
[edit]
Primary structure consists of a linear sequence of nucleotides that are linked together by phosphodiester bonds. It is this linear sequence of nucleotides that make up the primary structure of DNA or RNA. Nucleotides consist of 3 components:
- Nitrogenous base
- 5-carbon sugar which is called deoxyribose (found in DNA) and ribose (found in RNA).
- One or more phosphate groups.[1]
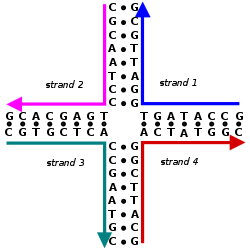
The nitrogen bases adenine and guanine are purine in structure and form a glycosidic bond between their 9 nitrogen and the 1' -OH group of the deoxyribose. Cytosine, thymine, and uracil are pyrimidines, hence the glycosidic bonds form between their 1 nitrogen and the 1' -OH of the deoxyribose. For both the purine and pyrimidine bases, the phosphate group forms a bond with the deoxyribose sugar through an ester bond between one of its negatively charged oxygen groups and the 5' -OH of the sugar.[2] The polarity in DNA and RNA is derived from the oxygen and nitrogen atoms in the backbone. Nucleic acids are formed when nucleotides come together through phosphodiester linkages between the 5' and 3' carbon atoms.[3] A nucleic acid sequence is the order of nucleotides within a DNA (GACT) or RNA (GACU) molecule that is determined by a series of letters. Sequences are presented from the 5' to 3' end and determine the covalent structure of the entire molecule. Sequences can be complementary to another sequence in that the base on each position is complementary as well as in the reverse order. An example of a complementary sequence to AGCT is TCGA. DNA is double-stranded containing both a sense strand and an antisense strand. Therefore, the complementary sequence will be to the sense strand.[4]
Complexes with alkali metal ions
[edit]There are three potential metal binding groups on nucleic acids: phosphate, sugar, and base moieties. Solid-state structure of complexes with alkali metal ions have been reviewed.[6]
Secondary structure
[edit]DNA
[edit]Secondary structure is the set of interactions between bases, i.e., which parts of strands are bound to each other. In DNA double helix, the two strands of DNA are held together by hydrogen bonds. The nucleotides on one strand base pairs with the nucleotide on the other strand. The secondary structure is responsible for the shape that the nucleic acid assumes. The bases in the DNA are classified as purines and pyrimidines. The purines are adenine and guanine. Purines consist of a double ring structure, a six-membered and a five-membered ring containing nitrogen. The pyrimidines are cytosine and thymine. It has a single ring structure, a six-membered ring containing nitrogen. A purine base always pairs with a pyrimidine base (guanine (G) pairs with cytosine (C) and adenine (A) pairs with thymine (T) or uracil (U)). DNA's secondary structure is predominantly determined by base-pairing of the two polynucleotide strands wrapped around each other to form a double helix. Although the two strands are aligned by hydrogen bonds in base pairs, the stronger forces holding the two strands together are stacking interactions between the bases. These stacking interactions are stabilized by Van der Waals forces and hydrophobic interactions, and show a large amount of local structural variability.[7] There are also two grooves in the double helix, which are called major groove and minor groove based on their relative size.
RNA
[edit]
The secondary structure of RNA consists of a single polynucleotide. Base pairing in RNA occurs when RNA folds between complementarity regions. Both single- and double-stranded regions are often found in RNA molecules.
The four basic elements in the secondary structure of RNA are:
- Helices
- Bulges
- Loops
- Junctions
The antiparallel strands form a helical shape.[3] Bulges and internal loops are formed by separation of the double helical tract on either one strand (bulge) or on both strands (internal loops) by unpaired nucleotides.
Stem-loop or hairpin loop is the most common element of RNA secondary structure.[8] Stem-loop is formed when the RNA chains fold back on themselves to form a double helical tract called the 'stem', the unpaired nucleotides forms single stranded region called the 'loop'. A tetraloop is a four-base pairs hairpin RNA structure. There are three common families of tetraloop in ribosomal RNA: UNCG, GNRA, and CUUG (N is one of the four nucleotides and R is a purine). UNCG is the most stable tetraloop.[9]
Pseudoknot is an RNA secondary structure first identified in turnip yellow mosaic virus.[10] It is minimally composed of two helical segments connected by single-stranded regions or loops. H-type fold pseudoknots are best characterized. In H-type fold, nucleotides in the hairpin-loop pair with the bases outside the hairpin stem forming second stem and loop. This causes formation of pseudoknots with two stems and two loops.[11] Pseudoknots are functional elements in RNA structure having diverse function and found in most classes of RNA.
Secondary structure of RNA can be predicted by experimental data on the secondary structure elements, helices, loops, and bulges. DotKnot-PW method is used for comparative pseudoknots prediction. The main points in the DotKnot-PW method is scoring the similarities found in stems, secondary elements and H-type pseudoknots.[12]
Tertiary structure
[edit]

Tertiary structure refers to the locations of the atoms in three-dimensional space, taking into consideration geometrical and steric constraints. It is a higher order than the secondary structure, in which large-scale folding in a linear polymer occurs and the entire chain is folded into a specific 3-dimensional shape. There are 4 areas in which the structural forms of DNA can differ.
