Recent from talks
Nothing was collected or created yet.
Nucleosome
View on Wikipedia
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone proteins[1] and resembles thread wrapped around a spool. The nucleosome is the fundamental subunit of chromatin. Each nucleosome is composed of a little less than two turns of DNA wrapped around a set of eight proteins called histones, which are known as a histone octamer. Each histone octamer is composed of two copies each of the histone proteins H2A, H2B, H3, and H4.
DNA must be compacted into nucleosomes to fit within the cell nucleus.[2] In addition to nucleosome wrapping, eukaryotic chromatin is further compacted by being folded into a series of more complex structures, eventually forming a chromosome. Each human cell contains about 30 million nucleosomes.[3]
Nucleosomes are thought to carry epigenetically inherited information in the form of covalent modifications of their core histones. Nucleosome positions in the genome are not random, and it is important to know where each nucleosome is located because this determines the accessibility of the DNA to regulatory proteins.[4]
Nucleosomes were first observed as particles in the electron microscope by Don and Ada Olins in 1974,[5] and their existence and structure (as histone octamers surrounded by approximately 200 base pairs of DNA) were proposed by Roger Kornberg.[6][7] The role of the nucleosome as a regulator of transcription was demonstrated by Lorch et al. in vitro[8] in 1987 and by Han and Grunstein[9] and Clark-Adams et al.[10] in vivo in 1988.
The nucleosome core particle consists of approximately 146 base pairs (bp) of DNA[11] wrapped in 1.67 left-handed superhelical turns around a histone octamer, consisting of 2 copies each of the core histones H2A, H2B, H3, and H4.[12] Core particles are connected by stretches of linker DNA, which can be up to about 80 bp long. Technically, a nucleosome is defined as the core particle plus one of these linker regions; however the word is often synonymous with the core particle.[13] Genome-wide nucleosome positioning maps are now available for many model organisms and human cells.[14]
Linker histones such as H1 and its isoforms are involved in chromatin compaction and sit at the base of the nucleosome near the DNA entry and exit binding to the linker region of the DNA.[15] Non-condensed nucleosomes without the linker histone resemble "beads on a string of DNA" under an electron microscope.[16]
In contrast to most eukaryotic cells, mature sperm cells largely use protamines to package their genomic DNA, most likely to achieve an even higher packaging ratio.[17] Histone equivalents and a simplified chromatin structure have also been found in Archaea,[18] suggesting that eukaryotes are not the only organisms that use nucleosomes.
Structure
[edit]Structure of the core particle
[edit]
Overview
[edit]Pioneering structural studies in the 1980s by Aaron Klug's group provided the first evidence that an octamer of histone proteins wraps DNA around itself in about 1.7 turns of a left-handed superhelix.[19] In 1997 the first near atomic resolution crystal structure of the nucleosome was solved by the Richmond group at the ETH Zurich, showing the most important details of the particle. The human alpha satellite palindromic DNA critical to achieving the 1997 nucleosome crystal structure was developed by the Bunick group at Oak Ridge National Laboratory in Tennessee.[20][21][22][23][24] The structures of over 20 different nucleosome core particles have been solved to date,[25] including those containing histone variants and histones from different species. The structure of the nucleosome core particle is remarkably conserved, and even a change of over 100 residues between frog and yeast histones results in electron density maps with an overall root mean square deviation of only 1.6Å.[26]
The nucleosome core particle (NCP)
[edit]The nucleosome core particle (shown in the figure) consists of about 146 base pair of DNA[11] wrapped in 1.67 left-handed superhelical turns around the histone octamer, consisting of 2 copies each of the core histones H2A, H2B, H3, and H4. Adjacent nucleosomes are joined by a stretch of free DNA termed linker DNA (which varies from 10 - 80 bp in length depending on species and tissue type[18]).The whole structure generates a cylinder of diameter 11 nm and a height of 5.5 nm.


Nucleosome core particles are observed when chromatin in interphase is treated to cause the chromatin to unfold partially. The resulting image, via an electron microscope, is "beads on a string". The string is the DNA, while each bead in the nucleosome is a core particle. The nucleosome core particle is composed of DNA and histone proteins.[29]
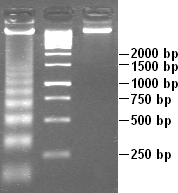
Partial DNAse digestion of chromatin reveals its nucleosome structure. Because DNA portions of nucleosome core particles are less accessible for DNAse than linking sections, DNA gets digested into fragments of lengths equal to multiplicity of distance between nucleosomes (180, 360, 540 base pairs etc.). Hence a very characteristic pattern similar to a ladder is visible during gel electrophoresis of that DNA.[27] Such digestion can occur also under natural conditions during apoptosis ("cell suicide" or programmed cell death), because autodestruction of DNA typically is its role.[30]
Protein interactions within the nucleosome
[edit]The core histone proteins contains a characteristic structural motif termed the "histone fold", which consists of three alpha-helices (α1-3) separated by two loops (L1-2). In solution, the histones form H2A-H2B heterodimers and H3-H4 heterotetramers. Histones dimerise about their long α2 helices in an anti-parallel orientation, and, in the case of H3 and H4, two such dimers form a 4-helix bundle stabilised by extensive H3-H3' interaction. The H2A/H2B dimer binds onto the H3/H4 tetramer due to interactions between H4 and H2B, which include the formation of a hydrophobic cluster.[12] The histone octamer is formed by a central H3/H4 tetramer sandwiched between two H2A/H2B dimers. Due to the highly basic charge of all four core histones, the histone octamer is stable only in the presence of DNA or very high salt concentrations.
