Nuclear envelope
View on Wikipedia| Nuclear envelope | |
|---|---|
 Human cell nucleus | |
| Identifiers | |
| TH | H1.00.01.2.01001 |
| FMA | 63888 |
| Anatomical terminology | |
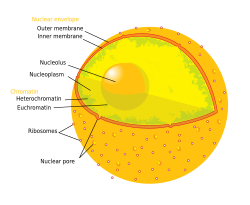
The nuclear envelope, also known as the nuclear membrane,[1][a] is made up of two lipid bilayer membranes that in eukaryotic cells surround the nucleus, which encloses the genetic material.
The nuclear envelope consists of two lipid bilayer membranes: an inner nuclear membrane and an outer nuclear membrane.[4] The space between the membranes is called the perinuclear space. It is usually about 10–50 nm wide.[5][6] The outer nuclear membrane is continuous with the endoplasmic reticulum membrane.[4] The nuclear envelope has many nuclear pores that allow materials to move between the cytosol and the nucleus.[4] Intermediate filament proteins called lamins form a structure called the nuclear lamina on the inner aspect of the inner nuclear membrane and give structural support to the nucleus.[4]
Structure
[edit]The nuclear envelope is made up of two lipid bilayer membranes, an inner nuclear membrane and an outer nuclear membrane. These membranes are connected to each other by nuclear pores. Two sets of intermediate filaments provide support for the nuclear envelope. An internal network forms the nuclear lamina on the inner nuclear membrane.[7] A looser network forms outside to give external support.[4] The actual shape of the nuclear envelope is irregular. It has invaginations and protrusions and can be observed with an electron microscope.

Outer membrane
[edit]The outer nuclear membrane also shares a common border with the endoplasmic reticulum.[8] While it is physically linked, the outer nuclear membrane contains proteins found in far higher concentrations than the endoplasmic reticulum.[9] All four nesprin proteins (nuclear envelope spectrin repeat proteins) present in mammals are expressed in the outer nuclear membrane.[10] Nesprin proteins connect cytoskeletal filaments to the nucleoskeleton.[11] Nesprin-mediated connections to the cytoskeleton contribute to nuclear positioning and to the cell's mechanosensory function.[12] KASH domain proteins of Nesprin-1 and -2 are part of a LINC complex (linker of nucleoskeleton and cytoskeleton) and can bind directly to cystoskeletal components, such as actin filaments, or can bind to proteins in the perinuclear space.[13][14] Nesprin-3 and -4 may play a role in unloading enormous cargo; Nesprin-3 proteins bind plectin and link the nuclear envelope to cytoplasmic intermediate filaments.[15] Nesprin-4 proteins bind the plus end directed motor kinesin-1.[16] The outer nuclear membrane is also involved in development, as it fuses with the inner nuclear membrane to form nuclear pores.[17]
Inner membrane
[edit]The inner nuclear membrane encloses the nucleoplasm, and is covered by the nuclear lamina, a mesh of intermediate filaments which stabilizes the nuclear membrane as well as being involved in chromatin function.[9] It is connected to the outer membrane by nuclear pores which penetrate the membranes. While the two membranes and the endoplasmic reticulum are linked, proteins embedded in the membranes tend to stay put rather than dispersing across the continuum.[18] It is lined with a fiber network called the nuclear lamina which is 10-40 nm thick and provides strength.[citation needed]
Mutations in the genes that encode for the inner nuclear membrane proteins can cause several laminopathies.[citation needed]
Nuclear pores
[edit]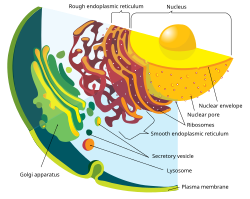
The nuclear envelope is punctured by around a thousand nuclear pore complexes, about 100 nm across, with an inner channel about 40 nm wide.[9] The complexes contain a number of nucleoporins, proteins that link the inner and outer nuclear membranes.[citation needed]
Cell division
[edit]During the G2 phase of interphase, the nuclear membrane increases its surface area and doubles its number of nuclear pore complexes.[9] In eukaryotes such as yeast which undergo closed mitosis, the nuclear membrane stays intact during cell division. The spindle fibers either form within the membrane, or penetrate it without tearing it apart.[9] In other eukaryotes (animals as well as plants), the nuclear membrane must break down during the prometaphase stage of mitosis to allow the mitotic spindle fibers to access the chromosomes inside. The breakdown and reformation processes are not well understood.
