Recent from talks
Nothing was collected or created yet.
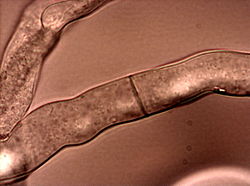
Neurospora crassa
View on Wikipedia
| Neurospora crassa | |
|---|---|
| |
| Scientific classification | |
| Kingdom: | Fungi |
| Division: | Ascomycota |
| Class: | Sordariomycetes |
| Order: | Sordariales |
| Family: | Sordariaceae |
| Genus: | Neurospora |
| Species: | N. crassa
|
| Binomial name | |
| Neurospora crassa Shear & B.O. Dodge
| |
Neurospora crassa is a type of red bread mold of the phylum Ascomycota. The genus name, meaning 'nerve spore' in Greek, refers to the characteristic striations on the spores. The first published account of this fungus was from an infestation of French bakeries in 1843.[1]
Neurospora crassa is used as a model organism because it is easy to grow and has a haploid life cycle that makes genetic analysis simple since recessive traits will show up in the offspring. Analysis of genetic recombination is facilitated by the ordered arrangement of the products of meiosis in Neurospora ascospores. Its entire genome of seven chromosomes has been sequenced.[2]
Neurospora was used by Edward Tatum and George Wells Beadle in their experiments for which they won the Nobel Prize in Physiology or Medicine in 1958. Beadle and Tatum exposed N. crassa to x-rays, causing mutations. They then observed failures in metabolic pathways caused by errors in specific enzymes. This led them to propose the "one gene, one enzyme" hypothesis that specific genes code for specific proteins. Their hypothesis was later elaborated to enzyme pathways by Norman Horowitz, also working on Neurospora. As Norman Horowitz reminisced in 2004,[3] "These experiments founded the science of what Beadle and Tatum called 'biochemical genetics'. In actuality, they proved to be the opening gun in what became molecular genetics and all developments that have followed from that."
In the 24 April 2003 issue of Nature, the genome of N. crassa was reported as completely sequenced.[4] The genome is about 43 megabases long and includes approximately 10,000 genes. There is a project underway to produce strains containing knockout mutants of every N. crassa gene.[5]
In its natural environment, N. crassa lives mainly in tropical and sub-tropical regions.[6] It can be found growing on dead plant matter after fires.
Neurospora is actively used in research around the world. It is important in the elucidation of molecular events involved in circadian rhythms, epigenetics and gene silencing, cell polarity, cell fusion, development, as well as many aspects of cell biology and biochemistry.
The sexual cycle
[edit]
Sexual fruiting bodies (perithecia) can only be formed when two mycelia of different mating type come together (see Figure). Like other Ascomycetes, N. crassa has two mating types that, in this case, are symbolized by A and a. There is no evident morphological difference between the A and a mating type strains. Both can form abundant protoperithecia, the female reproductive structure (see Figure). Protoperithecia are formed most readily in the laboratory when growth occurs on solid (agar) synthetic medium with a relatively low source of nitrogen.[7] Nitrogen starvation appears to be necessary for expression of genes involved in sexual development.[8] The protoperithecium consists of an ascogonium, a coiled multicellular hypha that is enclosed in a knot-like aggregation of hyphae. A branched system of slender hyphae, called the trichogyne, extends from the tip of the ascogonium projecting beyond the sheathing hyphae into the air. The sexual cycle is initiated (i.e. fertilization occurs) when a cell (usually a conidium) of opposite mating type contacts a part of the trichogyne (see Figure). Such contact can be followed by cell fusion leading to one or more nuclei from the fertilizing cell migrating down the trichogyne into the ascogonium. Since both A and a strains have the same sexual structures, neither strain can be regarded as exclusively male or female. However, as a recipient, the protoperithecium of both the A and a strains can be thought of as the female structure, and the fertilizing conidium can be thought of as the male participant.[citation needed]
The subsequent steps following fusion of A and a haploid cells, have been outlined by Fincham and Day[9] and Wagner and Mitchell.[10] After fusion of the cells, the further fusion of their nuclei is delayed. Instead, a nucleus from the fertilizing cell and a nucleus from the ascogonium become associated and begin to divide synchronously. The products of these nuclear divisions (still in pairs of unlike mating type, i.e. A/a) migrate into numerous ascogenous hyphae, which then begin to grow out of the ascogonium. Each of these ascogenous hypha bends to form a hook (or crozier) at its tip and the A and a pair of haploid nuclei within the crozier divide synchronously. Next, septa form to divide the crozier into three cells. The central cell in the curve of the hook contains one A and one a nucleus (see Figure). This binuclear cell initiates ascus formation and is called an "ascus-initial" cell. Next the two uninucleate cells on either side of the first ascus-forming cell fuse with each other to form a binucleate cell that can grow to form a further crozier that can then form its own ascus-initial cell. This process can then be repeated multiple times.