Histone - DNA interactions
[edit]The nucleosome contains over 120 direct protein-DNA interactions and several hundred water-mediated ones.[31] Direct protein - DNA interactions are not spread evenly about the octamer surface but rather located at discrete sites. These are due to the formation of two types of DNA binding sites within the octamer; the α1α1 site, which uses the α1 helix from two adjacent histones, and the L1L2 site formed by the L1 and L2 loops. Salt links and hydrogen bonding between both side-chain basic and hydroxyl groups and main-chain amides with the DNA backbone phosphates form the bulk of interactions with the DNA. This is important, given that the ubiquitous distribution of nucleosomes along genomes requires it to be a non-sequence-specific DNA-binding factor. Although nucleosomes tend to prefer some DNA sequences over others,[32] they are capable of binding practically to any sequence, which is thought to be due to the flexibility in the formation of these water-mediated interactions. In addition, non-polar interactions are made between protein side-chains and the deoxyribose groups, and an arginine side-chain intercalates into the DNA minor groove at all 14 sites where it faces the octamer surface. The distribution and strength of DNA-binding sites about the octamer surface distorts the DNA within the nucleosome core. The DNA is non-uniformly bent and also contains twist defects. The twist of free B-form DNA in solution is 10.5 bp per turn. However, the overall twist of nucleosomal DNA is only 10.2 bp per turn, varying from a value of 9.4 to 10.9 bp per turn.
Histone tail domains
[edit]The histone tail extensions constitute up to 30% by mass of histones, but are not visible in the crystal structures of nucleosomes due to their high intrinsic flexibility, and have been thought to be largely unstructured.[33] The N-terminal tails of histones H3 and H2B pass through a channel formed by the minor grooves of the two DNA strands, protruding from the DNA every 20 bp. The N-terminal tail of histone H4, on the other hand, has a region of highly basic amino acids (16–25), which, in the crystal structure, forms an interaction with the highly acidic surface region of a H2A-H2B dimer of another nucleosome, being potentially relevant for the higher-order structure of nucleosomes. This interaction is thought to occur under physiological conditions also, and suggests that acetylation of the H4 tail distorts the higher-order structure of chromatin.[citation needed]
Higher order structure
[edit]
The organization of the DNA that is achieved by the nucleosome cannot fully explain the packaging of DNA observed in the cell nucleus. Further compaction of chromatin into the cell nucleus is necessary, but it is not yet well understood. The current understanding[25] is that repeating nucleosomes with intervening "linker" DNA form a 10-nm-fiber, described as "beads on a string", and have a packing ratio of about five to ten.[18] A chain of nucleosomes can be arranged in a 30 nm fiber, a compacted structure with a packing ratio of ~50[18] and whose formation is dependent on the presence of the H1 histone.
A crystal structure of a tetranucleosome has been presented and used to build up a proposed structure of the 30 nm fiber as a two-start helix.[34] There is still a certain amount of contention regarding this model, as it is incompatible with recent electron microscopy data.[35] Beyond this, the structure of chromatin is poorly understood, but it is classically suggested that the 30 nm fiber is arranged into loops along a central protein scaffold to form transcriptionally active euchromatin. Further compaction leads to transcriptionally inactive heterochromatin.
Dynamics
[edit]Although the nucleosome is a very stable protein-DNA complex, it is not static and has been shown to undergo a number of different structural re-arrangements including nucleosome sliding and DNA site exposure. Depending on the context, nucleosomes can inhibit or facilitate transcription factor binding. Nucleosome positions are controlled by three major contributions: First, the intrinsic binding affinity of the histone octamer depends on the DNA sequence. Second, the nucleosome can be displaced or recruited by the competitive or cooperative binding of other protein factors. Third, the nucleosome may be actively translocated by ATP-dependent remodeling complexes.[36]
Nucleosome sliding
[edit]When incubated thermally, nucleosomes reconstituted onto the 5S DNA positioning sequence were able to reposition themselves translationally onto adjacent sequences.[37] This repositioning does not require disruption of the histone octamer but is consistent with nucleosomes being able to "slide" along the DNA in cis. CTCF binding sites act as nucleosome positioning anchors so that, when used to align various genomic signals, multiple flanking nucleosomes can be readily identified.[38] Although nucleosomes are intrinsically mobile, eukaryotes have evolved a large family of ATP-dependent chromatin remodelling enzymes to alter chromatin structure, many of which do so via nucleosome sliding. Nucleosome sliding is one of the possible mechanism for large scale tissue specific expression of genes. The transcription start site for genes expressed in a particular tissue, are nucleosome depleted while, the same set of genes in other tissue where they are not expressed, are nucleosome bound.[39]
DNA site exposure
[edit]Nucleosomal DNA is in equilibrium between a wrapped and unwrapped state. DNA within the nucleosome remains fully wrapped for only 250 ms before it is unwrapped for 10-50 ms and then rapidly rewrapped, as measured using time-resolved FRET.[40] This implies that DNA does not need to be actively dissociated from the nucleosome but that there is a significant fraction of time during which it is fully accessible. Introducing a DNA-binding sequence within the nucleosome increases the accessibility of adjacent regions of DNA when bound.[41]
This propensity for DNA within the nucleosome to "breathe" has important functional consequences for all DNA-binding proteins that operate in a chromatin environment.[40] In particular, the dynamic breathing of nucleosomes plays an important role in restricting the advancement of RNA polymerase II during transcription elongation.[42]
Nucleosome free region
[edit]Promoters of active genes have nucleosome free regions (NFR). This allows for promoter DNA accessibility to various proteins, such as transcription factors. Nucleosome free region typically spans for 200 nucleotides in S. cerevisiae[43] Well-positioned nucleosomes form boundaries of NFR. These nucleosomes are called +1-nucleosome and −1-nucleosome and are located at canonical distances downstream and upstream, respectively, from transcription start site.[44] +1-nucleosome and several downstream nucleosomes also tend to incorporate H2A.Z histone variant.[44]
Modulating nucleosome structure
[edit]Eukaryotic genomes are ubiquitously associated into chromatin; however, cells must spatially and temporally regulate specific loci independently of bulk chromatin. In order to achieve the high level of control required to co-ordinate nuclear processes such as DNA replication, repair, and transcription, cells have developed a variety of means to locally and specifically modulate chromatin structure and function. This can involve covalent modification of histones, the incorporation of histone variants, and non-covalent remodelling by ATP-dependent remodeling enzymes.