- Handedness – right or left
- Length of the helix turn
- Number of base pairs per turn
- Difference in size between the major and minor grooves[3]
The tertiary arrangement of DNA's double helix in space includes B-DNA, A-DNA, and Z-DNA. Triple-stranded DNA structures have been demonstrated in repetitive polypurine:polypyrimidine Microsatellite sequences and Satellite DNA.
B-DNA is the most common form of DNA in vivo and is a more narrow, elongated helix than A-DNA. Its wide major groove makes it more accessible to proteins. On the other hand, it has a narrow minor groove. B-DNA's favored conformations occur at high water concentrations; the hydration of the minor groove appears to favor B-DNA. B-DNA base pairs are nearly perpendicular to the helix axis. The sugar pucker which determines the shape of the a-helix, whether the helix will exist in the A-form or in the B-form, occurs at the C2'-endo.[13]
A-DNA, is a form of the DNA duplex observed under dehydrating conditions. It is shorter and wider than B-DNA. RNA adopts this double helical form, and RNA-DNA duplexes are mostly A-form, but B-form RNA-DNA duplexes have been observed.[14] In localized single strand dinucleotide contexts, RNA can also adopt the B-form without pairing to DNA.[15] A-DNA has a deep, narrow major groove which does not make it easily accessible to proteins. On the other hand, its wide, shallow minor groove makes it accessible to proteins but with lower information content than the major groove. Its favored conformation is at low water concentrations. A-DNAs base pairs are tilted relative to the helix axis, and are displaced from the axis. The sugar pucker occurs at the C3'-endo and in RNA 2'-OH inhibits C2'-endo conformation.[13] Long considered little more than a laboratory artifice, A-DNA is now known to have several biological functions.
Z-DNA is a relatively rare left-handed double-helix. Given the proper sequence and superhelical tension, it can be formed in vivo but its function is unclear. It has a more narrow, more elongated helix than A or B. Z-DNA's major groove is not really a groove, and it has a narrow minor groove. The most favored conformation occurs when there are high salt concentrations. There are some base substitutions but they require an alternating purine-pyrimidine sequence. The N2-amino of G H-bonds to 5' PO, which explains the slow exchange of protons and the need for the G purine. Z-DNA base pairs are nearly perpendicular to the helix axis. Z-DNA does not contain single base-pairs but rather a GpC repeat with P-P distances varying for GpC and CpG. On the GpC stack there is good base overlap, whereas on the CpG stack there is less overlap. Z-DNA's zigzag backbone is due to the C sugar conformation compensating for G glycosidic bond conformation. The conformation of G is syn, C2'-endo; for C it is anti, C3'-endo.[13]
A linear DNA molecule having free ends can rotate, to adjust to changes of various dynamic processes in the cell, by changing how many times the two chains of its double helix twist around each other. Some DNA molecules are circular and are topologically constrained. More recently circular RNA was described as well to be a natural pervasive class of nucleic acids, expressed in many organisms (see CircRNA).
A covalently closed, circular DNA (also known as cccDNA) is topologically constrained as the number of times the chains coiled around one other cannot change. This cccDNA can be supercoiled, which is the tertiary structure of DNA. Supercoiling is characterized by the linking number, twist and writhe. The linking number (Lk) for circular DNA is defined as the number of times one strand would have to pass through the other strand to completely separate the two strands. The linking number for circular DNA can only be changed by breaking of a covalent bond in one of the two strands. Always an integer, the linking number of a cccDNA is the sum of two components: twists (Tw) and writhes (Wr).[16]
Twists are the number of times the two strands of DNA are twisted around each other. Writhes are number of times the DNA helix crosses over itself. DNA in cells is negatively supercoiled and has the tendency to unwind. Hence the separation of strands is easier in negatively supercoiled DNA than in relaxed DNA. The two components of supercoiled DNA are solenoid and plectonemic. The plectonemic supercoil is found in prokaryotes, while the solenoidal supercoiling is mostly seen in eukaryotes.
Quaternary structure
[edit]The quaternary structure of nucleic acids is similar to that of protein quaternary structure. Although some of the concepts are not exactly the same, the quaternary structure refers to a higher-level of organization of nucleic acids. Moreover, it refers to interactions of the nucleic acids with other molecules. The most commonly seen form of higher-level organization of nucleic acids is seen in the form of chromatin which leads to its interactions with the small proteins histones. Also, the quaternary structure refers to the interactions between separate RNA units in the ribosome or spliceosome.[17]
See also
[edit]- Biomolecular structure
- Crosslinking of DNA
- DNA nanotechnology
- DNA supercoil
- Gene structure
- Non-helical models of DNA structure
- Nucleic acid design
- Nucleic acid double helix
- Nucleic acid structure determination (experimental)
- Nucleic acid structure prediction (computational)
- Nucleic acid thermodynamics
- Protein structure
- Satellite DNA
- Triple-stranded DNA
References
[edit]- ^ Krieger M, Scott MP, Matsudaira PT, Lodish HF, Darnell JE, Lawrence Z, Kaiser C, Berk A (2004). "Section 4.1: Structure of Nucleic Acids". Molecular cell biology. New York: W.H. Freeman and CO. ISBN 978-0-7167-4366-8.
