Phosphoglucomutase
View on Wikipedia| Phosphoglucomutase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
 | |||||||||
| Identifiers | |||||||||
| EC no. | 5.4.2.2 | ||||||||
| CAS no. | 9001-81-4 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Phosphoglucomutase (EC 5.4.2.2) is an enzyme that transfers a phosphate group on an α-D-glucose monomer from the 1 to the 6 position in the forward direction or the 6 to the 1 position in the reverse direction.
More precisely, it facilitates the interconversion of glucose 1-phosphate and glucose 6-phosphate.
Function
[edit]Role in glycogenolysis
[edit]After glycogen phosphorylase catalyzes the phosphorolytic cleavage of a glucosyl residue from the glycogen polymer, the freed glucose has a phosphate group on its 1-carbon. This glucose 1-phosphate molecule is not itself a useful metabolic intermediate, but phosphoglucomutase catalyzes the conversion of this glucose 1-phosphate to glucose 6-phosphate (see below for the mechanism of this reaction).
Glucose 6-phosphate’s metabolic fate depends on the needs of the cell at the time it is generated. If the cell is low on energy, then glucose 6-phosphate will travel down the glycolytic pathway, eventually yielding two molecules of adenosine triphosphate. If the cell is in need of biosynthetic intermediates, glucose 6-phosphate will enter the pentose phosphate pathway, where it will undergo a series of reactions to yield riboses and/or NADPH, depending on cellular conditions.
If glycogenolysis is taking place in the liver, glucose 6-phosphate can be converted to glucose by the enzyme glucose 6-phosphatase; the glucose produced in the liver is then released to the bloodstream for use in other organs. Muscle cells in contrast do not have the enzyme glucose 6-phosphatase, so they cannot share their glycogen stores with the rest of the body.
Role in glycogenesis
[edit]Phosphoglucomutase also acts in the opposite fashion when blood glucose levels are high. In this case, phosphoglucomutase catalyzes the conversion of glucose 6-phosphate (which is easily generated from glucose by the action of hexokinase) to glucose 1-phosphate.
This glucose-1-phosphate can then react with UTP to yield UDP-glucose in a reaction catalyzed by UDP-glucose-pyrophosphorylase. If activated by insulin, glycogen synthase will proceed to clip the glucose from the UDP-glucose complex onto a glycogen polymer.
Reaction mechanism
[edit]Phosphoglucomutase affects a phosphoryl group shift by exchanging a phosphoryl group with the substrate.[1] Isotopic labeling experiments have confirmed that this reaction proceeds through a glucose 1,6-bisphosphate intermediate.[2]
The first step in the forward reaction is the transfer of a phosphoryl group from the enzyme to glucose 1-phosphate, forming glucose 1,6-bisphosphate and leaving a dephosphorylated form of the enzyme.[2] The enzyme then undergoes a rapid diffusional reorientation to position the 1-phosphate of the bisphosphate intermediate properly relative to the dephosphorylated enzyme.[3] Substrate-velocity relationships and induced transport tests have revealed that the dephosphorylated enzyme then facilitates the transfer of a phosphoryl group from the glucose-1,6-bisphosphate intermediate to the enzyme, regenerating phosphorylated phosphoglucomutase and yielding glucose 6-phosphate (in the forward direction).[4][5] Later structural studies confirmed that the single site in the enzyme that becomes phosphorylated and dephosphorylated is the oxygen of the active-site serine residue (see diagram below).[6][7] A bivalent metal ion, usually magnesium or cadmium, is required for enzymatic activity and has been shown to complex directly with the phosphoryl group esterified to the active-site serine.[8]
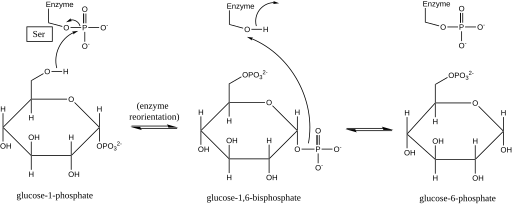
This formation of a glucose 1,6-bisphosphate intermediate is analogous to the interconversion of 2-phosphoglycerate and 3-phosphoglycerate catalyzed by phosphoglycerate mutase, in which 2,3-bisphosphoglycerate is generated as an intermediate.[9]
