Recent from talks
Nothing was collected or created yet.
Coenzyme Q – cytochrome c reductase
View on Wikipedia| Cytochrome b-c1 complex | |
|---|---|
 Crystal structure of mitochondrial cytochrome bc complex bound with ubiquinone.[1] | |
| Identifiers | |
| Symbol | (N/A) |
| SCOP2 | 1be3 / SCOPe / SUPFAM |
| TCDB | 3.D.3 |
| OPM superfamily | 92 |
| OPM protein | 3cx5 |
| Membranome | 258 |
| ubiquinol—cytochrome-c reductase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 7.1.1.8 | ||||||||
| CAS no. | 9027-03-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
The coenzyme Q : cytochrome c – oxidoreductase, sometimes called the cytochrome bc1 complex, and at other times complex III, is the third complex in the electron transport chain (EC 1.10.2.2), playing a critical role in biochemical generation of ATP (oxidative phosphorylation). Complex III is a multisubunit transmembrane protein encoded by both the mitochondrial (cytochrome b) and the nuclear genomes (all other subunits). Complex III is present in the mitochondria of all animals and all aerobic eukaryotes and the inner membranes of most bacteria. Mutations in Complex III cause exercise intolerance as well as multisystem disorders. The bc1 complex contains 11 subunits: 3 respiratory subunits (cytochrome b, cytochrome c1, Rieske protein), 2 core proteins, and 6 low-molecular-weight proteins.
Ubiquinol—cytochrome-c reductase catalyzes the chemical reaction
- QH2 + 2 ferricytochrome c Q + 2 ferrocytochrome c + 2 H+
Thus, the two substrates of this enzyme are quinol (QH2) and ferri- (Fe3+) cytochrome c, whereas its 3 products are quinone (Q), ferro- (Fe2+) cytochrome c, and H+.
This enzyme belongs to the family of oxidoreductases, specifically those acting on diphenols and related substances as donor with a cytochrome as acceptor. This enzyme participates in oxidative phosphorylation. It has four cofactors:[clarification needed] cytochrome c1, cytochrome b-562, cytochrome b-566, and a 2-Iron ferredoxin of the Rieske type.
Nomenclature
[edit]The systematic name of this enzyme class is ubiquinol:ferricytochrome-c oxidoreductase. Other names in common use include:
|
|
Structure
[edit]
Compared to the other major proton-pumping subunits of the electron transport chain, the number of subunits found can be small, as small as three polypeptide chains. This number does increase, and eleven subunits are found in higher animals.[2] Three subunits have prosthetic groups. The cytochrome b subunit has two b-type hemes (bL and bH), the cytochrome c subunit has one c-type heme (c1), and the Rieske Iron Sulfur Protein subunit (ISP) has a two iron, two sulfur iron-sulfur cluster (2Fe•2S).
Composition of complex
[edit]In vertebrates the bc1 complex, or Complex III, contains 11 subunits: 3 respiratory subunits, 2 core proteins and 6 low-molecular weight proteins.[3][4] Proteobacterial complexes may contain as few as three subunits.[5]
Table of subunit composition of complex III
[edit]| No. | Subunit name | Human gene symbol | Protein description from UniProt | Pfam family with Human protein |
|---|---|---|---|---|
| Respiratory subunit proteins | ||||
| 1 | MT-CYB / Cyt b | MT-CYB | Cytochrome b | Pfam PF13631 |
| 2 | CYC1 / Cyt c1 | CYC1 | Cytochrome c1, heme protein, mitochondrial | Pfam PF02167 |
| 3 | Rieske / UCR1 | UQCRFS1 | Cytochrome b-c1 complex subunit Rieske, mitochondrial EC 1.10.2.2 | Pfam PF02921 , Pfam PF00355 |
| Core protein subunits | ||||
| 4 | QCR1 / SU1 | UQCRC1 | Cytochrome b-c1 complex subunit 1, mitochondrial | Pfam PF00675, Pfam PF05193 |
| 5 | QCR2 / SU2 | UQCRC2 | Cytochrome b-c1 complex subunit 2, mitochondrial | Pfam PF00675, Pfam PF05193 |
| Low-molecular weight protein subunits | ||||
| 6 | QCR6 / SU6 | UQCRH | Cytochrome b-c1 complex subunit 6, mitochondrial | Pfam PF02320 |
| 7 | QCR7 / SU7 | UQCRB | Cytochrome b-c1 complex subunit 7 | Pfam PF02271 |
| 8 | QCR8 / SU8 | UQCRQ | Cytochrome b-c1 complex subunit 8 | Pfam PF02939 |
| 9 | QCR9 / SU9 | UQCRFS1a | (N-terminal of Rieske, no separate entry) | Pfam PF09165 |
| 10 | QCR10 / SU10 | UQCR10 | Cytochrome b-c1 complex subunit 9 | Pfam PF05365 |
| 11 | QCR11 / SU11 | UQCR11 | Cytochrome b-c1 complex subunit 10 | Pfam PF08997 |
- a In vertebrates, a cleavage product of 8 kDa from the N-terminus of the Rieske protein (Signal peptide) is retained in the complex as subunit 9. Thus subunits 10 and 11 correspond to fungal QCR9p and QCR10p.