- ^ "Structure of Nucleic Acids". SparkNotes.
- ^ a b c Anthony-Cahill SJ, Mathews CK, van Holde KE, Appling DR (2012). Biochemistry (4th ed.). Englewood Cliffs, N.J: Prentice Hall. ISBN 978-0-13-800464-4.
- ^ Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Wlater P (2002). Molecular Biology of the Cell (4th ed.). New York NY: Garland Science. ISBN 978-0-8153-3218-3.
- ^ Mao C (December 2004). "The emergence of complexity: lessons from DNA". PLOS Biology. 2 (12): e431. doi:10.1371/journal.pbio.0020431. PMC 535573. PMID 15597116.
- ^ Katsuyuki, Aoki; Kazutaka, Murayama; Hu, Ning-Hai (2016). "Solid State Structures of Alkali Metal Ion Complexes Formed by Low-Molecular-Weight Ligands of Biological Relevance". In Astrid, Sigel; Helmut, Sigel; Roland K.O., Sigel (eds.). The Alkali Metal Ions: Their Role for Life. Metal Ions in Life Sciences. Vol. 16. Springer. pp. 43–66. doi:10.1007/978-3-319-21756-7_3. ISBN 978-3-319-21755-0. PMID 26860299.
- ^ Sedova A, Banavali NK (2017). "Geometric Patterns for Neighboring Bases Near the Stacked State in Nucleic Acid Strands". Biochemistry. 56 (10): 1426–1443. doi:10.1021/acs.biochem.6b01101. PMID 28187685.
- ^ Tinoco I, Bustamante C (October 1999). "How RNA folds". Journal of Molecular Biology. 293 (2): 271–81. doi:10.1006/jmbi.1999.3001. PMID 10550208.
- ^ Hollyfield JG, Besharse JC, Rayborn ME (December 1976). "The effect of light on the quantity of phagosomes in the pigment epithelium". Experimental Eye Research. 23 (6): 623–35. doi:10.1016/0014-4835(76)90221-9. PMID 1087245.
- ^ Rietveld K, Van Poelgeest R, Pleij CW, Van Boom JH, Bosch L (March 1982). "The tRNA-like structure at the 3' terminus of turnip yellow mosaic virus RNA. Differences and similarities with canonical tRNA". Nucleic Acids Research. 10 (6): 1929–46. doi:10.1093/nar/10.6.1929. PMC 320581. PMID 7079175.
- ^ Staple DW, Butcher SE (June 2005). "Pseudoknots: RNA structures with diverse functions". PLOS Biology. 3 (6): e213. doi:10.1371/journal.pbio.0030213. PMC 1149493. PMID 15941360.
- ^ Sperschneider J, Datta A, Wise MJ (December 2012). "Predicting pseudoknotted structures across two RNA sequences". Bioinformatics. 28 (23): 3058–65. doi:10.1093/bioinformatics/bts575. PMC 3516145. PMID 23044552.
- ^ a b c Dickerson RE, Drew HR, Conner BN, Wing RM, Fratini AV, Kopka ML (April 1982). "The anatomy of A-, B-, and Z-DNA". Science. 216 (4545): 475–85. Bibcode:1982Sci...216..475D. doi:10.1126/science.7071593. PMID 7071593.
- ^ Chen X; Ramakrishnan B; Sundaralingam M (1995). "Crystal structures of B-form DNA-RNA chimers complexed with distamycin". Nature Structural Biology. 2 (9): 733–735. doi:10.1038/nsb0995-733. PMID 7552741. S2CID 6886088.
- ^ Sedova A, Banavali NK (2016). "RNA approaches the B-form in stacked single strand dinucleotide contexts". Biopolymers. 105 (2): 65–82. doi:10.1002/bip.22750. PMID 26443416. S2CID 35949700.
- ^ Mirkin SM (2001). "DNA Topology: Fundamentals". eLS. doi:10.1038/npg.els.0001038. ISBN 978-0470016176.
{{cite book}}:|journal=ignored (help) - ^ "Structural Biochemistry/Nucleic Acid/DNA/DNA structure". Retrieved 11 December 2012.