After formation of the ascus-initial cell, the A and a nucleus fuse with each other to form a diploid nucleus (see Figure). This nucleus is the only diploid nucleus in the entire life cycle of N. crassa. The diploid nucleus has 14 chromosomes formed from the two fused haploid nuclei that had 7 chromosomes each. Formation of the diploid nucleus is immediately followed by meiosis. The two sequential divisions of meiosis lead to four haploid nuclei, two of the A mating type and two of the a mating type. One further mitotic division leads to four A and four a nucleus in each ascus. Meiosis is an essential part of the life cycle of all sexually reproducing organisms, and in its main features, meiosis in N. crassa seems typical of meiosis generally.[citation needed]
As the above events are occurring, the mycelial sheath that had enveloped the ascogonium develops as the wall of the perithecium, becomes impregnated with melanin, and blackens. The mature perithecium has a flask-shaped structure.[citation needed]
A mature perithecium may contain as many as 300 asci, each derived from identical fusion diploid nuclei. Ordinarily, in nature, when the perithecia mature the ascospores are ejected rather violently into the air. These ascospores are heat resistant and, in the lab, require heating at 60 °C for 30 minutes to induce germination. For normal strains, the entire sexual cycle takes 10 to 15 days. In a mature ascus containing eight ascospores, pairs of adjacent spores are identical in genetic constitution, since the last division is mitotic, and since the ascospores are contained in the ascus sac that holds them in a definite order determined by the direction of nuclear segregations during meiosis. Since the four primary products are also arranged in sequence, a first division segregation pattern of genetic markers can be distinguished from a second division segregation pattern.[citation needed]
Fine structure genetic analysis
[edit]Because of the above features N. crassa was found to be very useful for the study of genetic events occurring in individual meioses. Mature asci from a perithecium can be separated on a microscope slide and the spores experimentally manipulated. These studies usually involved the separate culture of individual ascospores resulting from a single meiotic event and determining the genotype of each spore. Studies of this type, carried out in several different laboratories, established the phenomenon of "gene conversion" (e.g. see references[11][12][13]).
As an example of the gene conversion phenomenon, consider genetic crosses of two N. crassa mutant strains defective in gene pan-2. This gene is necessary for the synthesis of pantothenic acid (vitamin B5), and mutants defective in this gene can be experimentally identified by their requirement for pantothenic acid in their growth medium. The two pan-2 mutations B5 and B3 are located at different sites in the pan-2 gene, so that a cross of B5 ´ B3 yields wild-type recombinants at low frequency.[12] An analysis of 939 asci in which the genotypes of all meiotic products (ascospores) could be determined found 11 asci with an exceptional segregation pattern. These included six asci in which there was one wild-type meiotic product but no expected reciprocal double-mutant (B5B3) product. Furthermore, in three asci the ratio of meiotic products was 1B5:3B3, rather than in the expected 2:2 ratio. This study, as well as numerous additional studies in N. crassa and other fungi (reviewed by Whitehouse[14]), led to an extensive characterization of gene conversion. It became clear from this work that gene conversion events arise when a molecular recombination event happens to occur near the genetic markers under study (e.g. pan-2 mutations in the above example). Thus studies of gene conversion allowed insight into the details of the molecular mechanism of recombination. Over the decades since the original observations of Mary Mitchell in 1955,[11] a sequence of molecular models of recombination have been proposed based on both emerging genetic data from gene conversion studies and studies of the reaction capabilities of DNA. Current understanding of the molecular mechanism of recombination is discussed in the Wikipedia articles Gene conversion and Genetic recombination. An understanding of recombination is relevant to several fundamental biologic problems, such the role of recombination and recombinational repair in cancer (see BRCA1) and the adaptive function of meiosis.