Structure
[edit]
While rabbit muscle phosphoglucomutase has served as the prototype for much of the elucidation of this enzyme's structure, newer bacterium-derived crystal structures exhibit many of the same defining characteristics.[10] Each phosphoglucomutase monomer can be divided into four sequence domains, I-IV, based on the enzyme’s default spatial configuration (see image at right).[11]
Each monomer comprises four distinct α/β structural units, each of which contains one of the four strands in each monomer's β-sheet and is made up only of the residues in a given sequence domain (see image at right).[11] The burial of the active site (including Ser-116, the critical residue on the enzyme that is phosphorylated and dephosphorylated) in the hydrophobic interior of the enzyme serves to exclude water from counterproductively hydrolyzing critical phosphoester bonds while still allowing the substrate to access the active site.[12]
Disease relevance
[edit]Human muscle contains two isoenzymes of phosphoglucomutase with nearly identical catalytic properties, PGM I and PGM II.[13] One or the other of these forms is missing in some humans congenitally.[14] PGM1 deficiency is known as PGM1-CDG or CDG syndrome type 1t (CDG1T), formerly known as glycogen storage disease type 14 (GSD XIV).[15][16] The disease is both a glycogenosis and a congenital disorder of glycosylation.[17][18] It is also a metabolic myopathy and an inborn error of carbohydrate metabolism.[19]
PGM deficiency is an extremely rare condition that does not have a set of well-characterized physiological symptoms. This condition can be detected by an in vitro study of anaerobic glycolysis which reveals a block in the pathway toward lactic acid production after glucose 1-phosphate but before glucose 6-phosphate.[20] There are two forms of PGM1-CDG: 1.) exclusively myogenic, and 2.) multi-system (including muscles).[16]
The usual pathway for glycogen formation from blood glucose is blocked, as without phosphoglucomutase, glucose-6-phosphate cannot convert into glucose-1-phosphate. However, an alternative pathway from galactose can form glycogen by converting galactose → galactose-1-phosphate → glucose-1-phosphate. This allows glycogen to form, but without phosphoglucomutase, glucose-1-phosphate cannot convert into glucose-6-phosphate for glycolysis. This causes abnormal glycogen accumulation in muscle cells, observable in muscle biopsy.[16][21]
Although the phenotype and severity of the disease is highly variable, common symptoms include: exercise intolerance, exercise-induced hyperammonemia, abnormal glycogen accumulation in muscle biopsy, elevated serum CK, abnormal serum transferrin (loss of complete N-glycans), short stature, cleft palate, bifid uvula, and hepatopathy.[16][21]
A "second wind" phenomenon is observable in some, but not all, by measuring heart rate while on a treadmill.[16][22] At rest, muscle cells rely on blood glucose and free fatty acids; upon exertion, muscle glycogen is needed along with blood glucose and free fatty acids.[23][24] The reliance on muscle glycogen increases with higher-intensity aerobic exercise and all anaerobic exercise.[23][24]
Without being able to create ATP from stored muscle glycogen, during exercise there is a low ATP reservoir (ADP>ATP). Under such circumstances, the heart rate and breathing increases inappropriately given the exercise intensity, in an attempt to maximize the delivery of oxygen and blood borne fuels to the muscle cell. Free fatty acids are the slowest of the body's bioenergetic systems to produce ATP by oxidative phosphorylation, at approximately 10 minutes.[23] The relief of exercise intolerance symptoms, including a drop in heart rate of at least 10 BPM while going the same speed on the treadmill, after approximately 10 minutes of aerobic exercise is called "second wind," where increased ATP is being produced from free fatty acids.
