Recent from talks
Nothing was collected or created yet.
Spidroin
View on Wikipedia
This article needs attention from an expert in chemicals. The specific problem is: reason not known. (January 2015) |
| Spidroin, N-terminal | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Spidroin_N | ||||||||
| Pfam | PF16763 | ||||||||
| InterPro | IPR031913 | ||||||||
| CATH | 2lpj | ||||||||
| |||||||||
| Spidroin, C-terminal | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | Spidroin_MaSp | ||||||||
| Pfam | PF11260 | ||||||||
| InterPro | IPR021001 | ||||||||
| CATH | 2m0m | ||||||||
| |||||||||
| Spidroin-1 | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Organism | |||||||
| Symbol | ? | ||||||
| UniProt | P19837 | ||||||
| |||||||
| Spidroin-2 | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Organism | |||||||
| Symbol | ? | ||||||
| UniProt | P46804 | ||||||
| |||||||

Spidroins are the main proteins in spider silk. Different types of spider silk contain different spidroins, all of which are members of a single protein family.[1] The most-researched type of spidroins are the major ampullate silk proteins (MaSp) used in the construction of dragline silk, the strongest type of spider silk. Dragline silk fiber was originally thought to be made up of two types of spidroins, spidroin-1 (MaSp1) and spidroin-2 (MaSp2) however recent transcriptomic analysis of over 1000 spider species has revealed multiple spidroins are expressed making it much more complex.[2][3][4]
Spidroin is part of a large group of proteins called scleroproteins. This group includes other insoluble structural proteins such as collagen and keratin.
A fiber of dragline spidroin is as thick and resistant as one of steel but is more flexible. It can be stretched to approximately 135% of its original length without breaking. Its properties make it an excellent candidate for use in various scientific fields.[5]
Structure
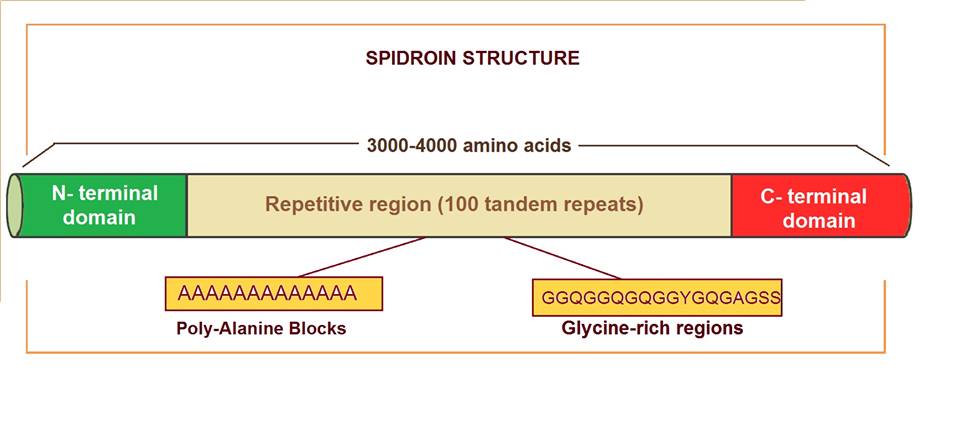
[edit]Major ampullate spidroins are large proteins with an extension of 250-350 kDa, with an average of 3500 amino acids. They represent a polymeric organization, mostly based on highly homogenized tandem repeats. There are 100 tandem copies of 30 to 40 amino acids which repeat sequence and they represent more than 90% of the protein sequence.[6] Alanine and glycine residues are the most abundant amino acids in these proteins. Alanine appears in blocks of six to fourteen units that form β-sheets. These alanine blocks can stack to create crystalline structures in the fiber, linking different protein molecules together. Glycine is present in different motifs, such as GGX and GPGXX (where X = A, L, Q, or Y), that also have specific secondary structures (310-helix and β-spiral, respectively). Glycine-rich regions are more amorphous and contribute to extensibility and flexibility. Some of the differences observed between spidroin 1 and spidroin 2 (the most important major ampullate spidroins) are the proline content, which is very low in the first one but significant in the second one, and the motifs. Motif (GGX)n is characteristic in spidroin 1, while GPG and QQ are typical in spidroin 2.