Histone post-translational modifications
[edit]
Since they were discovered in the mid-1960s, histone modifications have been predicted to affect transcription.[45] The fact that most of the early post-translational modifications found were concentrated within the tail extensions that protrude from the nucleosome core lead to two main theories regarding the mechanism of histone modification. The first of the theories suggested that they may affect electrostatic interactions between the histone tails and DNA to "loosen" chromatin structure. Later it was proposed that combinations of these modifications may create binding epitopes with which to recruit other proteins.[46] Recently, given that more modifications have been found in the structured regions of histones, it has been put forward that these modifications may affect histone-DNA[47] and histone-histone[48] interactions within the nucleosome core. Modifications (such as acetylation or phosphorylation) that lower the charge of the globular histone core are predicted to "loosen" core-DNA association; the strength of the effect depends on location of the modification within the core.[49] Some modifications have been shown to be correlated with gene silencing; others seem to be correlated with gene activation. Common modifications include acetylation, methylation, or ubiquitination of lysine; methylation of arginine; and phosphorylation of serine. The information stored in this way is considered epigenetic, since it is not encoded in the DNA but is still inherited to daughter cells. The maintenance of a repressed or activated status of a gene is often necessary for cellular differentiation.[18]
Histone variants
[edit]Although histones are remarkably conserved throughout evolution, several variant forms have been identified. This diversification of histone function is restricted to H2A and H3, with H2B and H4 being mostly invariant. H2A can be replaced by H2AZ (which leads to reduced nucleosome stability) or H2AX (which is associated with DNA repair and T cell differentiation), whereas the inactive X chromosomes in mammals are enriched in macroH2A. H3 can be replaced by H3.3 (which correlates with activate genes and regulatory elements) and in centromeres H3 is replaced by CENPA.[18]
ATP-dependent nucleosome remodeling
[edit]A number of distinct reactions are associated with the term ATP-dependent chromatin remodeling. Remodeling enzymes have been shown to slide nucleosomes along DNA,[50] disrupt histone-DNA contacts to the extent of destabilizing the H2A/H2B dimer[51][52] and to generate negative superhelical torsion in DNA and chromatin.[53] Recently, the Swr1 remodeling enzyme has been shown to introduce the variant histone H2A.Z into nucleosomes.[54] At present, it is not clear if all of these represent distinct reactions or merely alternative outcomes of a common mechanism. What is shared between all, and indeed the hallmark of ATP-dependent chromatin remodeling, is that they all result in altered DNA accessibility.
Studies looking at gene activation in vivo[55] and, more astonishingly, remodeling in vitro[56] have revealed that chromatin remodeling events and transcription-factor binding are cyclical and periodic in nature. While the consequences of this for the reaction mechanism of chromatin remodeling are not known, the dynamic nature of the system may allow it to respond faster to external stimuli. A recent study indicates that nucleosome positions change significantly during mouse embryonic stem cell development, and these changes are related to binding of developmental transcription factors.[57]
Dynamic nucleosome remodelling across the Yeast genome
[edit]Studies in 2007 have catalogued nucleosome positions in yeast and shown that nucleosomes are depleted in promoter regions and origins of replication.[58][59][60] About 80% of the yeast genome appears to be covered by nucleosomes[61] and the pattern of nucleosome positioning clearly relates to DNA regions that regulate transcription, regions that are transcribed and regions that initiate DNA replication.[62] Most recently, a new study examined dynamic changes in nucleosome repositioning during a global transcriptional reprogramming event to elucidate the effects on nucleosome displacement during genome-wide transcriptional changes in yeast (Saccharomyces cerevisiae).[63] The results suggested that nucleosomes that were localized to promoter regions are displaced in response to stress (like heat shock). In addition, the removal of nucleosomes usually corresponded to transcriptional activation and the replacement of nucleosomes usually corresponded to transcriptional repression, presumably because transcription factor binding sites became more or less accessible, respectively. In general, only one or two nucleosomes were repositioned at the promoter to effect these transcriptional changes. However, even in chromosomal regions that were not associated with transcriptional changes, nucleosome repositioning was observed, suggesting that the covering and uncovering of transcriptional DNA does not necessarily produce a transcriptional event. After transcription, the rDNA region has to protected from any damage, it suggested HMGB proteins play a major role in protecting the nucleosome free region.[64][65]
DNA Twist Defects
[edit]DNA twist defects are when the addition of one or a few base pairs from one DNA segment are transferred to the next segment resulting in a change of the DNA twist. This will not only change the twist of the DNA but it will also change the length.[66] This twist defect eventually moves around the nucleosome through the transferring of the base pair, this means DNA twists can cause nucleosome sliding.[67] Nucleosome crystal structures have shown that superhelix location 2 and 5 on the nucleosome are commonly found to be where DNA twist defects occur as these are common remodeler binding sites.[68] There are a variety of chromatin remodelers but all share the existence of an ATPase motor which facilitates chromatin sliding on DNA through the binding and hydrolysis of ATP.[69] ATPase has an open and closed state. When the ATPase motor is changing from open and closed states, the DNA duplex changes geometry and exhibits base pair tilting.[68] The initiation of the twist defects via the ATPase motor causes tension to accumulate around the remodeler site. The tension is released when the sliding of DNA has been completed throughout the nucleosome via the spread of two twist defects (one on each strand) in opposite directions.[69]
Nucleosome assembly in vitro
[edit]
Nucleosomes can be assembled in vitro by either using purified native or recombinant histones.[70][71] One standard technique of loading the DNA around the histones involves the use of salt dialysis. A reaction consisting of the histone octamers and a naked DNA template can be incubated together at a salt concentration of 2 M. By steadily decreasing the salt concentration, the DNA will equilibrate to a position where it is wrapped around the histone octamers, forming nucleosomes. In appropriate conditions, this reconstitution process allows for the nucleosome positioning affinity of a given sequence to be mapped experimentally.[72]
Disulfide crosslinked nucleosome core particles
[edit]A recent advance in the production of nucleosome core particles with enhanced stability involves site-specific disulfide crosslinks.[73] Two different crosslinks can be introduced into the nucleosome core particle. A first one crosslinks the two copies of H2A via an introduced cysteine (N38C) resulting in histone octamer which is stable against H2A/H2B dimer loss during nucleosome reconstitution. A second crosslink can be introduced between the H3 N-terminal histone tail and the nucleosome DNA ends via an incorporated convertible nucleotide.[74] The DNA-histone octamer crosslink stabilizes the nucleosome core particle against DNA dissociation at very low particle concentrations and at elevated salt concentrations.