Breakdown
[edit]
In mammals, the nuclear membrane can break down within minutes, following a set of steps during the early stages of mitosis. First, M-Cdk's phosphorylate nucleoporin polypeptides and they are selectively removed from the nuclear pore complexes. After that, the rest of the nuclear pore complexes break apart simultaneously. Biochemical evidence suggests that the nuclear pore complexes disassemble into stable pieces rather than disintegrating into small polypeptide fragments.[9] M-Cdk's also phosphorylate elements of the nuclear lamina (the framework that supports the envelope) leading to the disassembly of the lamina and hence the envelope membranes into small vesicles.[19] Electron and fluorescence microscopy has given strong evidence that the nuclear membrane is absorbed by the endoplasmic reticulum—nuclear proteins not normally found in the endoplasmic reticulum show up during mitosis.[9]
In addition to the breakdown of the nuclear membrane during the prometaphase stage of mitosis, the nuclear membrane also ruptures in migrating mammalian cells during the interphase stage of the cell cycle.[20] This transient rupture is likely caused by nuclear deformation. The rupture is rapidly repaired by a process dependent on "endosomal sorting complexes required for transport" (ESCRT) made up of cytosolic protein complexes.[20] During nuclear membrane rupture events, DNA double-strand breaks occur. Thus the survival of cells migrating through confined environments appears to depend on efficient nuclear envelope and DNA repair machineries.
Aberrant nuclear envelope breakdown has also been observed in laminopathies and in cancer cells leading to mislocalization of cellular proteins, the formation of micronuclei and genomic instability.[21][22][23]
Reformation
[edit]Exactly how the nuclear membrane reforms during telophase of mitosis is debated. Two theories exist[9]—
- Vesicle fusion — where vesicles of nuclear membrane fuse together to rebuild the nuclear membrane
- Re-shaping of the endoplasmic reticulum—where the parts of the endoplasmic reticulum containing the absorbed nuclear membrane envelop the nuclear space, reforming a closed membrane.
Origin of the nuclear membrane
[edit]A study of the comparative genomics, evolution and origins of the nuclear membrane led to the proposal that the nucleus emerged in the primitive eukaryotic ancestor (the "prekaryote"), and was triggered by the archaeo-bacterial symbiosis.[24] Several ideas have been proposed for the evolutionary origin of the nuclear membrane.[25] These ideas include the invagination of the plasma membrane in a prokaryote ancestor, or the formation of a genuine new membrane system following the establishment of proto-mitochondria in the archaeal host. The adaptive function of the nuclear membrane may have been to serve as a barrier to protect the genome from reactive oxygen species (ROS) produced by the cells' pre-mitochondria.[26][27]
Notes
[edit]References
[edit]- ^ Georgia State University. "Cell Nucleus and Nuclear Envelope". gsu.edu. Archived from the original on 2018-06-18. Retrieved 2014-01-21.
- ^ "Nuclear membrane". Biology Dictionary. Biology Online. Retrieved 7 December 2012.
- ^ "nuclear membrane". Merriam Webster. Retrieved 7 December 2012.
- ^ a b c d e Alberts, Bruce (2002). Molecular biology of the cell (4th ed.). New York [u.a.]: Garland. p. 197. ISBN 978-0-8153-4072-0.
- ^ "Perinuclear space". Dictionary. Biology Online. Retrieved 7 December 2012.
- ^ Berrios, Miguel, ed. (1998). Nuclear structure and function. San Diego: Academic Press. p. 4. ISBN 978-0-12-564155-5.
- ^ Coutinho, Henrique Douglas M; Falcão-Silva, Vivyanne S; Gonçalves, Gregório Fernandes; da Nóbrega, Raphael Batista (20 April 2009). "Molecular ageing in progeroid syndromes: Hutchinson-Gilford progeria syndrome as a model". Immunity & Ageing. 6: 4. doi:10.1186/1742-4933-6-4. PMC 2674425. PMID 19379495.
- ^ "Chloride channels in the Nuclear membrane" (PDF). Harvard.edu. Archived from the original (PDF) on 2 August 2010. Retrieved 7 December 2012.
- ^ a b c d e f g h Hetzer, Mertin (February 3, 2010). "The Nuclear Envelope". Cold Spring Harbor Perspectives in Biology. 2 (3) a000539. doi:10.1101/cshperspect.a000539. PMC 2829960. PMID 20300205.
- ^ Wilson, Katherine L.; Berk, Jason M. (2010-06-15). "The nuclear envelope at a glance". J Cell Sci. 123 (12): 1973–1978. doi:10.1242/jcs.019042. ISSN 0021-9533. PMC 2880010. PMID 20519579.
- ^ Burke, Brian; Roux, Kyle J. (2009-11-01). "Nuclei take a position: managing nuclear location". Developmental Cell. 17 (5): 587–597. doi:10.1016/j.devcel.2009.10.018. ISSN 1878-1551. PMID 19922864.
- ^ Uzer, Gunes; Thompson, William R.; Sen, Buer; Xie, Zhihui; Yen, Sherwin S.; Miller, Sean; Bas, Guniz; Styner, Maya; Rubin, Clinton T. (2015-06-01). "Cell Mechanosensitivity to Extremely Low-Magnitude Signals Is Enabled by a LINCed Nucleus". Stem Cells. 33 (6): 2063–2076. doi:10.1002/stem.2004. ISSN 1066-5099. PMC 4458857. PMID 25787126.
- ^ Crisp, Melissa; Liu, Qian; Roux, Kyle; Rattner, J. B.; Shanahan, Catherine; Burke, Brian; Stahl, Phillip D.; Hodzic, Didier (2006-01-02). "Coupling of the nucleus and cytoplasm: role of the LINC complex". The Journal of Cell Biology. 172 (1): 41–53. doi:10.1083/jcb.200509124. ISSN 0021-9525. PMC 2063530. PMID 16380439.