On the other hand, spidroins have non-repetitive amino (N) and carboxyl (C) terminal domains of approximately 150 and 100 amino acids respectively. N- and C-terminal domains share little resemblance, except that they are both rich in serine and both are largely amphipathic α-helical secondary structures. These domains are conserved not only between spidroin 1 and 2, but also among many silk types and spider species. Experimental data show the N- and C-terminal domains contribute to fiber assembly.[7] The C-terminal domain is involved in the organized transition from a soluble spidroin solution to an insoluble fiber during spinning.[8] In the N-terminal domain, there are signal peptides which regulate spidroin secretion from silk gland cells.[9][10]
Function
[edit]An individual spider spins a multitude of silk types, with each type emerging from its own distinctive set of abdominal silk glands. This complex silk machinery enables spiders to use task-specific silks (e.g., for web assembly, egg-case construction, prey wrapping, etc.).[10] The different types of silk (major ampullate silk, minor ampullate silk, flagelliform silk, aciniform silk, tubiliform silk, pyriform silk, and aggregate silk)[11] are composed of different types of proteins.
Dragline silk is mainly formed by spidroin proteins. It is a type of major ampullate silk and is produced in the major ampullate gland. Dragline silk is used not only to construct the outer frame and radii of the orb-shaped web but also as a hanging lifeline that allows the spider to evade and/or escape from predators.[12] The major ampullate gland that produces this silk is formed by three main sections: a central bag (B zone) flanked by a tail (A zone) and a duct heading towards the exit. The tail secretes most of the "spinning dope", a solution which contains the protein molecules that will constitute the silk fiber. The sac is the main storage repository.
The epithelium of the A zone is composed of tall columns of secretory cells of a single type, packed with secretory granules. The major component of these cells which secrete the fibroin solution is a 275kDa protein containing the polypeptides spidroin I and spidroin II. The output of these cells is an aqueous and highly viscous solution of about 50% protein (mostly spidroin). The product secreted makes up the dragline silk, the main structure.
This highly viscous protein emulsion flows into the B zone, where it is covered by glycoproteins. After exiting this bag, the liquid is funneled into the narrow duct. As the gelatinous protein solution moves into the duct, the integral spidroins and glycoproteins are gradually distorted into long, thin, aligned figures with the direction of the flow. Then, they are stretched and lined up in a way that will eventually allow them to create strong intermolecular links. After different processes the silk is extended in the spinning channel to form an extremely tough thread.
Industrial and biomedical applications
[edit]In the last decade, much research has been done about spidroin protein and spider silk in order to take advantage of some of its properties, such as its elasticity and strength. Spider silk is used in different industries, and its range of applications in biomedicine is increasing every day. For example, the military and defense industries use bulletproof vests made from these fibers.
Recombinant spidroin has been successfully obtained in both eukaryotic and prokaryotic cells although there were some difficulties in the procedure due to the length of the gene sequence. Thanks to expression and the cloning work, it is possible to obtain large-scale production of spidroin which provides new opportunities for the manufacture of new biomaterials.[13] There have been attempts to generate transgenic tobacco and potato plants that express remarkable amounts of recombinant Nephila clavipes dragline proteins.[14]
Furthermore, fibers developed from spidroin are tolerated in vitro, in cell culture, and in vivo, in animals like pigs, as no signs of either inflammatory response nor body reaction were shown to these fibers. These results suggest that they could be used in medicine without risk of biocompatibility issues and thus potentially lead to many new opportunities in tissue engineering and regenerative medicine.