Reaction
[edit]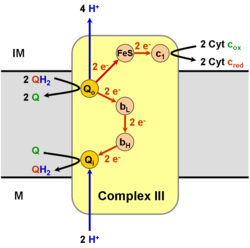
It catalyzes the reduction of cytochrome c by oxidation of coenzyme Q (CoQ) and the concomitant pumping of 4 protons from the mitochondrial matrix to the intermembrane space:
- QH2 + 2 cytochrome c (FeIII) + 2 H+
in → Q + 2 cytochrome c (FeII) + 4 H+
out
In the process called Q cycle,[6][7] two protons are consumed from the matrix (M), four protons are released into the inter membrane space (IM) and two electrons are passed to cytochrome c.
Reaction mechanism
[edit]
The reaction mechanism for complex III (cytochrome bc1, coenzyme Q: cytochrome C oxidoreductase) is known as the ubiquinone ("Q") cycle. In this cycle four protons get released into the positive "P" side (inter membrane space), but only two protons get taken up from the negative "N" side (matrix). As a result, a proton gradient is formed across the membrane. In the overall reaction, two ubiquinols are oxidized to ubiquinones and one ubiquinone is reduced to ubiquinol. In the complete mechanism, two electrons are transferred from ubiquinol to ubiquinone, via two cytochrome c intermediates.
Overall:
- 2 x QH2 oxidised to Q
- 1 x Q reduced to QH2
- 2 x Cyt c reduced
- 4 x H+ released into intermembrane space
- 2 x H+ picked up from matrix
The reaction proceeds according to the following steps:
Round 1:
- Cytochrome b binds a ubiquinol and a ubiquinone.
- The 2Fe/2S center and BL heme each pull an electron off the bound ubiquinol, releasing two protons into the intermembrane space.
- One electron is transferred to cytochrome c1 from the 2Fe/2S centre, whilst another is transferred from the BL heme to the BH Heme.
- Cytochrome c1 transfers its electron to cytochrome c (not to be confused with cytochrome c1), and the BH Heme transfers its electron to a nearby ubiquinone, resulting in the formation of a ubisemiquinone.
- Cytochrome c diffuses. The first ubiquinol (now oxidised to ubiquinone) is released, whilst the semiquinone remains bound.
Round 2:
- A second ubiquinol is bound by cytochrome b.
- The 2Fe/2S center and BL heme each pull an electron off the bound ubiquinol, releasing two protons into the intermembrane space.
- One electron is transferred to cytochrome c1 from the 2Fe/2S centre, whilst another is transferred from the BL heme to the BH Heme.
- Cytochrome c1 then transfers its electron to cytochrome c, whilst the nearby semiquinone produced from round 1 picks up a second electron from the BH heme, along with two protons from the matrix.
- The second ubiquinol (now oxidised to ubiquinone), along with the newly formed ubiquinol are released.[8]
Inhibitors of complex III
[edit]There are three distinct groups of Complex III inhibitors.
- Antimycin A binds to the Qi site and inhibits the transfer of electrons in Complex III from heme bH to oxidized Q (Qi site inhibitor).
- Myxothiazol and stigmatellin binds to the Qo site and inhibits the transfer of electrons from reduced QH2 to the Rieske Iron sulfur protein. Myxothiazol and stigmatellin bind to distinct but overlapping pockets within the Qo site.
- Myxothiazol binds nearer to cytochrome bL (hence termed a "proximal" inhibitor).
- Stigmatellin binds farther from heme bL and nearer the Rieske Iron sulfur protein, with which it strongly interacts.
Some have been commercialized as fungicides (the strobilurin derivatives, best known of which is azoxystrobin; QoI inhibitors) and as anti-malaria agents (atovaquone). Some Qo site inhibitors have been commercialized as insecticides (IRAC group 20).[9]
Also propylhexedrine inhibits cytochrome c reductase.[10]
Oxygen free radicals
[edit]A small fraction of electrons leave the electron transport chain before reaching complex IV. Premature electron leakage to oxygen results in the formation of superoxide. The relevance of this otherwise minor side reaction is that superoxide and other reactive oxygen species are highly toxic and are thought to play a role in several pathologies, as well as aging (the free radical theory of aging).[11] Electron leakage occurs mainly at the Qo site and is stimulated by antimycin A. Antimycin A locks the b hemes in the reduced state by preventing their re-oxidation at the Qi site, which, in turn, causes the steady-state concentrations of the Qo semiquinone to rise, the latter species reacting with oxygen to form superoxide. The effect of high membrane potential is thought to have a similar effect.[12] Superoxide produced at the Qo site can be released both into the mitochondrial matrix[13][14] and into the intermembrane space, where it can then reach the cytosol.[13][15] This could be explained by the fact that Complex III might produce superoxide as membrane permeable HOO• rather than as membrane impermeable O−.