Nucleosome assembly in vivo
[edit]
Nucleosomes are the basic packing unit of genomic DNA built from histone proteins around which DNA is coiled. They serve as a scaffold for formation of higher order chromatin structure as well as for a layer of regulatory control of gene expression. Nucleosomes are quickly assembled onto newly synthesized DNA behind the replication fork.
H3 and H4
[edit]Histones H3 and H4 from disassembled old nucleosomes are kept in the vicinity and randomly distributed on the newly synthesized DNA.[75] They are assembled by the chromatin assembly factor 1 (CAF-1) complex, which consists of three subunits (p150, p60, and p48).[76] Newly synthesized H3 and H4 are assembled by the replication coupling assembly factor (RCAF). RCAF contains the subunit Asf1, which binds to newly synthesized H3 and H4 proteins.[77] The old H3 and H4 proteins retain their chemical modifications which contributes to the passing down of the epigenetic signature. The newly synthesized H3 and H4 proteins are gradually acetylated at different lysine residues as part of the chromatin maturation process.[78] It is also thought that the old H3 and H4 proteins in the new nucleosomes recruit histone modifying enzymes that mark the new histones, contributing to epigenetic memory.
H2A and H2B
[edit]In contrast to old H3 and H4, the old H2A and H2B histone proteins are released and degraded; therefore, newly assembled H2A and H2B proteins are incorporated into new nucleosomes.[79] H2A and H2B are assembled into dimers which are then loaded onto nucleosomes by the nucleosome assembly protein-1 (NAP-1) which also assists with nucleosome sliding.[80] The nucleosomes are also spaced by ATP-dependent nucleosome-remodeling complexes containing enzymes such as Isw1 Ino80, and Chd1, and subsequently assembled into higher order structure.[81][82]
Gallery
[edit]The crystal structure of the nucleosome core particle (PDB: 1EQZ[28]) - different views showing details of histone folding and organization. Histones H2A, H2B, H3, H4 and DNA are coloured.
See also
[edit]References
[edit]- ^ Reece J, Campbell N (2006). Biology. San Francisco: Benjamin Cummings. ISBN 978-0-8053-6624-2.
- ^ Alberts B (2002). "Chromosomal DNA and Its Packaging in the Chromatin Fiber". Molecular biology of the cell (4th ed.). New York: Garland Science. p. 207. ISBN 978-0-8153-4072-0.
- ^ lberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002). Chromosomal DNA and Its Packaging in the Chromatin Fiber. Garland Science.
- ^ Teif VB, Clarkson CT (2019). "Nucleosome Positioning". Encyclopedia of Bioinformatics and Computational Biology. Vol. 2. pp. 308–317. doi:10.1016/B978-0-12-809633-8.20242-2. ISBN 978-0-12-811432-2. S2CID 43929234.
- ^ Olins AL, Olins DE (January 1974). "Spheroid chromatin units (v bodies)". Science. 183 (4122): 330–332. Bibcode:1974Sci...183..330O. doi:10.1126/science.183.4122.330. PMID 4128918. S2CID 83480762.
- ^ McDonald D (December 2005). "Milestone 9, (1973-1974) The nucleosome hypothesis: An alternative string theory". Nature Milestones: Gene Expression. doi:10.1038/nrm1798.
- ^ Kornberg RD (May 1974). "Chromatin structure: a repeating unit of histones and DNA". Science. 184 (4139): 868–871. Bibcode:1974Sci...184..868K. doi:10.1126/science.184.4139.868. PMID 4825889.
- ^ Lorch Y, LaPointe JW, Kornberg RD (April 1987). "Nucleosomes inhibit the initiation of transcription but allow chain elongation with the displacement of histones". Cell. 49 (2): 203–210. doi:10.1016/0092-8674(87)90561-7. PMID 3568125. S2CID 21270171.
- ^ Han M, Grunstein M (December 1988). "Nucleosome loss activates yeast downstream promoters in vivo". Cell. 55 (6): 1137–1145. doi:10.1016/0092-8674(88)90258-9. PMID 2849508. S2CID 41520634.
- ^ Clark-Adams CD, Norris D, Osley MA, Fassler JS, Winston F (February 1988). "Changes in histone gene dosage alter transcription in yeast". Genes & Development. 2 (2): 150–159. doi:10.1101/gad.2.2.150. PMID 2834270.