The way spiders produce spidroin in micelles has inspired a method of mass-producing recombinant proteins. By fusing a pH-insensitive, charge-reversed mutant of spidroin N-terminal domain to the proteins to produce, much more soluble proteins can be produced in E. coli.[15]
Other silk types
[edit]| Tubuliform (egg case) silk strands structural domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| Symbol | RP1-2 | ||||||||
| Pfam | PF12042 | ||||||||
| InterPro | IPR021915 | ||||||||
| CATH | 2mqa | ||||||||
| |||||||||
Silk proteins present in other spider silk types are also occasionally referred to as spidroin. These include tubuliform silk protein (TuSP), flagelliform silk protein (Flag; O44358-Q9NHW4-O44359), minor ampullate silk proteins (MiSp; K4MTL7), aciniform silk protein (AcSP), pyriform silk protein (PySp) and aggregate silk glue (ASG2/AgSp). These different silk proteins along with MaSP show some level of homology to each other, in protein domains, repeats, and in promoters, but also have their own unique features and variations on these parts to fulfill their different functions.[16][17][18] These commonalities point at a common origin of proteins found in all these different types of silks.[1][10]
Artificial production
[edit]
In July 2020 a team of Riken researchers report that they succeeded in using a genetically altered variant of R. sulfidophilum to produce spidroins.[19][20]
See also
[edit]References
[edit]- ^ a b Ayoub NA, Garb JE, Kuelbs A, Hayashi CY (March 2013). "Ancient properties of spider silks revealed by the complete gene sequence of the prey-wrapping silk protein (AcSp1)". Molecular Biology and Evolution. 30 (3): 589–601. doi:10.1093/molbev/mss254. PMC 3563967. PMID 23155003.
- ^ Motriuk-Smith D, Smith A, Hayashi CY, Lewis RV (2005). "Analysis of the conserved N-terminal domains in major ampullate spider silk proteins". Biomacromolecules. 6 (6): 3152–9. doi:10.1021/bm050472b. PMID 16283740.
- ^ Moisenovich MM, Pustovalova O, Shackelford J, Vasiljeva TV, Druzhinina TV, Kamenchuk YA, et al. (May 2012). "Tissue regeneration in vivo within recombinant spidroin 1 scaffolds". Biomaterials. 33 (15): 3887–98. doi:10.1016/j.biomaterials.2012.02.013. PMID 22364702.
- ^ Arakawa K, Kono N, Malay AD, Tateishi A, Ifuku N, Masunaga H, et al. (October 2022). "1000 spider silkomes: Linking sequences to silk physical properties". Science Advances. 8 (41) eabo6043. Bibcode:2022SciA....8O6043A. doi:10.1126/sciadv.abo6043. PMC 9555773. PMID 36223455.
- ^ Askarieh G, Hedhammar M, Nordling K, Saenz A, Casals C, Rising A, et al. (May 2010). "Self-assembly of spider silk proteins is controlled by a pH-sensitive relay". Nature. 465 (7295): 236–8. Bibcode:2010Natur.465..236A. doi:10.1038/nature08962. PMID 20463740. S2CID 4366005.
- ^ Xu M, Lewis RV (September 1990). "Structure of a protein superfiber: spider dragline silk". Proceedings of the National Academy of Sciences of the United States of America. 87 (18): 7120–4. Bibcode:1990PNAS...87.7120X. doi:10.1073/pnas.87.18.7120. PMC 54695. PMID 2402494.
- ^ Huemmerich D, Helsen CW, Quedzuweit S, Oschmann J, Rudolph R, Scheibel T (October 2004). "Primary structure elements of spider dragline silks and their contribution to protein solubility". Biochemistry. 43 (42): 13604–12. doi:10.1021/bi048983q. PMID 15491167.
- ^ Sponner A, Vater W, Rommerskirch W, Vollrath F, Unger E, Grosse F, et al. (December 2005). "The conserved C-termini contribute to the properties of spider silk fibroins". Biochemical and Biophysical Research Communications. 338 (2): 897–902. doi:10.1016/j.bbrc.2005.10.048. PMID 16253207.
- ^ Stark M, Grip S, Rising A, Hedhammar M, Engström W, Hjälm G, et al. (May 2007). "Macroscopic fibers self-assembled from recombinant miniature spider silk proteins". Biomacromolecules. 8 (5): 1695–701. doi:10.1021/bm070049y. PMID 17402782.
- ^ a b c Garb JE, Ayoub NA, Hayashi CY (August 2010). "Untangling spider silk evolution with spidroin terminal domains". BMC Evolutionary Biology. 10 (1): 243. Bibcode:2010BMCEE..10..243G. doi:10.1186/1471-2148-10-243. PMC 2928236. PMID 20696068.