2.[14]
Human gene names
[edit]This section may require cleanup to meet Wikipedia's quality standards. The specific problem is: Should get merged into table above. (December 2023) |
- MT-CYB: mtDNA encoded cytochrome b; mutations associated with exercise intolerance
- CYC1: cytochrome c1
- CYCS: cytochrome c
- UQCRFS1: Rieske iron sulfur protein
- UQCRB: Ubiquinone binding protein, mutation linked with mitochondrial complex III deficiency nuclear type 3
- UQCRH: hinge protein
- UQCRC2: Core 2, mutations linked to mitochondrial complex III deficiency, nuclear type 5
- UQCRC1: Core 1
- UQCR: 6.4KD subunit
- UQCR10: 7.2KD subunit
- TTC19: Newly identified subunit, mutations linked to complex III deficiency nuclear type 2. Helps remove the N-terminal fragment of UQCRFS1, which would otherwise interfere with complex III function.[16]
Mutations in complex III genes in human disease
[edit]Mutations in complex III-related genes typically manifest as exercise intolerance.[17][18] Other mutations have been reported to cause septo-optic dysplasia[19] and multisystem disorders.[20] However, mutations in BCS1L, a gene responsible for proper maturation of complex III, can result in Björnstad syndrome and the GRACILE syndrome, which in neonates are lethal conditions that have multisystem and neurologic manifestations typifying severe mitochondrial disorders. The pathogenicity of several mutations has been verified in model systems such as yeast.[21]
The extent to which these various pathologies are due to bioenergetic deficits or overproduction of superoxide is presently unknown.
See also
[edit]Additional images
[edit]References
[edit]- ^ PDB: 1ntz; Gao X, Wen X, Esser L, Quinn B, Yu L, Yu CA, Xia D (August 2003). "Structural basis for the quinone reduction in the bc1 complex: a comparative analysis of crystal structures of mitochondrial cytochrome bc1 with bound substrate and inhibitors at the Qi site". Biochemistry. 42 (30): 9067–80. doi:10.1021/bi0341814. PMID 12885240.
- ^ Iwata S, Lee JW, Okada K, Lee JK, Iwata M, Rasmussen B, Link TA, Ramaswamy S, Jap BK (July 1998). "Complete structure of the 11-subunit bovine mitochondrial cytochrome bc1 complex". Science. 281 (5373): 64–71. Bibcode:1998Sci...281...64I. doi:10.1126/science.281.5373.64. PMID 9651245.
- ^ Zhang Z, Huang L, Shulmeister VM, Chi YI, Kim KK, Hung LW, et al. (1998). "Electron transfer by domain movement in cytochrome bc1". Nature. 392 (6677): 677–84. Bibcode:1998Natur.392..677Z. doi:10.1038/33612. PMID 9565029. S2CID 4380033.
- ^ Hao GF, Wang F, Li H, Zhu XL, Yang WC, Huang LS, et al. (2012). "Computational discovery of picomolar Q(o) site inhibitors of cytochrome bc1 complex". J Am Chem Soc. 134 (27): 11168–76. Bibcode:2012JAChS.13411168H. doi:10.1021/ja3001908. PMID 22690928.
- ^ Yang XH, Trumpower BL (1986). "Purification of a three-subunit ubiquinol-cytochrome c oxidoreductase complex from Paracoccus denitrificans". J Biol Chem. 261 (26): 12282–9. doi:10.1016/S0021-9258(18)67236-9. PMID 3017970.
- ^ Kramer DM, Roberts AG, Muller F, Cape J, Bowman MK (2004). "Q-Cycle Bypass Reactions at the Qo Site of the Cytochrome bc1 (And Related) Complexes". Quinones and Quinone Enzymes, Part B. Methods in Enzymology. Vol. 382. pp. 21–45. doi:10.1016/S0076-6879(04)82002-0. ISBN 978-0-12-182786-1. PMID 15047094.
- ^ Crofts AR (2004). "The cytochrome bc1 complex: function in the context of structure". Annu. Rev. Physiol. 66: 689–733. doi:10.1146/annurev.physiol.66.032102.150251. PMID 14977419.
- ^ Ferguson SJ, Nicholls D, Ferguson S (2002). Bioenergetics (3rd ed.). San Diego: Academic. pp. 114–117. ISBN 978-0-12-518121-1.
- ^ Jeschke, Peter; Witschel, Matthias; Krämer, Wolfgang; Schirmer, Ulrich (25 January 2019). "32.3 Inhibitors of Mitochondrial Electron Transport: Acaricides and Insecticides". Modern Crop Protection Compounds (3rd ed.). Wiley-VCH. pp. 1156–1201. doi:10.1002/9783527699261. ISBN 9783527699261.
- ^ Holmes, J. H.; Sapeika, N; Zwarenstein, H (1975). "Inhibitory effect of anti-obesity drugs on NADH dehydrogenase of mouse heart homogenates". Research Communications in Chemical Pathology and Pharmacology. 11 (4): 645–6. PMID 241101.
- ^ Muller, F. L.; Lustgarten, M. S.; Jang, Y.; Richardson, A. & Van Remmen, H. (2007). "Trends in oxidative aging theories". Free Radic. Biol. Med. 43 (4): 477–503. doi:10.1016/j.freeradbiomed.2007.03.034. PMID 17640558.
- ^ Skulachev VP (May 1996). "Role of uncoupled and non-coupled oxidations in maintenance of safely low levels of oxygen and its one-electron reductants". Q. Rev. Biophys. 29 (2): 169–202. doi:10.1017/s0033583500005795. PMID 8870073. S2CID 40859585.