Nucleic acid structure
View on GrokipediaBasic components
Nucleobases
Nucleobases are the aromatic nitrogenous compounds that form the core informational components of nucleic acids, distinguishing DNA from RNA through specific variants. These molecules attach to sugar-phosphate backbones via glycosidic bonds and enable sequence-specific recognition through hydrogen bonding patterns. The primary nucleobases are classified into purines and pyrimidines based on their ring structures, with ionization properties governed by pKa values that ensure neutrality at physiological pH. Purine nucleobases, adenine (A) and guanine (G), possess a bicyclic structure comprising a six-membered pyrimidine ring fused to a five-membered imidazole ring, providing extended aromaticity and rigidity. Adenine, chemically known as 6-aminopurine, features an amino group at position 6, while guanine, or 2-amino-6-oxopurine, includes an amino group at position 2 and a keto group at position 6. Both exist predominantly in their amino-keto tautomeric forms under neutral conditions, with rare enol or imino tautomers occurring transiently and potentially influencing base pairing fidelity. The pKa values for these purines—approximately 4.15 for adenine (protonation at N1) and 9.2 for guanine (deprotonation at N1-H)—position them as neutral species at pH 7, minimizing electrostatic repulsion in nucleic acid polymers.[4][5] Pyrimidine nucleobases, cytosine (C), thymine (T) in DNA, and uracil (U) in RNA, are characterized by a single six-membered heterocyclic ring with nitrogens at positions 1 and 3. Cytosine is 4-amino-2-oxopyrimidine, bearing an amino group at position 4 and a keto group at position 2; thymine is 5-methyl-2,4-dioxopyrimidine, with keto groups at positions 2 and 4 and a methyl substituent at position 5; uracil is 2,4-dioxopyrimidine, identical to thymine but lacking the 5-methyl group. This methyl group in thymine enhances hydrophobic interactions and stability in DNA compared to uracil in RNA, contributing to distinct evolutionary roles in genetic storage versus expression. Their pKa values, around 4.5 for cytosine (protonation at N3), 9.7 for thymine, and 9.5 for uracil (deprotonation at N3-H), similarly favor neutral forms at physiological pH.[6][4] The hydrogen bonding capabilities of these nucleobases dictate complementary pairing: adenine forms two hydrogen bonds with thymine or uracil via its N1 acceptor and N6-H donor pairing with the O4 and N3-H of T/U, respectively, while guanine forms three hydrogen bonds with cytosine through its O6 and N1-H donors and N2-H donor interacting with cytosine's N3, O2, and N4-H. These patterns, illustrated in the base pair diagrams below, ensure specificity and stability in nucleic acid duplexes. Adenine-Thymine (or Uracil) Pair (2 H-bonds):- N6-H (A) ... O4 (T/U)
- N1 (A) ... H-N3 (T/U)
- O6 (G) ... H-N4 (C)
- N1-H (G) ... N3 (C)
- N2-H (G) ... O2 (C)
Sugars and phosphate backbone
The sugar-phosphate backbone forms the structural scaffold of nucleic acids, consisting of alternating deoxyribose (in DNA) or ribose (in RNA) sugars linked to phosphate groups via phosphodiester bonds. In DNA, the sugar is 2-deoxy-D-ribose, a pentose lacking a hydroxyl group at the 2' carbon position, while RNA incorporates D-ribose, which includes this 2'-OH group. Both sugars exist predominantly in the β-D-furanose (five-membered ring) conformation, with the anomeric carbon (C1') linked to the nucleobase via a β-glycosidic bond, ensuring a consistent orientation in the polymer chain. This furanose form provides rigidity to the backbone while allowing rotational flexibility around the C4'-C5' and C3'-O3' bonds. The absence of the 2'-OH in deoxyribose enhances DNA's chemical stability by preventing intramolecular nucleophilic attacks that could disrupt the phosphodiester linkages, making DNA suitable for long-term genetic storage. In contrast, ribose's 2'-OH group increases RNA's susceptibility to hydrolysis but also imparts greater conformational flexibility, particularly in single-stranded regions, enabling diverse folding motifs essential for RNA's functional roles. This structural difference influences overall polymer dynamics: RNA tends toward A-form helices with a wider major groove due to the 2'-OH's steric and hydrogen-bonding effects, while DNA favors the more elongated B-form. Phosphodiester bonds are formed through condensation polymerization, where the 5'-phosphate group of one nucleotide reacts with the 3'-OH group of another, eliminating a water molecule and creating a covalent linkage between the 5' carbon and 3' carbon across the phosphate. This unidirectional 5' to 3' polarity defines the orientation of nucleic acid chains, with synthesis and replication processes proceeding exclusively in this direction. The resulting backbone is polyanionic, as each phosphate carries a negative charge at physiological pH (pKa ≈ 1-2), necessitating counterions for charge neutralization and structural integrity. Monovalent cations such as Na⁺ and K⁺ serve as primary counterions, coordinating directly with the negatively charged phosphate oxygens to screen electrostatic repulsion and stabilize the helix. Molecular dynamics studies reveal that Na⁺ interacts more strongly with phosphate groups due to its higher charge density, often forming closer ion-phosphate contacts, whereas K⁺ prefers interactions with nucleobase atoms in the grooves, influencing hydration patterns and minor conformational adjustments. These ion-specific bindings are crucial for maintaining backbone solvation and preventing aggregation in cellular environments. The 2'-OH group in RNA uniquely enables base-catalyzed hydrolysis of the phosphodiester bond via a transesterification mechanism, where the deprotonated 2'-O⁻ acts as a nucleophile to attack the adjacent phosphorus, forming a 2',3'-cyclic phosphate intermediate and cleaving the chain. This reaction proceeds efficiently under alkaline conditions (pH > 7), with rate enhancements from general base catalysis, rendering RNA far less stable than DNA—phosphodiester bonds in DNA are approximately 100-200 times more resistant to such cleavage due to the missing 2'-OH. This inherent lability contributes to RNA's transient nature in vivo, contrasting with DNA's robustness.Nucleotides and polymerization