- ^ Humenik M, Scheibel T, Smith A (2011). Howorka S (ed.). "Spider silk: understanding the structure-function relationship of a natural fiber". Progress in Molecular Biology and Translational Science. 103. London: Elsevier Science: 131–85. doi:10.1016/B978-0-12-415906-8.00007-8. PMID 21999996.
- ^ Gaines WA, Marcotte WR (September 2008). "Identification and characterization of multiple Spidroin 1 genes encoding major ampullate silk proteins in Nephila clavipes". Insect Molecular Biology. 17 (5): 465–74. doi:10.1111/j.1365-2583.2008.00828.x. PMC 2831225. PMID 18828837.
- ^ Rising A, Widhe M, Johansson J, Hedhammar M (January 2011). "Spider silk proteins: recent advances in recombinant production, structure-function relationships and biomedical applications". Cellular and Molecular Life Sciences. 68 (2): 169–184. doi:10.1007/s00018-010-0462-z. PMC 11114806. PMID 20668909.
- ^ Scheller J, Gührs KH, Grosse F, Conrad U (June 2001). "Production of spider silk proteins in tobacco and potato". Nature Biotechnology. 19 (6): 573–7. doi:10.1038/89335. PMID 11385464. S2CID 517678.
- ^ Kronqvist N, Sarr M, Lindqvist A, Nordling K, Otikovs M, Venturi L, et al. (May 2017). "Efficient protein production inspired by how spiders make silk". Nature Communications. 8 (1) 15504. Bibcode:2017NatCo...815504K. doi:10.1038/ncomms15504. PMC 5457526. PMID 28534479.
- ^ Wen R, Liu X, Meng Q (December 2017). "Characterization of full-length tubuliform spidroin gene from Araneus ventricosus". International Journal of Biological Macromolecules. 105 (Pt 1): 702–710. doi:10.1016/j.ijbiomac.2017.07.086. PMID 28735001.
- ^ Dos Santos-Pinto JR, Arcuri HA, Esteves FG, Palma MS, Lubec G (October 2018). "Spider silk proteome provides insight into the structural characterization of Nephila clavipes flagelliform spidroin". Scientific Reports. 8 (1): 14674. Bibcode:2018NatSR...814674D. doi:10.1038/s41598-018-33068-9. PMC 6168590. PMID 30279551.
- ^ Collin MA, Clarke TH, Ayoub NA, Hayashi CY (February 2016). "Evidence from Multiple Species that Spider Silk Glue Component ASG2 is a Spidroin". Scientific Reports. 6 (1) 21589. Bibcode:2016NatSR...621589C. doi:10.1038/srep21589. PMC 4753498. PMID 26875681.
- ^ "Spider silk made by photosynthetic bacteria". phys.org. Retrieved 16 August 2020.
- ^ Foong CP, Higuchi-Takeuchi M, Malay AD, Oktaviani NA, Thagun C, Numata K (July 2020). "A marine photosynthetic microbial cell factory as a platform for spider silk production". Communications Biology. 3 (1). Springer Science and Business Media LLC: 357. doi:10.1038/s42003-020-1099-6. PMC 7343832. PMID 32641733.
 Text and images are available under a Creative Commons Attribution 4.0 International License.
Text and images are available under a Creative Commons Attribution 4.0 International License.
Further reading
[edit]- Li V. "Spider Silk: Chemical Structure". Molecule of the Month. University of Bristol.
- Altairac S (30 September 2008). "Spidroin". Protein Spotlight. SIB Swiss Institute of Bioinformatics.
- PDB: 3LR2: Askarieh G, Hedhammar M, Nordling K, Saenz A, Casals C, Rising A, et al. (May 2010). "Self-assembly of spider silk proteins is controlled by a pH-sensitive relay". Nature. 465 (7295): 236–8. Bibcode:2010Natur.465..236A. doi:10.1038/nature08962. PMID 20463740. S2CID 4366005.