- ^ a b Muller F (2000). "The nature and mechanism of superoxide production by the electron transport chain: Its relevance to aging". AGE. 23 (4): 227–253. doi:10.1007/s11357-000-0022-9. PMC 3455268. PMID 23604868.
- ^ a b Muller FL, Liu Y, Van Remmen H (November 2004). "Complex III releases superoxide to both sides of the inner mitochondrial membrane". J. Biol. Chem. 279 (47): 49064–73. doi:10.1074/jbc.M407715200. PMID 15317809.
- ^ Han D, Williams E, Cadenas E (January 2001). "Mitochondrial respiratory chain-dependent generation of superoxide anion and its release into the intermembrane space". Biochem. J. 353 (Pt 2): 411–6. doi:10.1042/0264-6021:3530411. PMC 1221585. PMID 11139407.
- ^ Bottani, E; Cerutti, R; Harbour, ME; Ravaglia, S; Dogan, SA; Giordano, C; Fearnley, IM; D'Amati, G; Viscomi, C; Fernandez-Vizarra, E; Zeviani, M (6 July 2017). "TTC19 Plays a Husbandry Role on UQCRFS1 Turnover in the Biogenesis of Mitochondrial Respiratory Complex III". Molecular Cell. 67 (1): 96–105.e4. doi:10.1016/j.molcel.2017.06.001. PMID 28673544.
- ^ DiMauro S (November 2006). "Mitochondrial myopathies" (PDF). Curr Opin Rheumatol. 18 (6): 636–41. doi:10.1097/01.bor.0000245729.17759.f2. PMID 17053512. S2CID 29140366.
- ^ DiMauro S (June 2007). "Mitochondrial DNA medicine". Biosci. Rep. 27 (1–3): 5–9. doi:10.1007/s10540-007-9032-5. PMID 17484047. S2CID 5849380.
- ^ Schuelke M, Krude H, Finckh B, Mayatepek E, Janssen A, Schmelz M, Trefz F, Trijbels F, Smeitink J (March 2002). "Septo-optic dysplasia associated with a new mitochondrial cytochrome b mutation". Ann. Neurol. 51 (3): 388–92. doi:10.1002/ana.10151. PMID 11891837. S2CID 12425236.
- ^ Wibrand F, Ravn K, Schwartz M, Rosenberg T, Horn N, Vissing J (October 2001). "Multisystem disorder associated with a missense mutation in the mitochondrial cytochrome b gene". Ann. Neurol. 50 (4): 540–3. doi:10.1002/ana.1224. PMID 11601507. S2CID 8944744.
- ^ Fisher N, Castleden CK, Bourges I, Brasseur G, Dujardin G, Meunier B (March 2004). "Human disease-related mutations in cytochrome b studied in yeast". J. Biol. Chem. 279 (13): 12951–8. doi:10.1074/jbc.M313866200. PMID 14718526.
Further reading
[edit]- Marres CM, Slater EC (1977). "Polypeptide composition of purified QH2:cytochrome c oxidoreductase from beef-heart mitochondria". Biochim. Biophys. Acta. 462 (3): 531–548. doi:10.1016/0005-2728(77)90099-8. PMID 597492.
- Rieske JS (1976). "Composition, structure, and function of complex III of the respiratory chain". Biochim. Biophys. Acta. 456 (2): 195–247. doi:10.1016/0304-4173(76)90012-4. PMID 788795.
- Wikstrom M, Krab K, Saraste M (1981). "Proton-translocating cytochrome complexes". Annu. Rev. Biochem. 50: 623–655. doi:10.1146/annurev.bi.50.070181.003203. PMID 6267990.