Adaptive function of mating type
[edit]That mating in N. crassa can only occur between strains of different mating types suggests that some degree of outcrossing is favored by natural selection. In haploid multicellular fungi, such as N. crassa, meiosis occurring in the brief diploid stage is one of their most complex processes. Although physically much larger than the diploid stage, the haploid multicellular vegetative stage characteristically has a simple modular construction with little differentiation. In N. crassa, recessive mutations affecting the diploid stage of the life cycle are quite frequent in natural populations.[15] These mutations, when homozygous in the diploid stage, often cause spores to have maturation defects or to produce barren fruiting bodies with few ascospores (sexual spores). Most of these homozygous mutations cause abnormal meiosis (e.g., disturbed chromosome pairing or pachytene or diplotene).[16] The number of genes affecting the diploid stage was estimated to be at least 435[15] (about 4% of the total number of 9,730 genes). Thus, outcrossing, promoted by the necessity for the union of opposite mating types, likely provides the benefit of masking recessive mutations that would otherwise be harmful to sexual spore formation (see Complementation (genetics)).
Current research
[edit]Neurospora crassa is not only a model organism for the study of phenotypic types in knock-out variants, but a particularly useful organism widely used in computational biology and the circadian clock. It has a natural reproductive cycle of 22 hours and is influenced by external factors such as light and temperature. Knock out variants of wild type N. crassa are widely studied to determine the influence of particular genes (see Frequency (gene)).
Use in Food
[edit]A meat substitute made from this species is sold under the brand name Meati.[17]
See also
[edit]- Charles Yanofsky – American geneticist (1925–2018)
- David Perkins – American geneticist
- Edward Tatum – American geneticist
- Fluffy transcription factor – Gene of Neurospora crassa required for asexual sporulation
- George Beadle – American geneticist (1903–1989)
- Norman Horowitz – American geneticist
- One gene–one enzyme hypothesis – Theory in genetics
- Robert Metzenberg – American geneticist
Notes and references
[edit]- ^ Davis, Perkins (2002). "Neurospora: a model of model microbes". Nature Reviews Genetics. 3 (5): 397–403. doi:10.1038/nrg797. PMID 11988765. S2CID 15642417.
- ^ Trans-NIH Neurospora Initiative
- ^ Horowitz NH, Berg P, Singer M, et al. (January 2004). "A centennial: George W. Beadle, 1903–1989". Genetics. 166 (1): 1–10. doi:10.1534/genetics.166.1.1. PMC 1470705. PMID 15020400.
- ^ Galagan J.; Calvo S.; Borkovich K.; Selker E.; Read N. D.; et al. (2003). "The genome sequence of the filamentous fungus Neurospora crassa". Nature. 422 (6934): 859–868. Bibcode:2003Natur.422..859G. doi:10.1038/nature01554. PMID 12712197.
- ^ Colot H.V.; Park G.; Turner G.E.; Ringleberg C.; Crew C.M.; Litvinkova L.; Weiss R.L.; Borkovitch K.A.; Dunlap J.C.; et al. (2006). "A high-throughput gene knockout procedure for Neurospora reveals functions for multiple transcription factors". Proceedings of the National Academy of Sciences, USA. 103 (27): 10352–10357. Bibcode:2006PNAS..10310352C. doi:10.1073/pnas.0601456103. PMC 1482798. PMID 16801547.
- ^ Perkins D. D.; Turner B. C. (1988). "Neurospora from natural populations: Toward the population biology of a haploid eukaryote". Experimental Mycology. 12 (2): 91–131. doi:10.1016/0147-5975(88)90001-1.
- ^ Westergaard M, Mitchell HK (1947). "Neurospora. V. "A synthetic medium favoring sexual reproduction". Am J Bot. 34 (10): 573–577. doi:10.2307/2437339. JSTOR 2437339.
- ^ Nelson MA, Metzenberg RL (September 1992). "Sexual development genes of Neurospora crassa". Genetics. 132 (1): 149–62. doi:10.1093/genetics/132.1.149. PMC 1205113. PMID 1356883.
- ^ Fincham, John Robert Stanley (1965). Fungal genetics. Oxford, Blackwell Scientific Publications.
- ^ Wagner, Robert P.; Mitchell, Herschel K. (1964). Genetics and metabolism. New York, J. Wiley.
- ^ a b Mitchell MB (April 1955). "ABERRANT RECOMBINATION OF PYRIDOXINE MUTANTS OF Neurospora". Proc. Natl. Acad. Sci. U.S.A. 41 (4): 215–20. Bibcode:1955PNAS...41..215M. doi:10.1073/pnas.41.4.215. PMC 528059. PMID 16589648.
- ^ a b Case ME, Giles NH (May 1958). "EVIDENCE FROM TETRAD ANALYSIS FOR BOTH NORMAL AND ABERRANT RECOMBINATION BETWEEN ALLELIC MUTANTS IN Neurospora Crassa". Proc. Natl. Acad. Sci. U.S.A. 44 (5): 378–90. Bibcode:1958PNAS...44..378C. doi:10.1073/pnas.44.5.378. PMC 335434. PMID 16590210.