- ^ a b In different crystals, values of 146 and 147 basepairs were observed
- ^ a b Luger K, Mäder AW, Richmond RK, Sargent DF, Richmond TJ (September 1997). "Crystal structure of the nucleosome core particle at 2.8 A resolution". Nature. 389 (6648): 251–260. Bibcode:1997Natur.389..251L. doi:10.1038/38444. PMID 9305837. S2CID 4328827.
- ^ Alberts B (2007). Molecular Biology of the Cell (5th ed.). New York: Garland Science. p. 211. ISBN 978-0-8153-4106-2.
- ^ Shtumpf M, Piroeva KV, Agrawal SP, Jacob DR, Teif VB (June 2022). "NucPosDB: a database of nucleosome positioning in vivo and nucleosomics of cell-free DNA". Chromosoma. 131 (1–2): 19–28. doi:10.1007/s00412-021-00766-9. PMC 8776978. PMID 35061087.
- ^ Zhou YB, Gerchman SE, Ramakrishnan V, Travers A, Muyldermans S (September 1998). "Position and orientation of the globular domain of linker histone H5 on the nucleosome". Nature. 395 (6700): 402–405. Bibcode:1998Natur.395..402Z. doi:10.1038/26521. PMID 9759733. S2CID 204997317.
- ^ Thoma F, Koller T, Klug A (November 1979). "Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin". The Journal of Cell Biology. 83 (2 Pt 1): 403–427. doi:10.1083/jcb.83.2.403. PMC 2111545. PMID 387806.
- ^ Clarke HJ (1992). "Nuclear and chromatin composition of mammalian gametes and early embryos". Biochemistry and Cell Biology. 70 (10–11): 856–866. Bibcode:1992BCB....70..856C. doi:10.1139/o92-134. PMID 1297351.
- ^ a b c d e f Felsenfeld G, Groudine M (January 2003). "Controlling the double helix". Nature. 421 (6921): 448–453. Bibcode:2003Natur.421..448F. doi:10.1038/nature01411. PMID 12540921.
- ^ Richmond TJ, Finch JT, Rushton B, Rhodes D, Klug A (1984). "Structure of the nucleosome core particle at 7 A resolution". Nature. 311 (5986): 532–7. Bibcode:1984Natur.311..532R. doi:10.1038/311532a0. PMID 6482966. S2CID 4355982.
- ^ Harp JM, Palmer EL, York MH, Gewiess A, Davis M, Bunick GJ (October 1995). "Preparative separation of nucleosome core particles containing defined-sequence DNA in multiple translational phases". Electrophoresis. 16 (10): 1861–1864. doi:10.1002/elps.11501601305. PMID 8586054. S2CID 20178479.
- ^ Palmer EL, Gewiess A, Harp JM, York MH, Bunick GJ (October 1995). "Large-scale production of palindrome DNA fragments". Analytical Biochemistry. 231 (1): 109–114. doi:10.1006/abio.1995.1509. PMID 8678288.
- ^ Harp JM, Uberbacher EC, Roberson AE, Palmer EL, Gewiess A, Bunick GJ (March 1996). "X-ray diffraction analysis of crystals containing twofold symmetric nucleosome core particles". Acta Crystallographica. Section D, Biological Crystallography. 52 (Pt 2): 283–288. Bibcode:1996AcCrD..52..283H. doi:10.1107/S0907444995009139. PMID 15299701.
- ^ Harp JM, Hanson BL, Timm DE, Bunick GJ (December 2000). "Asymmetries in the nucleosome core particle at 2.5 A resolution". Acta Crystallographica. Section D, Biological Crystallography. 56 (Pt 12): 1513–1534. doi:10.1107/s0907444900011847. PMID 11092917.
- ^ Hanson BL, Alexander C, Harp JM, Bunick GJ (2004). "Preparation and crystallization of nucleosome core particle". Chromatin and Chromatin Remodeling Enzymes, Part A. Methods in Enzymology. Vol. 375. pp. 44–62. doi:10.1016/s0076-6879(03)75003-4. ISBN 978-0-12-182779-3. PMID 14870658.
- ^ a b Chakravarthy S, Park YJ, Chodaparambil J, Edayathumangalam RS, Luger K (February 2005). "Structure and dynamic properties of nucleosome core particles". FEBS Letters. 579 (4): 895–898. Bibcode:2005FEBSL.579..895C. doi:10.1016/j.febslet.2004.11.030. PMID 15680970. S2CID 41706403.
- ^ White CL, Suto RK, Luger K (September 2001). "Structure of the yeast nucleosome core particle reveals fundamental changes in internucleosome interactions". The EMBO Journal. 20 (18): 5207–5218. doi:10.1093/emboj/20.18.5207. PMC 125637. PMID 11566884.
- ^ a b Stryer L (1995). Biochemistry (fourth ed.). New York - Basingstoke: W. H. Freeman and Company. ISBN 978-0-7167-2009-6.
- ^ a b Harp JM, Hanson BL, Timm DE, Bunick GJ (December 2000). "Asymmetries in the nucleosome core particle at 2.5 A resolution". Acta Crystallographica. Section D, Biological Crystallography. 56 (Pt 12): 1513–1534. doi:10.1107/S0907444900011847. PMID 11092917. PDB ID: 1EQZ.
- ^ Alberts B (2009). Essential Cell Biology (2nd ed.). New York: Garland Science.
- ^ Allen, Paul D.; Newland, Adrian C. (1 June 1998). "Electrophoretic DNA analysis for the detection of apoptosis". Molecular Biotechnology. 9 (3): 247–251. doi:10.1007/BF02915798. ISSN 1559-0305. PMID 9718585.