Nucleotides consist of a nucleobase linked to a pentose sugar (ribose in RNA or deoxyribose in DNA) via a β-N-glycosidic bond, with one to three phosphate groups attached to the 5'-oxygen of the sugar, forming nucleoside monophosphates (NMPs), diphosphates (NDPs), or triphosphates (NTPs).[9] The triphosphate forms are the primary substrates for nucleic acid synthesis: deoxyribonucleoside triphosphates (dNTPs) for DNA and ribonucleoside triphosphates (NTPs) for RNA, providing the energy needed for polymerization through cleavage of the high-energy phosphoanhydride bonds.[10] In cells, NTP levels are maintained higher than NMP or NDP levels to support efficient synthesis, with kinases such as nucleoside diphosphate kinase catalyzing the transfer of phosphate from ATP to NDPs.[11] Nucleic acid polymerization occurs enzymatically via DNA or RNA polymerases, which catalyze the template-directed addition of nucleotides to a growing chain. DNA polymerase, first isolated by Arthur Kornberg in 1956, incorporates dNTPs complementary to a DNA template, forming a phosphodiester bond between the 3'-hydroxyl of the primer terminus and the 5'-phosphate of the incoming dNTP, with concomitant release of pyrophosphate (PPi) that drives the reaction forward. RNA polymerase follows a similar mechanism but initiates de novo without a primer, using NTPs to synthesize RNA complementary to a DNA template starting from a promoter sequence, also releasing PPi; this process was elucidated through studies of bacterial enzymes like E. coli RNA polymerase.[12] Both enzymes require a template strand to ensure base-pairing specificity, with the incoming nucleotide selected via hydrogen bonding to the template base in the active site.[13] Polymerization proceeds exclusively in the 5' to 3' direction, where new nucleotides are added to the 3'-hydroxyl end of the chain, resulting in a linear polymer with 5' phosphate and 3' hydroxyl termini.[14] This directionality arises from the chemical mechanism of nucleophilic attack by the 3'-OH on the α-phosphate of the incoming NTP or dNTP, preventing 3' to 5' synthesis.[15] The release of PPi is often hydrolyzed by pyrophosphatases to shift the equilibrium toward polymer elongation.[16] DNA and RNA synthesis differ in substrates and fidelity: DNA polymerases use dNTPs lacking the 2'-hydroxyl group, enabling a more stable double helix, while RNA polymerases incorporate rNTPs with the 2'-OH, which introduces greater flexibility but higher reactivity.[17] DNA polymerases possess 3' to 5' exonuclease proofreading activity, achieving error rates of approximately 10^{-7} to 10^{-9} per nucleotide, whereas RNA polymerases lack robust proofreading, resulting in higher error rates of about 10^{-4} to 10^{-5}, suitable for transient RNA molecules.[18][19]Primary structure
Nucleotide sequence and composition
The primary structure of nucleic acids is defined as the precise linear sequence of nucleotides, determined by the specific order of their nitrogenous bases—adenine (A), cytosine (C), guanine (G), and thymine (T) in deoxyribonucleic acid (DNA), or uracil (U) replacing thymine in ribonucleic acid (RNA)—connected via phosphodiester bonds in the 5' to 3' direction.[20] This sequence encodes genetic information and is conventionally denoted as a string of single-letter symbols, such as 5'-ATGC-3' for a short DNA segment.[21] The nucleotide composition, particularly the GC content (the percentage of guanine and cytosine bases), significantly affects the stability and thermal properties of the nucleic acid. In double-stranded DNA, higher GC content correlates with increased melting temperature (Tm), the point at which half the double helix dissociates into single strands, due to the stronger hydrogen bonding between G-C pairs compared to A-T pairs. For oligonucleotides shorter than 20 bases under standard PCR conditions (e.g., 50 mM monovalent salt), an approximate Tm is given by the Wallace rule: °C.[22] Chargaff's rules describe the equimolar base ratios observed in most double-stranded DNA molecules: the quantity of adenine equals thymine (A = T), and guanine equals cytosine (G = C), arising from complementary base pairing along the two strands.[23] These parity relationships, established through biochemical analyses of DNA from various organisms, do not apply universally; exceptions occur in single-stranded DNA or certain viral genomes where base pairing is absent or incomplete.[24] Sequence motifs represent recurring patterns within the primary structure, such as simple tandem repeats. A prominent example is the poly-A tail in eukaryotic messenger RNA (mRNA), a homopolymeric stretch of 50–250 adenine residues added post-transcriptionally at the 3' end.[25]Chemical modifications and stability