External links
[edit]- cytochrome bc1 complex site (Edward A. Berry) at the Wayback Machine (archived October 9, 2006) at lbl.gov
- cytochrome bc1 complex site (Antony R. Crofts) Archived 2007-09-17 at the Wayback Machine at uiuc.edu
- PROMISE Database: cytochrome bc1 complex at archive.today (archived August 27, 1999) at scripps.edu
- Interactive Molecular Model of Complex III at the Wayback Machine (archived January 12, 2009) (Requires MDL Chime)
- UMich Orientation of Proteins in Membranes families/superfamily-3 - Calculated positions of bc1 and related complexes in membranes
- Coenzyme+Q-Cytochrome-c+Reductase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- #124000 MITOCHONDRIAL COMPLEX III DEFICIENCY, NUCLEAR TYPE 1; MC3DN1 on OMIM; lists all other types of complex III deficiency
Coenzyme Q – cytochrome c reductase
View on GrokipediaNomenclature and overview
Nomenclature
Coenzyme Q – cytochrome c reductase, also known as ubiquinol:cytochrome-c oxidoreductase, is the systematic name for this enzyme, reflecting its role in oxidizing ubiquinol (the reduced form of coenzyme Q) while reducing cytochrome c.[4] It is classified under the Enzyme Commission number EC 7.1.1.8, which denotes its function as a membrane-bound oxidoreductase transferring electrons from an iron-sulfur protein to a quinone or related compound, an update from its prior designation as EC 1.10.2.2 in 2018 to better align with classifications for proton-translocating complexes.[5][6] Common alternative names include Complex III, referring to its position as the third protein complex in the mitochondrial electron transport chain; the bc1 complex, highlighting its core cytochromes b and c1; and coenzyme Q:cytochrome c oxidoreductase, emphasizing the electron transfer from coenzyme Q to cytochrome c.[4][5] The nomenclature evolved from early studies of mitochondrial electron transfer in the 1960s, when researchers isolated and characterized respiratory chain components from beef heart mitochondria, initially describing it as a reduced coenzyme Q-cytochrome c reductase based on its catalytic activity. This functional naming persisted as subsequent work delineated its distinct subunits and cofactors, solidifying terms like Complex III within the broader framework of respiratory complexes established by that era's biochemical fractionations. It is distinguished from other respiratory complexes—such as Complex I (NADH:ubiquinone oxidoreductase, EC 7.1.1.2), Complex II (succinate:ubiquinone oxidoreductase, EC 1.3.5.1), and Complex IV (cytochrome c oxidase, EC 7.1.1.9)—by its specific mediation of electron transfer between ubiquinol and cytochrome c, without involvement in NADH or succinate oxidation or final oxygen reduction.[4][5]Biological role
Coenzyme Q – cytochrome c reductase, also known as mitochondrial complex III or the bc1 complex, is embedded in the inner mitochondrial membrane where it functions as a central component of the electron transport chain (ETC) in aerobic respiration.[7] This positioning allows it to facilitate the sequential transfer of electrons through the respiratory chain, linking upstream complexes like complex I and II to downstream cytochrome c and complex IV.[8] The primary biological role of the enzyme involves oxidizing ubiquinol (QH2), the reduced form of coenzyme Q, and reducing cytochrome c, thereby shuttling electrons from the lipid-soluble ubiquinone pool to the water-soluble cytochrome c in the intermembrane space.[7] This electron transfer is coupled to the translocation of protons across the inner mitochondrial membrane via the Q cycle mechanism, generating a proton motive force that establishes an electrochemical gradient essential for energy conservation.[9] In mammalian mitochondria, complex III assembles into higher-order respiratory supercomplexes, such as I1III2IV1 and I2III2IV2, which enhance channeling of electrons and substrates for efficient respiration; recent cryo-EM structures from 2022 have resolved these assemblies in situ, revealing how they optimize electron flow while minimizing reactive oxygen species production.[10] Beyond its canonical function in the ETC, complex III indirectly supports non-canonical pathways through interactions with the coenzyme Q pool, such as the oxidation of hydrogen sulfide (H2S) mediated by sulfide:quinone oxidoreductase (SQOR). SQOR transfers electrons from H2S to ubiquinone, producing persulfides and feeding into the Q pool for subsequent oxidation by complex III, thereby regulating sulfide homeostasis and preventing toxicity while contributing to mitochondrial redox balance, as highlighted in 2021 reviews on CoQ dynamics.[8] This process underscores the enzyme's broader role in cellular signaling and detoxification.[11] The proton-pumping activity of complex III, which translocates four protons per two electrons transferred in the Q cycle, directly contributes to the proton gradient that drives ATP synthesis by ATP synthase (complex V) in oxidative phosphorylation.[12] This coupling ensures efficient energy production from nutrient oxidation, with disruptions in complex III function leading to impaired ATP generation and mitochondrial dysfunction.[13]Structure and composition
Overall architecture