- ^ Davey CA, Sargent DF, Luger K, Maeder AW, Richmond TJ (June 2002). "Solvent mediated interactions in the structure of the nucleosome core particle at 1.9 a resolution". Journal of Molecular Biology. 319 (5): 1097–1113. doi:10.1016/S0022-2836(02)00386-8. PMID 12079350.
- ^ Segal E, Fondufe-Mittendorf Y, Chen L, Thåström A, Field Y, Moore IK, et al. (August 2006). "A genomic code for nucleosome positioning". Nature. 442 (7104): 772–778. Bibcode:2006Natur.442..772S. doi:10.1038/nature04979. PMC 2623244. PMID 16862119.
- ^ Zheng C, Hayes JJ (April 2003). "Structures and interactions of the core histone tail domains". Biopolymers. 68 (4): 539–546. doi:10.1002/bip.10303. PMID 12666178.
- ^ Schalch T, Duda S, Sargent DF, Richmond TJ (July 2005). "X-ray structure of a tetranucleosome and its implications for the chromatin fibre". Nature. 436 (7047): 138–141. Bibcode:2005Natur.436..138S. doi:10.1038/nature03686. PMID 16001076. S2CID 4387396.
- ^ Robinson PJ, Fairall L, Huynh VA, Rhodes D (April 2006). "EM measurements define the dimensions of the "30-nm" chromatin fiber: evidence for a compact, interdigitated structure". Proceedings of the National Academy of Sciences of the United States of America. 103 (17): 6506–6511. Bibcode:2006PNAS..103.6506R. doi:10.1073/pnas.0601212103. PMC 1436021. PMID 16617109.
- ^ Teif VB, Rippe K (September 2009). "Predicting nucleosome positions on the DNA: combining intrinsic sequence preferences and remodeler activities". Nucleic Acids Research. 37 (17): 5641–5655. doi:10.1093/nar/gkp610. PMC 2761276. PMID 19625488.
- ^ Pennings S, Muyldermans S, Meersseman G, Wyns L (May 1989). "Formation, stability and core histone positioning of nucleosomes reassembled on bent and other nucleosome-derived DNA". Journal of Molecular Biology. 207 (1): 183–192. doi:10.1016/0022-2836(89)90449-X. PMID 2738923.
- ^ Fu Y, Sinha M, Peterson CL, Weng Z (July 2008). Van Steensel B (ed.). "The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome". PLOS Genetics. 4 (7) e1000138. doi:10.1371/journal.pgen.1000138. PMC 2453330. PMID 18654629.
- ^ Bargaje R, Alam MP, Patowary A, Sarkar M, Ali T, Gupta S, et al. (October 2012). "Proximity of H2A.Z containing nucleosome to the transcription start site influences gene expression levels in the mammalian liver and brain". Nucleic Acids Research. 40 (18): 8965–8978. doi:10.1093/nar/gks665. PMC 3467062. PMID 22821566.
- ^ a b Li G, Levitus M, Bustamante C, Widom J (January 2005). "Rapid spontaneous accessibility of nucleosomal DNA". Nature Structural & Molecular Biology. 12 (1): 46–53. doi:10.1038/nsmb869. PMID 15580276. S2CID 14540078.
- ^ Li G, Widom J (August 2004). "Nucleosomes facilitate their own invasion". Nature Structural & Molecular Biology. 11 (8): 763–769. doi:10.1038/nsmb801. PMID 15258568. S2CID 11299024.
- ^ Hodges C, Bintu L, Lubkowska L, Kashlev M, Bustamante C (July 2009). "Nucleosomal fluctuations govern the transcription dynamics of RNA polymerase II". Science. 325 (5940): 626–628. Bibcode:2009Sci...325..626H. doi:10.1126/science.1172926. PMC 2775800. PMID 19644123.
- ^ Yuan GC, Liu YJ, Dion MF, Slack MD, Wu LF, Altschuler SJ, Rando OJ (July 2005). "Genome-scale identification of nucleosome positions in S. cerevisiae". Science. 309 (5734): 626–630. Bibcode:2005Sci...309..626Y. doi:10.1126/science.1112178. PMID 15961632. S2CID 43625066.
- ^ a b Lai WK, Pugh BF (September 2017). "Understanding nucleosome dynamics and their links to gene expression and DNA replication". Nature Reviews. Molecular Cell Biology. 18 (9): 548–562. doi:10.1038/nrm.2017.47. PMC 5831138. PMID 28537572.
- ^ Allfrey VG, Faulkner R, Mirsky AE (May 1964). "Acetylation and Methylation of Histones and Their Possible Role in the Regulation of RNA Synthesis". Proceedings of the National Academy of Sciences of the United States of America. 51 (5): 786–794. Bibcode:1964PNAS...51..786A. doi:10.1073/pnas.51.5.786. PMC 300163. PMID 14172992.
- ^ Strahl BD, Allis CD (January 2000). "The language of covalent histone modifications". Nature. 403 (6765): 41–45. Bibcode:2000Natur.403...41S. doi:10.1038/47412. PMID 10638745. S2CID 4418993.
- ^ Cosgrove MS, Boeke JD, Wolberger C (November 2004). "Regulated nucleosome mobility and the histone code". Nature Structural & Molecular Biology. 11 (11): 1037–1043. doi:10.1038/nsmb851. PMID 15523479. S2CID 34704745.
- ^ Ye J, Ai X, Eugeni EE, Zhang L, Carpenter LR, Jelinek MA, et al. (April 2005). "Histone H4 lysine 91 acetylation a core domain modification associated with chromatin assembly". Molecular Cell. 18 (1): 123–130. doi:10.1016/j.molcel.2005.02.031. PMC 2855496. PMID 15808514.