Chemical modifications to the primary structure of nucleic acids involve the addition of functional groups to nucleobases or the sugar-phosphate backbone, altering their chemical properties without changing the underlying nucleotide sequence. In DNA, one of the most prevalent modifications is 5-methylcytosine (5mC), which occurs primarily at CpG dinucleotides and plays a central role in epigenetic regulation by influencing gene expression through chromatin remodeling and transcriptional repression.[26] This modification is catalyzed by DNA methyltransferases and is essential for processes such as genomic imprinting and X-chromosome inactivation.[27] Another significant DNA modification is N6-methyladenine (m6A), which is widespread in bacterial genomes where it contributes to restriction-modification systems that protect against foreign DNA invasion. In bacteria like Xanthomonas oryzae, m6A is installed by methyltransferases such as Dam and helps regulate replication and repair pathways.[28] In RNA, over 170 distinct chemical modifications have been identified, particularly in ribosomal RNA (rRNA) and transfer RNA (tRNA), where they fine-tune structure and function.[29] [30] Key examples include pseudouridine (Ψ), formed by isomerization of uridine, which enhances base stacking and hydrogen bonding stability; N6-methyladenosine (m6A), the most abundant internal modification in eukaryotic mRNA that affects splicing, export, and translation; and 2'-O-methylation (Nm), which protects against degradation and modulates ribosome assembly.[30] These modifications are enzymatically installed by writer proteins, such as pseudouridine synthases (PUS enzymes) that catalyze the reversible C-C glycosidic bond formation in Ψ without requiring cofactors.[31] For instance, families like TruA and TruB in bacteria and Pus1-Pus10 in eukaryotes target specific sites in tRNA and rRNA.[31] These modifications significantly impact nucleic acid stability by conferring resistance to enzymatic degradation and modulating helical properties. In therapeutic applications, such as small interfering RNA (siRNA), incorporation of 2'-fluoro substitutions at the 2' position of the ribose sugar enhances nuclease resistance, allowing prolonged activity in vivo while maintaining RNA interference efficacy.[32] Similarly, 5mC in DNA reduces backbone flexibility, increasing helix rigidity and protecting against hydrolytic cleavage.[33] In RNA, Ψ and Nm stabilize secondary structures by improving thermodynamic stability and shielding against endonucleases like RNase A.[34] Detection of these modifications relies on specialized techniques that preserve and identify the altered bases. For 5mC in DNA, bisulfite sequencing is a cornerstone method that converts unmethylated cytosines to uracils via sulfonation and deamination, while 5mC remains resistant, enabling differentiation through subsequent PCR amplification and sequencing.[35] This approach provides genome-wide mapping but requires careful optimization to minimize DNA fragmentation. Base composition, particularly CpG density, influences the prevalence of modifiable sites like those for 5mC.[27]Secondary structure
Base pairing rules and hydrogen bonding
In nucleic acids, base pairing refers to the specific association between nucleobases that stabilizes secondary structures through hydrogen bonding. In DNA, the canonical Watson-Crick base pairing rules dictate that adenine (A) pairs with thymine (T), and guanine (G) pairs with cytosine (C). These pairings occur between a purine on one strand and a pyrimidine on the complementary strand, ensuring geometric uniformity in the double helix. The specificity arises from complementary hydrogen bond donor and acceptor sites on the bases, which form precise interactions: the A-T pair involves two hydrogen bonds, while the G-C pair forms three, contributing to greater stability in G-C rich regions.[3][36] The hydrogen bonds in these pairs are typically N-H···O or N-H···N types, involving the Watson-Crick faces of the bases. For A-T, one bond forms between the N1 of adenine (donor) and N3 of thymine (acceptor), and the second between the amino group at C6 of adenine (donor) and the carbonyl at C4 of thymine (acceptor). In G-C pairing, the three bonds are: N1 of guanine to N3 of cytosine, the amino group at C2 of guanine to the carbonyl at C2 of cytosine, and the amino group at C4 of cytosine (donor) to the carbonyl at C6 of guanine (acceptor). These interactions not only dictate pairing fidelity but also influence melting temperatures, with each additional G-C bond increasing duplex stability by approximately 1-2 kcal/mol compared to A-T.[37][38] In RNA, the base pairing rules are analogous but substitute uracil (U) for thymine, forming A-U pairs with two hydrogen bonds (N1 of adenine to N3 of uracil, and amino at C6 of adenine to carbonyl at C4 of uracil) and retaining G-C pairs with three. RNA often adopts single-stranded conformations with intramolecular base pairing to form stems in hairpin loops or other motifs, where hydrogen bonding patterns remain similar but allow for greater flexibility. The G-C pair's stronger bonding (due to the third hydrogen bond) promotes more stable RNA duplexes, as evidenced by higher thermal denaturation temperatures in GC-rich sequences.[39][40] Beyond strict Watson-Crick pairing, non-canonical interactions like wobble base pairs occur, particularly in RNA. Proposed by Francis Crick, the wobble hypothesis describes relaxed base pairing at the third position of codons during translation, allowing a single tRNA to recognize multiple synonymous codons. For instance, guanine in the anticodon can pair with either cytosine or uracil in the mRNA via two hydrogen bonds, shifting the geometry to accommodate the "wobble" without disrupting overall specificity. G-U wobble pairs, common in RNA structures, feature two hydrogen bonds (N1 of G to O2 of U, and O6 of G to N3 of U) and introduce functional diversity, such as in ribosomal RNA where they influence decoding accuracy and structural dynamics. These wobble interactions are weaker than canonical pairs but essential for the degeneracy of the genetic code, reducing the required number of tRNAs from 61 to about 40.[41][42]| Base Pair | Molecule | Hydrogen Bonds | Key Donors/Acceptors |