Another consequence of a low ATP reservoir (ADP>ATP) during exercise, due to not being able to produce ATP from muscle glycogen, is increased use of the myokinase (adenylate kinase) reaction and the purine nucleotide cycle. The myokinase reaction produces AMP (2 ADP → ATP + AMP), and then the purine nucleotide cycle both uses AMP and produces more AMP along with fumarate (the fumarate is then converted and produces ATP via oxidative phosphorylation). Ammonia (NH3) is a byproduct in the purine nucleotide cycle when AMP is converted into IMP. During a non-ischemic forearm test, PGM1-CDG individuals show exercise-induced elevated serum ammonia (hyperammonemia) and normal serum lactate rise.[16][18][19]
Studies in other diseases that have a glycolytic block have shown during ischemic and non-ischemic forearm exercise tests, that not only does ammonia rise, but after exercise, rises also in serum inosine, hypoxanthine, and uric acid.[25][26] These studies supported that when the exercise is stopped or sufficient ATP is produced from other fuels (such as free fatty acids), then the ATP reservoir normalizes and the buildup of AMP and other nucleotides covert into nucleosides and leave the muscle cell to be converted into uric acid, known as myogenic hyperuricemia. AMP → IMP → Inosine → Hypoxanthine → Xanthine → Uric acid. Unfortunately, the studies on PGM1-CDG only tested for serum ammonia and lactate, so it is currently unknown definitively whether PGM1-CDG individuals also experience myogenic hyperuricemia.[16][18][19]
Genes
[edit]See also
[edit]References
[edit]- ^ Jagannathan V, Luck JM (June 1949). "Phosphoglucomutase; mechanism of action". The Journal of Biological Chemistry. 179 (2): 569–575. doi:10.1016/S0021-9258(19)51252-2. PMID 18149991.
- ^ a b Najjar VA, Pullman ME (May 1954). "The occurrence of a group transfer involving enzyme (phosphoglucomutase) and substrate". Science. 119 (3097): 631–634. Bibcode:1954Sci...119..631N. doi:10.1126/science.119.3097.631. PMID 13156640.
- ^ Ray Jr WJ, Peck EJ (1972). "Phosphomutases". In Boyer PD (ed.). The Enzymes. Vol. 6 (3rd ed.). New York: Academic Press. pp. 407–477. doi:10.1016/S1874-6047(08)60047-5. ISBN 978-0-12-122706-7.
- ^ Ray WJ, Roscelli GA (April 1964). "A Kinetic Study of the Phosphoglucomutase Pathway". The Journal of Biological Chemistry. 239 (4): 1228–1236. doi:10.1016/S0021-9258(18)91416-X. PMID 14165931.
- ^ Britton HG, Clarke JB (November 1968). "The mechanism of the phosphoglucomutase reaction. Studies on rabbit muscle phosphoglucomutase with flux techniques". The Biochemical Journal. 110 (2): 161–180. doi:10.1042/bj1100161. PMC 1187194. PMID 5726186.
- ^ Ray WJ, Mildvan AS, Grutzner JB (December 1977). "Phosphorus nuclear magnetic resonance studies of phosphoglucomutase and its metal ion complexes". Archives of Biochemistry and Biophysics. 184 (2): 453–463. doi:10.1016/0003-9861(77)90455-6. PMID 23074.
- ^ Ray WJ, Hermodson MA, Puvathingal JM, Mahoney WC (August 1983). "The complete amino acid sequence of rabbit muscle phosphoglucomutase". The Journal of Biological Chemistry. 258 (15): 9166–9174. doi:10.1016/S0021-9258(17)44646-1. PMID 6223925.
- ^ Rhyu GI, Ray WJ, Markley JL (January 1984). "Enzyme-bound intermediates in the conversion of glucose 1-phosphate to glucose 6-phosphate by phosphoglucomutase. Phosphorus NMR studies". Biochemistry. 23 (2): 252–260. doi:10.1021/bi00297a013. PMID 6230103.
- ^ Sutherland EW, Cohn M (October 1949). "The mechanism of the phosphoglucomutase reaction". The Journal of Biological Chemistry. 180 (3): 1285–1295. doi:10.1016/S0021-9258(19)51242-X. PMID 18148026.
- ^ Mehra-Chaudhary R, Mick J, Tanner JJ, Henzl MT, Beamer LJ (April 2011). "Crystal structure of a bacterial phosphoglucomutase, an enzyme involved in the virulence of multiple human pathogens". Proteins. 79 (4): 1215–1229. doi:10.1002/prot.22957. PMC 3066478. PMID 21246636.
- ^ a b Dai JB, Liu Y, Ray WJ, Konno M (March 1992). "The crystal structure of muscle phosphoglucomutase refined at 2.7-angstrom resolution". The Journal of Biological Chemistry. 267 (9): 6322–6337. doi:10.1016/S0021-9258(18)42699-3. PMID 1532581.
- ^ Ray WJ, Puvathingal JM, Liu YW (July 1991). "Formation of substrate and transition-state analogue complexes in crystals of phosphoglucomutase after removing the crystallization salt". Biochemistry. 30 (28): 6875–6885. doi:10.1021/bi00242a011. PMID 1829964.