- ^ Fenley AT, Adams DA, Onufriev AV (September 2010). "Charge state of the globular histone core controls stability of the nucleosome". Biophysical Journal. 99 (5): 1577–1585. Bibcode:2010BpJ....99.1577F. doi:10.1016/j.bpj.2010.06.046. PMC 2931741. PMID 20816070.
- ^ Whitehouse I, Flaus A, Cairns BR, White MF, Workman JL, Owen-Hughes T (August 1999). "Nucleosome mobilization catalysed by the yeast SWI/SNF complex". Nature. 400 (6746): 784–787. Bibcode:1999Natur.400..784W. doi:10.1038/23506. PMID 10466730. S2CID 2841873.
- ^ Kassabov SR, Zhang B, Persinger J, Bartholomew B (February 2003). "SWI/SNF unwraps, slides, and rewraps the nucleosome". Molecular Cell. 11 (2): 391–403. doi:10.1016/S1097-2765(03)00039-X. PMID 12620227.
- ^ Bruno M, Flaus A, Stockdale C, Rencurel C, Ferreira H, Owen-Hughes T (December 2003). "Histone H2A/H2B dimer exchange by ATP-dependent chromatin remodeling activities". Molecular Cell. 12 (6): 1599–1606. doi:10.1016/S1097-2765(03)00499-4. PMC 3428624. PMID 14690611.
- ^ Havas K, Flaus A, Phelan M, Kingston R, Wade PA, Lilley DM, Owen-Hughes T (December 2000). "Generation of superhelical torsion by ATP-dependent chromatin remodeling activities". Cell. 103 (7): 1133–1142. doi:10.1016/S0092-8674(00)00215-4. PMID 11163188. S2CID 7911590.
- ^ Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C (January 2004). "ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex". Science. 303 (5656): 343–348. Bibcode:2004Sci...303..343M. doi:10.1126/science.1090701. PMID 14645854. S2CID 9881829.
- ^ Métivier R, Penot G, Hübner MR, Reid G, Brand H, Kos M, Gannon F (December 2003). "Estrogen receptor-alpha directs ordered, cyclical, and combinatorial recruitment of cofactors on a natural target promoter". Cell. 115 (6): 751–763. doi:10.1016/S0092-8674(03)00934-6. PMID 14675539. S2CID 145525.
- ^ Nagaich AK, Walker DA, Wolford R, Hager GL (April 2004). "Rapid periodic binding and displacement of the glucocorticoid receptor during chromatin remodeling". Molecular Cell. 14 (2): 163–174. doi:10.1016/S1097-2765(04)00178-9. PMID 15099516.
- ^ Teif VB, Vainshtein Y, Caudron-Herger M, Mallm JP, Marth C, Höfer T, Rippe K (November 2012). "Genome-wide nucleosome positioning during embryonic stem cell development". Nature Structural & Molecular Biology. 19 (11): 1185–1192. doi:10.1038/nsmb.2419. PMID 23085715. S2CID 34509771.
- ^ Albert I, Mavrich TN, Tomsho LP, Qi J, Zanton SJ, Schuster SC, Pugh BF (March 2007). "Translational and rotational settings of H2A.Z nucleosomes across the Saccharomyces cerevisiae genome". Nature. 446 (7135): 572–576. Bibcode:2007Natur.446..572A. doi:10.1038/nature05632. PMID 17392789. S2CID 4416890.
- ^ Li B, Carey M, Workman JL (February 2007). "The role of chromatin during transcription". Cell. 128 (4): 707–719. doi:10.1016/j.cell.2007.01.015. PMID 17320508. S2CID 1773333.
- ^ Whitehouse I, Rando OJ, Delrow J, Tsukiyama T (December 2007). "Chromatin remodelling at promoters suppresses antisense transcription". Nature. 450 (7172): 1031–1035. Bibcode:2007Natur.450.1031W. doi:10.1038/nature06391. PMID 18075583. S2CID 4305576.
- ^ Lee W, Tillo D, Bray N, Morse RH, Davis RW, Hughes TR, Nislow C (October 2007). "A high-resolution atlas of nucleosome occupancy in yeast". Nature Genetics. 39 (10): 1235–1244. doi:10.1038/ng2117. PMID 17873876. S2CID 12816925.
- ^ Eaton ML, Galani K, Kang S, Bell SP, MacAlpine DM (April 2010). "Conserved nucleosome positioning defines replication origins". Genes & Development. 24 (8): 748–753. doi:10.1101/gad.1913210. PMC 2854390. PMID 20351051.
- ^ Shivaswamy S, Bhinge A, Zhao Y, Jones S, Hirst M, Iyer VR (March 2008). "Dynamic remodeling of individual nucleosomes across a eukaryotic genome in response to transcriptional perturbation". PLOS Biology. 6 (3) e65. doi:10.1371/journal.pbio.0060065. PMC 2267817. PMID 18351804.
- ^ Murugesapillai D, McCauley MJ, Huo R, Nelson Holte MH, Stepanyants A, Maher LJ, et al. (August 2014). "DNA bridging and looping by HMO1 provides a mechanism for stabilizing nucleosome-free chromatin". Nucleic Acids Research. 42 (14): 8996–9004. doi:10.1093/nar/gku635. PMC 4132745. PMID 25063301.
- ^ Murugesapillai D, McCauley MJ, Maher LJ, Williams MC (February 2017). "Single-molecule studies of high-mobility group B architectural DNA bending proteins". Biophysical Reviews. 9 (1): 17–40. doi:10.1007/s12551-016-0236-4. PMC 5331113. PMID 28303166.
- ^ Winger J, Nodelman IM, Levendosky RF, Bowman GD (May 2018). "A twist defect mechanism for ATP-dependent translocation of nucleosomal DNA". eLife. 7 e34100. doi:10.7554/eLife.34100. PMC 6031429. PMID 29809147.