|---|---|---|---|
| A-T | DNA | 2 | A(N1)-T(N3); A(N6)-T(O4) |
| G-C | DNA/RNA | 3 | G(N1)-C(N3); G(N2)-C(O2); G(O6)-C(N4) |
| A-U | RNA | 2 | A(N1)-U(N3); A(N6)-U(O4) |
| G-U (wobble) | RNA | 2 | G(N1)-U(O2); G(O6)-U(N3) |
DNA double helix configurations
The DNA double helix, formed through complementary base pairing between adenine-thymine and guanine-cytosine, manifests in several distinct configurations that influence its overall geometry and biological function. These variants arise primarily from differences in backbone conformation, base stacking, and hydration levels, with the B-form representing the predominant structure under physiological conditions.[3] B-DNA is a right-handed helix characterized by a smooth, elongated structure with approximately 10.5 base pairs per turn, a helical pitch of 3.4 nm, a rise of 0.34 nm per base pair, and a twist angle of 36° between adjacent base pairs. This configuration features distinct major and minor grooves, which facilitate interactions with proteins for processes such as replication and transcription. The structure was first proposed by Watson and Crick based on X-ray diffraction data, with refined parameters derived from fiber diffraction studies.[3][43] A-DNA, also right-handed but shorter and wider than B-DNA, adopts a more compact form with 11 base pairs per turn, a pitch of about 2.8 nm, a rise of 0.23 nm per base pair, and a twist of approximately 33°. In this conformation, the base pairs are tilted relative to the helix axis, resulting in a deep, narrow minor groove and a shallow major groove. A-DNA is favored under low-humidity conditions, such as in dehydrated fibers, and is commonly observed in DNA-RNA hybrids.[43] Z-DNA represents a left-handed helix with a zig-zag phosphate backbone, accommodating 12 base pairs per turn, a pitch of 4.5 nm, a rise of 0.37 nm per base pair, and a twist of -30°. Unlike the right-handed forms, Z-DNA has a single deep groove and no distinct major/minor distinction, with syn glycosidic conformations for purines and anti for pyrimidines. This form is stabilized in sequences rich in alternating purine-pyrimidine tracts, particularly GC repeats, and was first identified through crystallographic analysis of synthetic oligonucleotides. The prevalence of these helical forms is modulated by environmental factors, including hydration, ionic strength, and sequence composition. B-DNA predominates in aqueous, physiological environments with moderate salt concentrations (e.g., ~150 mM NaCl), while A-DNA emerges at relative humidities below 75% or in the presence of alcohols that reduce water activity. Z-DNA formation is promoted by high salt concentrations (e.g., >2 M NaCl) or multivalent cations like Mg²⁺, which screen phosphate repulsions, and is further enhanced in negatively supercoiled contexts or by specific protein binding, though the latter influences are secondary to ionic effects. Sequence motifs, such as AT-rich regions favoring B-DNA stability through optimal stacking, and GC-rich segments predisposing to Z-DNA via favorable syn-anti alternations, also play a key role. pH variations can induce transitions, with acidic conditions occasionally stabilizing A-like forms by protonating bases and altering hydrogen bonding.[43]| Helix Type | Handedness | Base Pairs per Turn | Pitch (nm) | Rise per Base Pair (nm) | Twist Angle (°) | Key Features |
|---|---|---|---|---|---|---|
| B-DNA | Right | 10.5 | 3.4 | 0.34 | 36 | Major/minor grooves; physiological form |
| A-DNA | Right | 11 | 2.8 | 0.23 | 33 | Tilted bases; low humidity |
| Z-DNA | Left | 12 | 4.5 | 0.37 | -30 | Zig-zag backbone; high salt/GC-rich |
RNA folding motifs
RNA folding motifs are local secondary structural elements that arise from base pairing within single-stranded RNA molecules, enabling diverse functions such as catalysis, regulation, and molecular recognition. Unlike the continuous double helix of DNA, these motifs feature discontinuous helical regions interrupted by unpaired nucleotides, forming compact architectures stabilized by hydrogen bonding and stacking interactions.[44] The most fundamental motif is the stem-loop, consisting of a double-stranded helical stem formed by complementary base pairing and an unpaired loop of 4-7 nucleotides at the apex. Stem-loops can be further diversified by bulges and internal loops, where unpaired nucleotides protrude from one or both strands, respectively, disrupting the continuity of the helix and introducing flexibility or binding sites. For instance, bulge loops with a single unpaired nucleotide on one strand facilitate sharp turns in the RNA backbone, while internal loops with unpaired residues on both strands allow for asymmetric expansions that accommodate tertiary contacts.