Coenzyme Q – cytochrome c reductase, commonly known as Complex III or the cytochrome bc1 complex, functions as a symmetric dimer embedded in the inner mitochondrial membrane, with each monomer comprising 11 distinct subunits in humans.[14] The dimeric organization enhances stability and coordinates electron transfer across the two monomers, spanning approximately 100 Å in height and 70 Å in width. At the core of each monomer lie three essential redox-active subunits: cytochrome b, the Rieske iron-sulfur protein (ISP), and cytochrome c1, which form the catalytic heart of the complex. Cytochrome b is a transmembrane protein harboring the Qo site on the intermembrane space (positive) side and the Qi site on the matrix (negative) side, enabling bifurcated electron flow in the Q-cycle. The Rieske ISP, with its [2Fe-2S] cluster, and cytochrome c1, featuring a c-type heme, are anchored peripherally on the positive side to facilitate docking of ubiquinol and cytochrome c, respectively. Proton translocation pathways consist of hydrophilic channels lined by charged residues, such as histidines in the Rieske ISP and conserved water networks near the Qo site, allowing the uptake and release of four protons per Q-cycle. Lipid interactions, particularly with cardiolipin molecules at the dimer interface and subunit boundaries, rigidify the structure and influence conformational dynamics essential for catalysis.[15][16] Cryo-electron microscopy (cryo-EM) has elucidated the overall architecture with atomic-level detail. Structures from the 2010s, including bovine Complex III resolved at around 3.5 Å, first delineated the symmetric dimer and key binding pockets for quinones and inhibitors. Subsequent 2024 in situ cryo-EM studies on porcine mitochondria refined these to higher resolutions (better than 3 Å locally), revealing native lipid densities and subtle asymmetries in monomer orientations that support functional dimerism. In apicomplexan organisms, such as Toxoplasma gondii, 2025 cryo-EM structures of the Complex III2 dimer at 3.2 Å resolution underscore conserved features like the cytochrome b core and Rieske positioning, despite additional clade-specific subunits expanding the monomer to 14 components.[15][17] Complex III further integrates into larger respiratory supercomplexes within the membrane, promoting efficient electron channeling. Notably, the I2+III2 supercomplex, visualized by cryo-EM at 3.7 Å resolution in 2025, forms a compact assembly where the III2 dimer bridges two Complex I monomers, stabilizing the cristae architecture and mitigating defects in electron transport under oxidative stress.[18]Subunit composition
Coenzyme Q – cytochrome c reductase in humans consists of 11 protein subunits arranged in a monomeric unit that dimerizes to form the functional complex. The three core catalytic subunits—cytochrome b, cytochrome c₁, and the Rieske iron-sulfur protein (ISP)—house the redox-active cofactors responsible for electron transfer from ubiquinol to cytochrome c, while the two core proteins and six supernumerary subunits provide structural integrity, stability, and regulatory roles.[15] The redox cofactors are integral to the catalytic mechanism: cytochrome b binds two low-potential hemes (b_H and b_L) for bifurcated electron transfer in the Q cycle; cytochrome c₁ contains a high-potential heme c₁ for transferring electrons to cytochrome c; and the ISP harbors a [2Fe–2S] cluster that shuttles electrons between the quinol oxidation site and cytochrome c₁.[15][19] Stoichiometry is one copy of each subunit per monomer, with the dimer exhibiting twofold symmetry.| Subunit Name | Gene Symbol | Mass (kDa) | Function/Cofactor | Notes (Organism Comparison) |
|---|---|---|---|---|
| Cytochrome b | MT-CYB | 42 | Transmembrane electron carrier; binds hemes b_H and b_L for Q cycle | Mitochondrial-encoded; conserved across species |
| Cytochrome c₁ | CYC1 | 35 | Electron transfer to cytochrome c; binds heme c₁ | Nuclear-encoded; conserved |
| Rieske ISP | UQCRFS1 | 24 (mature) | Mobile electron shuttle; [2Fe–2S] cluster | Nuclear-encoded; processed post-translationally; conserved |
| Core protein 1 | UQCRC1 | 53 | Structural scaffold; protease-like activity | Nuclear-encoded; conserved |
| Core protein 2 | UQCRC2 | 48 | Structural scaffold; protease-like activity | Nuclear-encoded; conserved |
| Subunit VII (QCR7) | UQCRB | 13 | Stabilizes Qi site; ubiquinol binding | Nuclear-encoded; conserved |
| Subunit VIII (QCR8) | UQCRQ | 10 | Membrane anchor; structural | Nuclear-encoded; conserved |
| Hinge protein (QCR6) | UQCRH | 10 | Regulates Rieske domain mobility | Nuclear-encoded; conserved |
| Subunit IX (QCR9) | UQCR10 | 7 | Structural stabilization | Nuclear-encoded; conserved |
| Subunit X (QCR10) | UQCR11 | 6 | Structural stabilization | Nuclear-encoded; conserved |
| Subunit XI (QCR11) | UQCR11 | 6.5 | Structural stabilization of dimer interface | Nuclear-encoded; absent in yeast (10 subunits total) |
Reaction and mechanism
Catalyzed reaction
Coenzyme Q – cytochrome c reductase, also known as the cytochrome bc₁ complex or Complex III, catalyzes the transfer of electrons from ubiquinol (QH₂) to cytochrome c, coupled to the translocation of protons across the inner mitochondrial membrane. The overall stoichiometry of the reaction, as part of the Q-cycle mechanism, is represented by the equation: This net reaction involves the oxidation of one molecule of ubiquinol to ubiquinone (Q) via the Qo site on the intermembrane space side, reduction of two molecules of oxidized cytochrome c, uptake of two protons from the matrix, and release of four protons into the intermembrane space.[24] The Q-cycle mechanism, proposed by Peter Mitchell, enables this process by bifurcating the two electrons from ubiquinol: one electron follows the high-potential chain through the Rieske iron-sulfur protein and cytochrome c₁ to reduce cytochrome c, while the other traverses the low-potential chain via the b cytochromes to reduce a ubiquinone molecule at the Qi site, regenerating a semiquinone intermediate that completes the cycle. The net effect is the transfer of four protons across the membrane per two electrons transferred from ubiquinol to cytochrome c, doubling the proton-to-electron stoichiometry compared to a linear mechanism.