- ^ Bowman GD (August 2019). "Uncovering a New Step in Sliding Nucleosomes". Trends in Biochemical Sciences. 44 (8): 643–645. doi:10.1016/j.tibs.2019.05.001. PMC 7092708. PMID 31171402.
- ^ a b Nodelman IM, Bowman GD (May 2021). "Biophysics of Chromatin Remodeling". Annual Review of Biophysics. 50 (1): 73–93. doi:10.1146/annurev-biophys-082520-080201. PMC 8428145. PMID 33395550.
- ^ a b Brandani GB, Takada S (November 2018). Onufriev A (ed.). "Chromatin remodelers couple inchworm motion with twist-defect formation to slide nucleosomal DNA". PLOS Computational Biology. 14 (11) e1006512. Bibcode:2018PLSCB..14E6512B. doi:10.1371/journal.pcbi.1006512. PMC 6237416. PMID 30395604.
- ^ Hayes JJ, Lee KM (May 1997). "In vitro reconstitution and analysis of mononucleosomes containing defined DNAs and proteins". Methods. 12 (1): 2–9. doi:10.1006/meth.1997.0441. PMID 9169189.
- ^ Dyer PN, Edayathumangalam RS, White CL, Bao Y, Chakravarthy S, Muthurajan UM, Luger K (2004). "Reconstitution of nucleosome core particles from recombinant histones and DNA". Chromatin and Chromatin Remodeling Enzymes, Part A. Methods in Enzymology. Vol. 375. pp. 23–44. doi:10.1016/s0076-6879(03)75002-2. ISBN 978-0-12-182779-3. PMID 14870657.
- ^ Yenidunya A, Davey C, Clark D, Felsenfeld G, Allan J (April 1994). "Nucleosome positioning on chicken and human globin gene promoters in vitro. Novel mapping techniques". Journal of Molecular Biology. 237 (4): 401–414. doi:10.1006/jmbi.1994.1243. PMID 8151701.
- ^ Frouws TD, Barth PD, Richmond TJ (January 2018). "Site-Specific Disulfide Crosslinked Nucleosomes with Enhanced Stability". Journal of Molecular Biology. 430 (1): 45–57. doi:10.1016/j.jmb.2017.10.029. PMC 5757783. PMID 29113904.
- ^ Ferentz AE, Verdine GL (1994). "The Convertible Nucleoside Approach: Structural Engineering of Nucleic Acids by Disulfide Cross-Linking". In Eckstein F, Lilley DM (eds.). Nucleic Acids and Molecular Biology. Vol. 8. pp. 14–40. doi:10.1007/978-3-642-78666-2_2. ISBN 978-3-642-78668-6.
- ^ Yamasu K, Senshu T (January 1990). "Conservative segregation of tetrameric units of H3 and H4 histones during nucleosome replication". Journal of Biochemistry. 107 (1): 15–20. doi:10.1093/oxfordjournals.jbchem.a122999. PMID 2332416.
- ^ Kaufman PD, Kobayashi R, Kessler N, Stillman B (June 1995). "The p150 and p60 subunits of chromatin assembly factor I: a molecular link between newly synthesized histones and DNA replication". Cell. 81 (7): 1105–1114. doi:10.1016/S0092-8674(05)80015-7. PMID 7600578. S2CID 13502921.
- ^ Tyler JK, Adams CR, Chen SR, Kobayashi R, Kamakaka RT, Kadonaga JT (December 1999). "The RCAF complex mediates chromatin assembly during DNA replication and repair". Nature. 402 (6761): 555–560. Bibcode:1999Natur.402..555T. doi:10.1038/990147. PMID 10591219. S2CID 205097512.
- ^ Benson LJ, Gu Y, Yakovleva T, Tong K, Barrows C, Strack CL, et al. (April 2006). "Modifications of H3 and H4 during chromatin replication, nucleosome assembly, and histone exchange". The Journal of Biological Chemistry. 281 (14): 9287–9296. doi:10.1074/jbc.M512956200. PMID 16464854.
- ^ Louters L, Chalkley R (June 1985). "Exchange of histones H1, H2A, and H2B in vivo". Biochemistry. 24 (13): 3080–3085. doi:10.1021/bi00334a002. PMID 4027229.
- ^ Park YJ, Chodaparambil JV, Bao Y, McBryant SJ, Luger K (January 2005). "Nucleosome assembly protein 1 exchanges histone H2A-H2B dimers and assists nucleosome sliding". The Journal of Biological Chemistry. 280 (3): 1817–1825. doi:10.1074/jbc.M411347200. PMID 15516689.
- ^ Vincent JA, Kwong TJ, Tsukiyama T (May 2008). "ATP-dependent chromatin remodeling shapes the DNA replication landscape". Nature Structural & Molecular Biology. 15 (5): 477–484. doi:10.1038/nsmb.1419. PMC 2678716. PMID 18408730.
- ^ Yadav T, Whitehouse I (April 2016). "Replication-Coupled Nucleosome Assembly and Positioning by ATP-Dependent Chromatin-Remodeling Enzymes". Cell Reports. 15 (4): 715–723. doi:10.1016/J.CELREP.2016.03.059. PMC 5063657. PMID 27149855.
External links
[edit]- MBInfo - What are nucleosomes
- Nucleosomes on the Richmond Lab website
- Proteopedia Nucleosomes
- Nucleosome at the PDB
- Dynamic Remodeling of Individual Nucleosomes Across a Eukaryotic Genome in Response to Transcriptional Perturbation
- Nucleosome positioning data and tools online (annotated list, constantly updated)
- Histone protein structure
- HistoneDB 2.0 - Database of histones and variants at NCBI