[44][45] Hairpins represent a specific class of stem-loops where the loop size and sequence confer exceptional stability, particularly tetraloops (four-nucleotide loops) with consensus sequences like GNRA, which exhibit enhanced thermodynamic stability due to non-canonical base interactions and stacking. These tetraloops are among the most stable loop configurations, with free energy contributions up to 4-6 kcal/mol more favorable than larger loops, as determined from optical melting studies. Magnesium ions (Mg²⁺) further stabilize these motifs by bridging negatively charged phosphate groups in loops and stems, reducing electrostatic repulsion and promoting compact folding, especially in bulged or internal regions.[46][47] Beyond simple stem-loops, pseudoknots form when a single-stranded region base-pairs with a complementary sequence outside an existing stem, creating interleaved helices that cross over like a knot and often enhance mechanical rigidity or signaling. Kissing loops occur when the apical loops of two separate hairpins interact via complementary base pairing, forming transient or stable intermolecular contacts that mediate dimerization or regulatory switching.[48][49] In transfer RNA (tRNA), the cloverleaf secondary structure exemplifies the integration of multiple stem-loops, including the acceptor stem, D-loop, anticodon arm, and T-loop, which collectively form four helical arms connected by loops for amino acid attachment and codon recognition. Riboswitches, regulatory RNA elements in bacterial mRNAs, frequently incorporate pseudoknots and kissing loops alongside stems to sense metabolites like thiamine or guanine, undergoing conformational changes that control gene expression.[50][51] Computational tools such as mfold and the ViennaRNA package enable prediction and identification of these motifs by minimizing free energy using dynamic programming algorithms that account for base-pairing rules, loop penalties, and stacking energies. Mfold, developed by Zuker, computes optimal and suboptimal foldings for sequences up to several hundred nucleotides, while ViennaRNA extends this with advanced features like pseudoknot prediction and covariance models for motif detection in alignments.[52][53]Tertiary structure
DNA supercoiling and topology
DNA supercoiling represents a key aspect of the tertiary structure of closed circular DNA molecules, where the double helix, serving as the substrate, undergoes additional coiling beyond its intrinsic helical twist to achieve compaction and facilitate biological processes. In covalently closed circular DNA, such as bacterial plasmids or viral genomes, the topology is invariant unless broken and resealed, leading to superhelical tension that influences DNA accessibility and function.[54] The topological state of supercoiled DNA is quantified by the linking number (Lk), defined as the sum of the twist (Tw), which measures the helical turns of the two strands around each other, and the writhe (Wr), which captures the coiling of the helical axis in space:This relationship, established through mathematical analysis of ribbon topology, holds for any closed DNA duplex.[55] The relaxed linking number (Lk₀) corresponds to the state without supercoiling, typically about 10.5 base pairs per turn in B-form DNA. Supercoiling arises when Lk deviates from Lk₀, quantified by ΔLk = Lk - Lk₀; negative ΔLk indicates underwinding (negative supercoiling), while positive ΔLk indicates overwinding (positive supercoiling). Negative supercoiling predominates in vivo, promoting DNA unwinding for processes like replication and transcription, whereas positive supercoiling can accumulate ahead of progressing polymerases.[54][56] To manage superhelical tension, cells employ DNA topoisomerases, enzymes that transiently break and rejoin DNA strands to alter Lk. Type I topoisomerases, such as the Escherichia coli enzyme discovered in 1971, relax supercoils by nicking one strand, changing Lk in steps of ±1 without requiring ATP; they preferentially relieve negative supercoils.[57] Type II topoisomerases, including DNA gyrase, act on both strands, altering Lk in steps of ±2 and often requiring ATP; while most type II enzymes relax supercoils bidirectionally, gyrase uniquely introduces negative supercoils using ATP hydrolysis, counteracting the positive supercoils generated during transcription. In bacteria, gyrase maintains an overall negative superhelical density (σ ≈ -0.06) essential for chromosomal compaction within the confined nucleoid space.[58] Supercoiled DNA adopts distinct three-dimensional configurations to partition the writhe component. Plectonemic supercoils form right-handed interwound structures where the DNA axis coils around itself, typical in unconstrained bacterial DNA and allowing dynamic partitioning of twist and writhe. In contrast, toroidal (or solenoidal) supercoils involve the DNA wrapping left-handedly around a core, as seen in eukaryotic nucleosomes where approximately 147 base pairs wrap 1.65–1.7 turns around the histone octamer, contributing about -1 supercoil per nucleosome.[59] This wrapping constrains negative supercoils, reducing free writhe and aiding chromatin compaction. In bacteria, negative supercoiling driven by gyrase compacts the genome by favoring plectonemic structures and branched domains, while also linking to replication by relieving torsional stress at forks and to transcription by enhancing promoter opening and RNA polymerase progression.[60][61]
.png/250px-DNA_RNA_structure_(full).png)
.png)