[24] The reaction is thermodynamically favorable due to the difference in standard redox potentials between the substrates and products. The midpoint redox potential (E_m) for the ubiquinol/ubiquinone couple is approximately +60 mV, while that for the cytochrome c (oxidized/reduced) couple is about +250 mV, providing a sufficient driving force (ΔE ≈ +190 mV) for electron transfer under physiological conditions.[9] This proton translocation contributes to the generation of the proton motive force (Δp), comprising a pH gradient (ΔpH) and a membrane potential (Δψ), which drives ATP synthesis via ATP synthase in the electron transport chain. The efficiency of this energy conservation mechanism underscores the role of the Q-cycle in mitochondrial oxidative phosphorylation.[9]Detailed mechanism
The detailed mechanism of coenzyme Q–cytochrome c reductase, also known as the cytochrome bc₁ complex, operates through the Q-cycle, a bifurcated electron transfer process that couples the oxidation of ubiquinol (QH₂) to the reduction of cytochrome c while translocating protons across the inner mitochondrial membrane.[25] In the initial step of the Q-cycle, QH₂ binds at the quinol oxidation site (Qo site) on the positive (P) side of the membrane, where it undergoes two-electron oxidation. One electron is transferred to the Rieske iron-sulfur protein (ISP), which then relays it through the high-potential chain to cytochrome c₁ and subsequently to cytochrome c, while the second electron is directed to the low-potential chain via heme b_L to heme b_H.[26] This bifurcation generates a transient semiquinone anion (SQ⁻) at the Qo site, and two scalar protons are released into the intermembrane space (P-side) during QH₂ deprotonation.[25] The electron from the b hemes reaches the quinone reduction site (Qi site) on the negative (N) side, where it reduces ubiquinone (Q) to a semiquinone anion (SQ⁻), with no net proton uptake in this half-cycle.[26] A second turnover of the cycle is required for completion: another QH₂ molecule binds at the Qo site, repeating the bifurcation process, with its high-potential electron again reducing cytochrome c and its low-potential electron reducing the SQ⁻ at the Qi site to ubiquinol (QH₂). This second reduction at Qi incorporates two protons from the matrix (N-side), completing the vectorial proton translocation.[25] Overall, the full Q-cycle results in the net oxidation of one QH₂ at Qo, reduction of one Q to QH₂ at Qi, transfer of two electrons to cytochrome c, release of four scalar protons to the P-side, and uptake of two vectorial protons from the N-side, establishing a proton gradient for ATP synthesis.[26] Central to the Qo site reaction is the conformational dynamics of the ISP, whose extrinsic head domain—containing the [2Fe-2S] cluster—undergoes a large-scale movement of approximately 20–30 Å via a flexible hinge, swinging from a position proximal to the Qo site (b-position) to one adjacent to cytochrome c₁ (c-position) to facilitate electron transfer.[27] This mobility ensures efficient bifurcation by positioning the ISP for initial electron acceptance from QH₂ before relocating for delivery to c₁, with the domain's redox state and quinone occupancy regulating the transition.[27] The distinction between scalar and vectorial proton transport in the Q-cycle underscores its protonmotive efficiency: scalar protons are directly released from substrate oxidation on the P-side, whereas vectorial protons are effectively pumped by the asymmetric uptake at Qi, driven by the redox span across the membrane.[25] Recent cryo-EM structures of the inhibitor-bound III₂–IV supercomplex containing the cytochrome bc₁ complex from the apicomplexan parasite Toxoplasma gondii (2025) provide mechanistic insights into the Qo and Qi sites in a near-native state.[28]Inhibitors and reactive oxygen species
Inhibitors
Coenzyme Q – cytochrome c reductase, also known as the cytochrome bc1 complex or Complex III, is inhibited by compounds that target its two distinct quinone-binding sites: the Qo site (or Qp site) on the positive (intermembrane space) side and the Qi site (or Qn site) on the negative (matrix) side. These inhibitors disrupt the Q-cycle mechanism by blocking electron transfer from ubiquinol (QH2) or to ubiquinone (Q), respectively.[1] Qo site inhibitors, such as stigmatellin and myxothiazol, bind near the iron-sulfur protein (ISP) subunit and prevent the initial oxidation of ubiquinol, thereby halting the bifurcated electron transfer pathway. Stigmatellin coordinates with the ISP histidine ligand and a conserved glutamate residue, stabilizing the complex in a conformation that impedes Qo site occupancy.[1] Myxothiazol, a methoxyacrylate, occupies a hydrophobic pocket adjacent to the Qo site, displacing the ISP and inhibiting its movement essential for electron donation to cytochrome c1.[29] Qi site inhibitors, exemplified by antimycin A, bind within the Qi pocket of cytochrome b, blocking the reduction of ubiquinone to semiquinone and preventing electron flow from the low-potential b heme (bH) to Q. This inhibition traps electrons in the complex, disrupting the overall proton translocation coupled to the Q-cycle.[1] Ilicicolin H acts similarly at the Qi site but with lower affinity than antimycin A; it is antimycin-insensitive in certain yeast assays, allowing its use to probe residual activity in antimycin-treated complexes.