- ^ Stadler DR (July 1959). "Gene Conversion of Cysteine Mutants in Neurospora". Genetics. 44 (4): 647–56. doi:10.1093/genetics/44.4.647. PMC 1209971. PMID 17247847.
- ^ Whitehouse, HLK. (1982). Genetic Recombination. Wiley, New York ISBN 978-0471102052
- ^ a b Leslie JF, Raju NB (December 1985). "Recessive mutations from natural populations of Neurospora crassa that are expressed in the sexual diplophase". Genetics. 111 (4): 759–77. doi:10.1093/genetics/111.4.759. PMC 1202670. PMID 2933298.
- ^ Raju NB, Leslie JF (October 1992). "Cytology of recessive sexual-phase mutants from wild strains of Neurospora crassa". Genome. 35 (5): 815–26. doi:10.1139/g92-124. PMID 1427061.
- ^ "Mycelium 101: What is Mycelium?".
References
[edit]- Perkins, D; Davis, R (December 2000), "Evidence for Safety of Neurospora Species for Academic and Commercial Uses", Applied and Environmental Microbiology, vol. 66, no. 12, pp. 5107–5109, Bibcode:2000ApEnM..66.5107P, doi:10.1128/aem.66.12.5107-5109.2000, PMC 92429, PMID 11097875.
- Osherov, N; May, GS (30 May 2001), "The molecular mechanisms of conidial germination", FEMS Microbiol Lett, vol. 199, no. 2, pp. 153–60, doi:10.1111/j.1574-6968.2001.tb10667.x, PMID 11377860
- Froehlich, AC; Noh, B; Vierstra, RD, Loros J & Dunlap JC (December 2005), "Genetic and molecular analysis of Phytochromes from the filamentous fungus Neurospora crassa", Eukaryot Cell, vol. 4, no. 12, pp. 2140–52, doi:10.1128/ec.4.12.2140-2152.2005, PMC 1317490, PMID 16339731
{{citation}}: CS1 maint: multiple names: authors list (link) - Horowitz, NH (April 1991), "Fifty years ago: the Neurospora revolution", Genetics, vol. 127, no. 4, pp. 631–5, doi:10.1093/genetics/127.4.631, PMC 1204391, PMID 1827628
- Horowitz, NH; Berg, P; Singer, M, Lederberg J, Susman M, Doebley J & Crow JF. (January 2004), "A centennial: George W. Beadle, 1903-1989", Genetics, vol. 166, no. 1, pp. 1–10, doi:10.1534/genetics.166.1.1, PMC 1470705, PMID 15020400
{{citation}}: CS1 maint: multiple names: authors list (link) - Kaldi, K; Gonzalez, BH; Brunner, M (23 December 2005), "Transcriptional regulation of the Neurospora circadian clock gene wc-1 affects the phase of circadian output", EMBO Rep, vol. 7, no. 2, pp. 199–204, doi:10.1038/sj.embor.7400595, PMC 1369249, PMID 16374510
- Pittalwala, Iqbal (29 April 2003), "UC Riverside scientists contribute to study that unveils genome sequence of bread mold", Newsroom (University of California, Riverside), archived from the original on 18 May 2008, retrieved 28 December 2005.
- Ruoff, P; Loros, JJ; Dunlap, JC (6 December 2005), "The relationship between FRQ-protein stability and temperature compensation in the Neurospora circadian clock", Proc Natl Acad Sci USA, vol. 102, no. 49, pp. 17681–6, Bibcode:2005PNAS..10217681R, doi:10.1073/pnas.0505137102, PMC 1308891, PMID 16314576
External links
[edit]- Neurospora crassa genome
- "The Neurospora Homepage". Fungal Genetics Stock Center (FGSC). Retrieved 27 December 2005.
- "The Neurospora Compendium". Fungal Genetics Stock Center (FGSC). Retrieved 27 December 2005.
- "The Neurospora-Fungal Genome Initiative". Neurospora Genome Project. Archived from the original on 4 March 2016. Retrieved 12 June 2015.
- "Trans-NIH Neurospora Initiative". National Institutes of Health (NIH — United States). Archived from the original on 28 October 2005. Retrieved 27 December 2005.
- [1] Montenegro-Montero A. (2010) "The Almighty Fungi: The Revolutionary Neurospora crassa". A historical view of the many contributions of this organism to molecular biology.