External links
[edit]Spidroin
View on GrokipediaBiological Context
Definition and Classification
Spidroins are a family of scleroproteins synthesized exclusively by spiders, serving as the primary structural components of all spider silk fibers and comprising over 90% of their mass. These large, modular proteins are characterized by extensive repetitive domains that confer the unique mechanical properties of silk, analogous to the fibroin proteins found in other arthropod silks but adapted for the diverse functional roles in spider biology.[3][4] The term "spidroin," derived from "spider" and "fibroin," was introduced in the early 1990s following the molecular characterization of the first spidroin sequences from the dragline silk of the golden orb-weaver spider Nephila clavipes. The initial partial cDNA for the major ampullate spidroin 1 (MaSp1) was isolated and sequenced in 1990, revealing its highly repetitive alanine- and glycine-rich structure, while a second dragline protein, MaSp2, was identified shortly thereafter in 1992. These discoveries marked the beginning of systematic studies on spider silk proteins, distinguishing them from previously known insect silks. Spidroins are classified into seven principal types based on the specialized silk-producing glands in which they are expressed, reflecting the functional diversity of spider silks. These include MaSp1 and MaSp2 from the major ampullate glands, which form the tough dragline silk used for bridging and safety lines; MiSp from the minor ampullate glands, contributing to temporary scaffolding; AcSp from the aciniform glands, used in prey wrapping; TuSp from the tubuliform glands for egg case construction; Flag from the flagelliform glands for capture spiral threads; Pyr from the pyriform glands for attachment discs; and AgSp from the aggregate glands, which produce adhesive glue on cribellar silk. This gland-specific nomenclature—using two-letter abbreviations followed by "Sp" for spidroin—standardizes the identification of isoforms across species.[1][5] Both spidroins and the fibroin heavy chain produced by silkworms (Bombyx mori) feature conserved non-repetitive N-terminal (NT) and C-terminal (CT) domains that bookend their central repetitive sequences, but spidroin terminal domains exhibit higher conservation across types and species, playing critical roles in protein dimerization, alignment, and fiber assembly during silk extrusion—a key evolutionary adaptation distinguishing spider silk architecture.[6]Diversity Across Species
Spiders, with over 53,000 described species worldwide as of November 2025, exhibit remarkable diversity in spidroin production, reflecting adaptations to varied ecological roles from web-building to hunting.[7] Comprehensive silkomics analyses, including a 2022 study sequencing transcriptomes from 1,098 species across 76 families, have identified more than 11,000 putative spidroin genes, underscoring the genetic basis for silk specialization.[8] In model orb-weaving species like Trichonephila clavipes, genomic sequencing has revealed at least 28 distinct spidroin genes, encoding proteins for multiple silk types such as dragline and capture spirals, with recent studies noting 10 major ampullate spidroin (MaSp) paralogs.[9][10] This diversity manifests in phylogenetic variations between major spider clades. Mygalomorph spiders, including tarantulas, typically possess simpler MaSp isoforms with fewer paralogs—estimated at around four duplications—resulting in less specialized silk repertoires suited to burrowing or sheet webs.[11] In contrast, araneomorph spiders, which dominate modern ecosystems, display extensive gene duplications—up to 10 or more events—enabling a broader array of specialized silks for complex behaviors like orb-weaving or cobweb construction.[11] These expansions, occurring post-divergence from mygalomorphs approximately 300 million years ago, have facilitated silk adaptations to diverse niches, such as the tensile draglines of aerial hunters versus the adhesive threads of ground-dwellers.[12] Evolutionary patterns reveal conserved structural elements amid sequence divergence, driving silk functionalization. Spidroin repetitive cores maintain a modular architecture of tandem motifs that provide mechanical versatility, yet their amino acid compositions diverge to tune properties like stiffness or extensibility across lineages.[13] Terminal domains, however, exhibit high conservation, with non-repetitive N- and C-terminal regions shared across spidroins to regulate solubility and assembly, while subtle variations contribute to clade-specific innovations.