- ^ Zeng, X; et al. (2018). "Nuclear Envelope-Associated Chromosome Dynamics during Meiotic Prophase I." Frontiers in Cell and Developmental Biology. 5: 121. doi:10.3389/fcell.2017.00121. PMC 5767173. PMID 29376050.
- ^ Wilhelmsen, Kevin; Litjens, Sandy H. M.; Kuikman, Ingrid; Tshimbalanga, Ntambua; Janssen, Hans; van den Bout, Iman; Raymond, Karine; Sonnenberg, Arnoud (2005-12-05). "Nesprin-3, a novel outer nuclear membrane protein, associates with the cytoskeletal linker protein plectin". The Journal of Cell Biology. 171 (5): 799–810. doi:10.1083/jcb.200506083. ISSN 0021-9525. PMC 2171291. PMID 16330710.
- ^ Roux, Kyle J.; Crisp, Melissa L.; Liu, Qian; Kim, Daein; Kozlov, Serguei; Stewart, Colin L.; Burke, Brian (2009-02-17). "Nesprin 4 is an outer nuclear membrane protein that can induce kinesin-mediated cell polarization". Proceedings of the National Academy of Sciences of the United States of America. 106 (7): 2194–2199. Bibcode:2009PNAS..106.2194R. doi:10.1073/pnas.0808602106. ISSN 1091-6490. PMC 2650131. PMID 19164528.
- ^ Fichtman, Boris; Ramos, Corinne; Rasala, Beth; Harel, Amnon; Forbes, Douglass J. (2010-12-01). "Inner/Outer Nuclear Membrane Fusion in Nuclear Pore Assembly". Molecular Biology of the Cell. 21 (23): 4197–4211. doi:10.1091/mbc.E10-04-0309. ISSN 1059-1524. PMC 2993748. PMID 20926687.
- ^ Georgatos, S. D. (April 19, 2001). "The inner nuclear membrane: simple, or very complex?". The EMBO Journal. 20 (12): 2989–2994. doi:10.1093/emboj/20.12.2989. PMC 150211. PMID 11406575.
- ^ Alberts; et al. (2008). "Chapter 17: The Cell Cycle". Molecular Biology of The Cell (5th ed.). New York: Garland Science. pp. 1079–1080. ISBN 978-0-8153-4106-2.
- ^ a b Raab M, Gentili M, de Belly H, Thiam HR, Vargas P, Jimenez AJ, Lautenschlaeger F, Voituriez R, Lennon-Duménil AM, Manel N, Piel M (2016). "ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death". Science. 352 (6283): 359–62. Bibcode:2016Sci...352..359R. doi:10.1126/science.aad7611. PMID 27013426. S2CID 28544308.
- ^ Vargas; et al. (2012). "Transient nuclear envelope rupturing during interphase in human cancer cells". Nucleus (Austin, Tex.). 3 (1). Nucleus: 88–100. doi:10.4161/nucl.18954. PMC 3342953. PMID 22567193.
- ^ Lim; et al. (2016). "Nuclear envelope rupture drives genome instability in cancer". Molecular Biology of the Cell. 27 (21). MBoC: 3210–3213. doi:10.1091/mbc.E16-02-0098. PMC 5170854. PMID 27799497.
- ^ Hatch; et al. (2016). "Nuclear envelope rupture is induced by actin-based nucleus confinement". Journal of Cell Biology. 215 (1). JCB: 27–36. doi:10.1083/jcb.201603053. PMC 5057282. PMID 27697922. Retrieved 24 March 2019.
- ^ Mans BJ, Anantharaman V, Aravind L, Koonin EV (2004). "Comparative genomics, evolution and origins of the nuclear envelope and nuclear pore complex". Cell Cycle. 3 (12): 1612–37. doi:10.4161/cc.3.12.1316. PMID 15611647.
- ^ Martin W (2005). "Archaebacteria (Archaea) and the origin of the eukaryotic nucleus". Curr. Opin. Microbiol. 8 (6): 630–7. doi:10.1016/j.mib.2005.10.004. PMID 16242992.
- ^ Speijer D (2015). "Birth of the eukaryotes by a set of reactive innovations: New insights force us to relinquish gradual models". BioEssays. 37 (12): 1268–76. doi:10.1002/bies.201500107. PMID 26577075. S2CID 20068849.
- ^ Bernstein H, Bernstein C. Sexual communication in archaea, the precursor to meiosis. pp. 103-117 in Biocommunication of Archaea (Guenther Witzany, ed.) 2017. Springer International Publishing ISBN 978-3-319-65535-2 DOI 10.1007/978-3-319-65536-9
External links
[edit]- Histology image: 20102loa – Histology Learning System at Boston University
- Animations of nuclear pores and transport through the nuclear envelope Archived 2009-02-07 at the Wayback Machine
- Illustrations of nuclear pores and transport through the nuclear membrane Archived 2009-02-07 at the Wayback Machine
- Nuclear+membrane at the U.S. National Library of Medicine Medical Subject Headings (MeSH)