[30][31] Atovaquone, a hydroxynaphthoquinone, targets the Qo site in Plasmodium falciparum's bc1 complex and is a frontline antimalarial drug that collapses the mitochondrial membrane potential in parasites. Recent cryo-EM structures from 2025 reveal atovaquone binding in the Toxoplasma gondii III2–IV supercomplex, showing it occupies the Qo pocket and interacts with key residues of cytochrome b, confirming its conserved mechanism across apicomplexans.[17] Antifungal agents targeting the bc1 complex, such as natural product-derived Qi site inhibitors like ilicicolin H and synthetic analogs (e.g., AS2077715), selectively impair fungal respiration and virulence by exploiting differences in the Qo or Qi sites between fungi and mammals.[32][33] Endogenous regulation of Complex III activity involves cardiolipin, a mitochondrial phospholipid that binds to the complex and stabilizes its dimeric assembly, enhancing electron transfer efficiency at both Qo and Qi sites. Cardiolipin deficiency impairs supercomplex formation with Complexes IV and reduces overall respiratory chain activity, underscoring its role as a non-competitive modulator.[34][35]Production of reactive oxygen species
Coenzyme Q – cytochrome c reductase, also known as complex III, generates superoxide (O₂⁻•) primarily at the quinol oxidation site (Qo site) through the unstable semiquinone intermediate formed during ubiquinol oxidation.[36] This semiquinone radical (SQ⁻) reacts with molecular oxygen, leading to superoxide production as a byproduct of electron leakage in the electron transport chain.[37] The process occurs via both forward electron transfer (from ubiquinol to cytochrome c) and reverse electron transfer pathways, where high proton motive force drives electrons backward, enhancing SQ⁻ stability and ROS output.[37][38] Under physiological conditions, ROS from complex III serve as signaling molecules, such as in hypoxia, where reduced oxygen availability stabilizes the hypoxia-inducible factor-1α (HIF-1α) via superoxide-mediated oxidation of prolyl hydroxylases.[39][40] This signaling promotes adaptive responses like angiogenesis and glycolysis. In contrast, excessive ROS contribute to cellular damage, including lipid peroxidation in ferroptosis, where the Nrf2/FSP1/CoQ10 axis suppresses ferroptosis by scavenging radicals and preventing iron-dependent propagation.[41] Reverse electron transport, often triggered by high succinate levels, and hypoxic conditions further elevate ROS production at the Qo site by altering electron flux and SQ⁻ lifetime.[38][39] Mitochondrial coenzyme Q (CoQ) acts as a primary radical scavenger, intercepting superoxide and other ROS near the Qo site to mitigate oxidative stress.[42] By regenerating its reduced form (ubiquinol), CoQ prevents propagation of radical chain reactions within the inner membrane. Recent studies demonstrate that CoQ10 supplementation reduces mitochondrial ROS levels in models of metabolic disorders, such as CDKL5 deficiency, by restoring antioxidant capacity and normalizing glutathione balance.[43] In neuronal models of oxidative stress, CoQ10 pretreatment significantly lowers superoxide production, highlighting its therapeutic potential in ROS-related pathologies.[42]Genetics and human pathology
Encoding genes
The Coenzyme Q – cytochrome c reductase, also known as Complex III or the cytochrome bc1 complex, is composed of 11 protein subunits in humans, with their encoding genes distributed between the mitochondrial and nuclear genomes. Only one subunit, cytochrome b, is encoded by the mitochondrial genome, while the remaining ten subunits are encoded by nuclear genes. These nuclear-encoded subunits include two core catalytic subunits (cytochrome c1 and the Rieske iron-sulfur protein) and eight other nuclear-encoded subunits (including core proteins and supernumerary subunits) that contribute to stability, assembly, and regulation.[44][45] The mitochondrially encoded gene is MT-CYB (also known as MTCYB), located on the mitochondrial DNA (mtDNA) at positions 14,748–15,826 (revised Cambridge reference sequence). It produces the cytochrome b protein, a transmembrane subunit essential for ubiquinol oxidation and the Q-cycle mechanism within Complex III.[45] All other subunits are nuclear-encoded, synthesized in the cytosol, and imported into the mitochondria for assembly with cytochrome b. The nuclear genes encoding the Complex III subunits are as follows:| Gene Symbol | Chromosome Location | Protein Product | Function |
|---|---|---|---|
| CYC1 | 8q24.3 | Cytochrome c1 | Heme-containing subunit that accepts electrons from the Rieske iron-sulfur protein and transfers them to cytochrome c.[46][47] |
| UQCRFS1 | 19p13.3 | Rieske iron-sulfur protein | Contains a [2Fe-2S] cluster that facilitates electron transfer from ubiquinol to cytochrome c1.[48][49] |
| UQCRC1 | 3p21.31 | Core protein 1 | Structural core subunit involved in complex assembly and stabilization.[50][51] |
| UQCRC2 | 16q13 | Core protein 2 | Structural core subunit that supports dimerization and overall complex integrity.[52][53] |
| UQCRB | 8q22.1 | Ubiquinol-cytochrome c reductase binding protein (QP-C) | Hinge protein that aids in cytochrome c binding and electron transfer.[44][54] |
| UQCRH | 1p33 | Hinge protein | Assists in cytochrome c docking and complex stability.[55][56] |
| UQCRQ | 5q31.1 | Complex III subunit VII (QCR8) | Small supernumerary subunit associated with the core and involved in ubiquinone binding.[57][58] |
| UQCR10 | 22q11.21 | Complex III subunit X (QCR9) | Low-molecular-weight supernumerary subunit contributing to structural integrity.[59][60] |
| UQCR11 | 19p13.3 | Complex III subunit XI (QCR10) | Smallest supernumerary subunit aiding in complex assembly and stability.[61][62] |