[13] This balance has enabled silk diversification, as seen in orb-weavers (Araneidae and Nephilidae) evolving elastic draglines for prey capture in flight paths, versus hunting spiders (Theridiidae) developing sticky threads for terrestrial ambushes.[8] Representative examples highlight these adaptations. In the black widow Latrodectus hesperus (Theridiidae), aggregate spidroin (AgSp) forms the glycoprotein glue coating gumfoot capture threads, providing adhesive properties for irregular cobwebs that ensnare ground prey.[14] Similarly, in Nephila species (Nephilidae), MaSp2 variants enriched with proline (up to ~9% content) in GPGXX motifs impart exceptional elasticity to dragline silks, allowing webs to withstand impacts from flying insects without breaking.[15]Molecular Structure
Primary Sequence and Motifs
Spidroins are large proteins typically ranging from 250 to 350 kDa in molecular weight, comprising approximately 3000 to 4000 amino acid residues.[16][17] Their primary sequences are dominated by a few key amino acids, with glycine accounting for 40-50% and alanine for 20-30% of the composition, alongside notable proportions of serine and proline.[18] These residues form the basis for the protein's repetitive architecture, which constitutes over 90% of the sequence and enables the material's unique properties. The core of spidroin sequences consists of extensive repetitive regions characterized by specific motifs that dictate local secondary structures. Poly-alanine blocks, denoted as (A)_n where n typically ranges from 3 to 9, promote the formation of β-sheet crystallites responsible for structural rigidity.[16] Glycine-proline-glycine motifs, such as GPGXX (e.g., GPGQQ where X represents glutamine or other residues), facilitate β-turns that contribute to flexibility.[19] Additionally, GGX motifs, with X as a variable amino acid, serve as amorphous spacers that maintain chain entropy and prevent excessive ordering.[20] Flanking these repetitive domains are non-repetitive N-terminal (NT) and C-terminal (CT) domains, which act as structural caps. The NT domain spans approximately 130-150 amino acids, while the CT domain is around 100 amino acids.[16] Sequence variability exists among spidroin types; for instance, major ampullate spidroin 1 (MaSp1) is enriched in alanine, enhancing tensile strength, whereas major ampullate spidroin 2 (MaSp2) incorporates higher levels of proline (about 15 mol%) and serine for greater elasticity.[15]Domains and Hierarchical Assembly
Spidroins exhibit a modular architecture featuring distinct N-terminal (NT) and C-terminal (CT) domains that flank a large repetitive core region, enabling controlled folding and assembly into hierarchical silk structures. The NT domain, approximately 14-15 kDa in size, adopts a compact five-helix bundle fold, as determined by NMR spectroscopy, which maintains solubility in the neutral pH environment of the silk gland storage compartment. At neutral pH (around 7-8), the NT exists as a stable monomer with a characteristic distribution of charged residues that prevents premature aggregation. Upon acidification to pH ~6.2 during extrusion, the NT undergoes rapid dimerization through hydrophobic interactions and hydrogen bonding at specific interfaces, such as the helix 2-3 loop, facilitating the initial alignment of spidroin monomers. This pH-triggered dimerization is further enhanced by shear forces in the spinning duct, promoting parallel orientation of the protein chains essential for subsequent fiber formation.[21] The CT domain, roughly 10-12 kDa, plays a complementary role in modulating assembly through pH-sensitive conformational dynamics. In solution at neutral pH, the CT forms a compact dimeric five-helix bundle structure, shielding the repetitive regions to inhibit aggregation.[1] As pH drops during spinning, a structural transition exposes hydrophobic surfaces and drives multimerization, including disulfide-linked dimers. A 2024 structural study using solid-state NMR, FTIR spectroscopy, and other methods on assembled fibers revealed that the CT undergoes an α-helix to β-sheet transition, integrating into the silk matrix with mixed secondary structures and partial unfolding, contributing to the transition from soluble precursors to insoluble fibrils. This conversion underscores the CT's function as a molecular switch, coordinating the timely alignment and polymerization of spidroins.