- ^ Joshi JG, Handler P (June 1969). "Phosphoglucomutase. VI. Purification and properties of phosphoglucomutases from human muscle". The Journal of Biological Chemistry. 244 (12): 3343–3351. doi:10.1016/S0021-9258(18)93132-7. PMID 4978319.
- ^ Brown DH (1986). "Glycogen metabolism and glycolysis in muscle". Myology: Basic and Clinical. New York: McGraw-Hill. pp. 673–95. ISBN 978-0-07-079570-9.
- ^ "Orphanet: Glycogen storage disease due to phosphoglucomutase deficiency". www.orpha.net. Retrieved May 13, 2021.
- ^ a b c d e f g Altassan R, Radenkovic S, Edmondson AC, Barone R, Brasil S, Cechova A, et al. (January 2021). "International consensus guidelines for phosphoglucomutase 1 deficiency (PGM1-CDG): Diagnosis, follow-up, and management". Journal of Inherited Metabolic Disease. 44 (1): 148–163. doi:10.1002/jimd.12286. PMC 7855268. PMID 32681750.
- ^ Tegtmeyer LC, Rust S, van Scherpenzeel M, Ng BG, Losfeld ME, Timal S, et al. (February 2014). "Multiple phenotypes in phosphoglucomutase 1 deficiency". The New England Journal of Medicine. 370 (6): 533–542. doi:10.1056/NEJMoa1206605. PMC 4373661. PMID 24499211.
- ^ a b c Stojkovic T, Vissing J, Petit F, Piraud M, Orngreen MC, Andersen G, et al. (July 2009). "Muscle glycogenosis due to phosphoglucomutase 1 deficiency". The New England Journal of Medicine. 361 (4): 425–427. doi:10.1056/NEJMc0901158. PMID 19625727.
- ^ a b c Hogrel JY, Janssen JB, Ledoux I, Ollivier G, Béhin A, Stojkovic T, et al. (October 2017). "The diagnostic value of hyperammonaemia induced by the non-ischaemic forearm exercise test" (PDF). Journal of Clinical Pathology. 70 (10): 896–898. doi:10.1136/jclinpath-2017-204324. PMID 28400468. S2CID 36935686.
- ^ Sugie H, Kobayashi J, Sugie Y, Ichimura M, Miyamoto R, Ito T, et al. (April 1988). "Infantile muscle glycogen storage disease: phosphoglucomutase deficiency with decreased muscle and serum carnitine levels". Neurology. 38 (4): 602–605. doi:10.1212/WNL.38.4.602. PMID 2965317. S2CID 11491932.
- ^ a b "Congenital Disorder of Glycosylation, Type It; CDG1T". Online Mendelian Inheritance in Man. 2012-07-11.
- ^ Preisler N, Cohen J, Vissing CR, Madsen KL, Heinicke K, Sharp LJ, et al. (November 2017). "Impaired glycogen breakdown and synthesis in phosphoglucomutase 1 deficiency". Molecular Genetics and Metabolism. 122 (3): 117–121. doi:10.1016/j.ymgme.2017.08.007. PMID 28882528.
- ^ a b c "Berne and Levy Physiology, 6th ed 38. Hormonal Regulation of Energy Metabolism".
- ^ a b van Loon LJ, Greenhaff PL, Constantin-Teodosiu D, Saris WH, Wagenmakers AJ (October 2001). "The effects of increasing exercise intensity on muscle fuel utilisation in humans". The Journal of Physiology. 536 (Pt 1): 295–304. doi:10.1111/j.1469-7793.2001.00295.x. PMC 2278845. PMID 11579177.
- ^ Mineo I, Kono N, Hara N, Shimizu T, Yamada Y, Kawachi M, et al. (July 1987). "Myogenic hyperuricemia. A common pathophysiologic feature of glycogenosis types III, V, and VII". The New England Journal of Medicine. 317 (2): 75–80. doi:10.1056/NEJM198707093170203. PMID 3473284.
- ^ Mineo I, Tarui S (1995). "Myogenic hyperuricemia: what can we learn from metabolic myopathies?". Muscle & Nerve. Supplement. 3: S75 – S81. doi:10.1002/mus.880181416. PMID 7603532. S2CID 41588282.
External links
[edit]- Phosphoglucomutase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