[22] Hierarchical assembly of spidroins proceeds from individual monomer folding to supramolecular architectures, yielding the macroscopic silk fiber. At the molecular level, folded NT and CT domains anchor the repetitive sequences, which contain poly-alanine and glycine-rich motifs that enable secondary structure transitions. These repetitive regions primarily form crystalline β-sheet nanocrystals (typically 2-5 nm in cross-section), embedded within an amorphous matrix of random coils and α-helices, achieving an overall crystallinity of 30-50% that balances strength and elasticity. Multimers stabilized by NT-CT interactions further align into protofibrils or nanofibrils (2-5 nm diameter), as observed via cryo-TEM, which bundle into microfibrils (20-100 nm) and ultimately coalesce into macroscale fibers exhibiting a skin-core morphology, where the outer skin layer is more oriented and crystalline than the viscoelastic core. This multi-scale organization, driven by pH gradients, shear, and ionic shifts, ensures the efficient transformation of dilute spidroin solutions into robust silk threads.[23][24]Natural Biosynthesis
Genetic Expression
Spidroin genes form a multigene family in spider genomes, with multiple paralogs encoding proteins for specific silk types; for instance, major ampullate spidroins (MaSp) typically include 2-7 paralogs per species, such as at least three functional MaSp1 copies in black widow spiders and up to nine MaSp2 variants in orb-weavers like Argiope. These genes are often organized in genomic clusters, spanning 1-1.5 Mb and containing 10-14 MaSp loci in species such as Argiope aurantia, reflecting tandem duplications and gene conversion events that homogenize repetitive regions. Across spider genomes, the total number of spidroin genes ranges from approximately 20 to 35, as cataloged in species like Pardosa pseudoannulata (20 genes), Nephila clavipes (28 genes), and Larinioides sclopetarius (35 genes).[25][10][9][26] The transcriptional regulation of spidroin genes is governed by gland-specific promoters that restrict expression to epithelial cells within distinct silk glands, enabling the production of specialized silks. In orb-weaving spiders, these regulatory elements ensure that MaSp genes are predominantly active in major ampullate glands, while paralogs like flagelliform spidroins (Flag) are confined to flagelliform glands. Expression levels are modulated by environmental and physiological cues, including the spider's molting cycle and nutritional status; during post-molt phases, major ampullate glands exhibit heightened secretory activity with increased Type-M and Type-S granule production, while protein limitation from low feeding reduces proline and glutamine content in MaSp transcripts, shifting expression toward variants with altered amino acid motifs. Post-transcriptional modifications, such as alternative splicing, occur occasionally (e.g., in MaSp-f isoforms), but are generally minimal, preserving the repetitive structure of mature mRNA.[27][28][29][9] High-level transcription of spidroin mRNA occurs primarily in the secretory epithelium of ampullate glands, where single-cell RNA sequencing has identified specialized cell types zoned along the gland: Zone A cells express MaSp1 and MaSp2 at peak levels, Zone B cells produce MaSp3 and ampullate-like spidroins, and Zone C cells transcribe low-molecular-weight components like SpiCE-LMa. This zonation supports the assembly of complex silk dopes, with mRNA abundance correlating to protein output in epithelial cells rather than other tissues.[30] The evolutionary genetics of spidroins trace back to ancient duplication events approximately 240-300 million years ago, coinciding with or predating the estimated Mygalomorphae-Araneomorphae divergence (around 240-310 MYA based on recent phylogenomic analyses). Subsequent expansions, with at least 10 duplications in araneomorphs versus four in mygalomorphs, drove diversification tied to silk gland specialization. Recent phylogenomic analyses across Araneae, incorporating genomes from 28 species, reveal ongoing gene birth-death dynamics, with clade-specific expansions (e.g., MaSp2 in orb-weavers) and hox cluster restructuring mirroring spider diversification patterns.[11][31]Glandular Synthesis and Extrusion
Spider silk production occurs primarily in specialized abdominal glands, with the major ampullate glands serving as the key site for synthesizing the high-strength dragline silk used in lifelines and web frames. These glands, in species such as Argiope aurantia, consist of a tail region lined with secretory epithelium containing rough endoplasmic reticulum for protein synthesis, a central storage sac, a narrowing duct for processing, and an external spigot for extrusion.[32][33] The duct features distinct zones that sequentially dope the silk precursor with ions like Na⁺ and Cl⁻ in the initial segments to maintain solubility, followed by acid secretion to lower pH and facilitate structural transitions.[33][34] Spidroins, the primary structural proteins, are translated on ribosomes associated with the rough endoplasmic reticulum in the gland's epithelial cells of the tail zone before being packaged into secretory vesicles. These vesicles release the proteins into the glandular lumen, forming an aqueous spinning dope stored in the sac at concentrations of 30-50% spidroin by weight and a neutral pH of approximately 6.9.[33][17] As the dope moves into the duct, high shear forces from the narrowing geometry, combined with a pH drop to around 6.3 driven by proton secretion, trigger molecular alignment and partial folding of the spidroins; this process is mediated by pH-sensitive conformational changes in the N- and C-terminal domains, promoting dimerization and polymerization.[34][18][35] During extrusion, the converging geometry of the S-shaped duct induces extensional flow, aligning the spidroin molecules into a liquid-crystalline phase that enhances fiber orientation.[36] Rapid solidification follows as carbonic anhydrase enzymes in the duct generate CO₂ and H⁺, with CO₂ drawdown dehydrating the dope and stabilizing the nascent fiber at the spigot.[37] This coordinated mechanism ensures the dope transforms into a solid fiber under ambient conditions without external heat or solvents.[38] Variations in glandular processes adapt the dope for specific silk types; for instance, flagelliform glands, which are smaller with shorter ducts, produce a softer, more viscous dope rich in glycine-rich spidroins tailored for the highly elastic capture spirals in orb webs.[34][39] This contrasts with the firmer dope of major ampullate glands, highlighting gland-specific tuning of ion composition and flow dynamics to yield silks with distinct extensibility.[40]Physicochemical Properties
Mechanical Characteristics
Spidroin-based spider silks exhibit exceptional mechanical properties, particularly in tensile strength and toughness, arising from their protein composition. Major ampullate spidroin (MaSp) dragline silk typically displays a tensile strength of 1.0–1.7 GPa, comparable to high-tensile steel (1.5 GPa) but at approximately one-sixth the density, enabling superior strength-to-weight ratios.[2][41] Elasticity in these fibers reaches 20–30% strain before failure, allowing significant deformation without fracture.[2] This combination contributes to a toughness of 150–350 MJ/m³, surpassing synthetic materials like Kevlar (50 MJ/m³) and nylon (80 MJ/m³), with energy absorption facilitated by sliding and deformation within β-sheet nanocrystals in the hierarchical structure.[41][42] Type-specific variations highlight functional specialization in spidroin silks. MaSp dragline silk prioritizes high strength and moderate extensibility for load-bearing roles, achieving up to 1.65 GPa tensile strength in species like Caerostris darwini.[41] In contrast, flagelliform spidroin (Flag) silk demonstrates extreme extensibility of 300–600% strain at lower tensile strengths of 0.1–0.5 GPa, enabling effective energy capture through elastic recovery.[2][41] These properties stem from distinct spidroin isoforms, with MaSp featuring alanine-rich motifs for crystalline strength and Flag incorporating glycine-proline repeats for rubber-like behavior.[41] Mechanical characterization relies on micromechanical tensile assays using instruments like linear extensometers to measure stress-strain curves under controlled conditions.[43] Properties are influenced by extrusion parameters, such as drawing speed during silk production, which increases Young's modulus up to ~10 times and yield stress ~7 times at rates of 10–20 mm/s by enhancing molecular alignment and reducing crystallite size.[42] Humidity also modulates performance; relative humidity levels from 25% to 85% linearly increase breaking strain from approximately 20% to 35-40% while decreasing elastic modulus from 10-14 GPa to about 1.4 GPa, due to water-mediated weakening of hydrogen bonds in the protein matrix.[43]| Property | MaSp Dragline Silk | Flag Silk | Kevlar | Nylon |
|---|---|---|---|---|
| Tensile Strength (GPa) | 1.0–1.7 | 0.1–0.5 | 3.6 | 0.75–0.95 |
| Extensibility (% Strain) | 20–30 | 300–600 | 2.7 | 18 |
| Toughness (MJ/m³) | 150–350 | 27–283 | 50 | 80 |