Nucleoplasm
View on Wikipedia
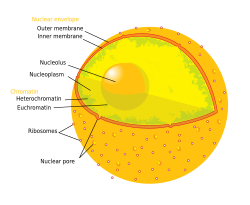
The nucleoplasm, also known as karyoplasm,[1] is the type of protoplasm that makes up the cell nucleus, the most prominent organelle of the eukaryotic cell. It is enclosed by the nuclear envelope, also known as the nuclear membrane.[2] The nucleoplasm resembles the cytoplasm of a eukaryotic cell in that it is a gel-like substance found within a membrane, although the nucleoplasm only fills out the space in the nucleus and has its own unique functions. The nucleoplasm suspends structures within the nucleus that are not membrane-bound and is responsible for maintaining the shape of the nucleus.[2] The structures suspended in the nucleoplasm include chromosomes, various proteins, nuclear bodies, the nucleolus, nucleoporins, nucleotides, and nuclear speckles.[2]
The soluble, liquid portion of the nucleoplasm is called the karyolymph[3] nucleosol,[4] or nuclear hyaloplasm.
History
[edit]
The existence of the nucleus, including the nucleoplasm, was first documented as early as 1682 by the Dutch microscopist Leeuwenhoek and was later described and drawn by Franz Bauer.[5] However, the cell nucleus was not named and described in detail until Robert Brown's presentation to the Linnean Society in 1831.[6] The nucleoplasm, while described by Bauer and Brown, was not specifically isolated as a separate entity until its naming in 1882 by Polish-German scientist Eduard Strasburger, one of the most famous botanists of the 19th century, and the first person to discover mitosis in plants.[7]
Role
[edit]Many important cell functions take place in the nucleus, more specifically in the nucleoplasm. The main function of the nucleoplasm is to provide the proper environment for essential processes that take place in the nucleus, serving as the suspension substance for all organelles inside the nucleus, and storing the structures that are used in these processes.[2] 34% of proteins encoded in the human genome are ones that localize to the nucleoplasm.[2] These proteins take part in RNA transcription and gene regulation in the nucleoplasm.[2] Proteins located in the nucleoplasm are involved in the activation of genes that are used in the cell cycle.[8] Some nucleoporins which typically make up the nuclear pore, can be mobile and participate in the regulation of gene expression in the nucleoplasm.[8][9] The nuclear pore is where molecules travel from inside the nucleoplasm to the cytoplasm and vice versa.[9] The nucleoplasm is also a route for many molecules to travel through.[9] Smaller molecules are able to pass freely through the nuclear pore to get into and out of the nucleoplasm, while larger proteins need the help of receptors on the surface of the nuclear envelope.[9] The nuclear matrix is also believed to be contained in the nucleoplasm where it functions to maintain the size and shape of the nucleus, in a role similar to that of the cytoskeleton found in the cytoplasm.[10] However, the existence and the exact function of the nuclear matrix remain unclear and heavily debated.
Composition
[edit]The nucleoplasm is a highly viscous liquid that is enveloped by the nuclear membrane and consists mainly of water, proteins, dissolved ions, and a variety of other substances including nucleic acids and minerals.
Proteins
[edit]There are around 20,000 protein-coding genes in humans,[11] and nearly a third of these have been found to localize to the nucleoplasm via targeting by a nuclear localization sequence (NLS).[2] Cytosolic proteins, known as importins, act as receptors for the NLS, escorting the protein to a nuclear pore complex to be transported into the nucleoplasm.[12] Proteins in the nucleoplasm are mainly tasked with participating in and regulating cellular functions that are DNA-dependent, including transcription, RNA splicing, DNA repair, DNA replication, and a variety of metabolic processes.[2] These proteins are divided into histone proteins, a class of proteins that bind to DNA and give chromosomes their shape and regulate gene activity,[13] and non-histone proteins.
The nucleoplasm contains many enzymes that are instrumental in the synthesis of DNA and RNA, including DNA polymerase and RNA polymerase which function in DNA replication and RNA transcription, respectively. Additionally, the nucleoplasm is host to many of the enzymes that play essential roles in cellular metabolism. NAD+ synthase is stored in the nucleoplasm and functions in electron transport and redox reactions involved with the electron transport chain and synthesis of adenosine triphosphate (ATP).[14] Pyruvate kinase is also found in the nucleoplasm in significant quantities; this enzyme is involved in the final step of glycolysis, catalyzing the conversion of phosphoenolpyruvate (PEP) to pyruvate along with the phosphorylation of adenosine diphosphate (ADP) to ATP.[15] Importantly, the nucleoplasm contains co-factors and co-enzymes, including acetyl-CoA, which plays a vital role in the citric acid cycle,[16] and ATP, which is involved in energy storage and transfer.
Ions
[edit]
The ionic composition of the nucleoplasm is crucial in maintaining homeostasis within the cell and the organism as a whole. Ions that have been documented in the nucleoplasm include sodium, potassium, calcium, phosphorus, and magnesium. These ions are key players in a variety of biological functions. Sodium and potassium play key roles in the sodium-potassium pump, a transmembrane ATPase that pumps three sodium ions out of the cell for every two potassium ions it pumps into the cell, creating an ionic gradient.[17] While this pump is generally considered to be a plasma membrane protein, its presence has been recorded in the nuclear envelope, controlling the ionic gradient between the cytoplasm and nucleoplasm of the cell and contributing to the homeostasis of calcium within the cell.[18] These ions also determine the concentration gradient that exists between the cytoplasm and nucleoplasm, serving to control the flow of ions across the nuclear envelope.[19] They are important in maintaining the osmolarity of the nucleoplasm which in turn provides structural integrity to the nuclear envelope as well as the organelles suspended in the dense nucleoplasm.
Similarity to cytoplasm
[edit]Nucleoplasm is quite similar to the cytoplasm, with the main difference being that nucleoplasm is found inside the nucleus while the cytoplasm is located inside the cell, outside of the nucleus. Their ionic compositions are nearly identical due to the ion pumps and permeability of the nuclear envelope, however, the proteins in these two fluids differ greatly. Proteins in the cytoplasm are termed cytosolic proteins which are produced by free ribosomes while proteins that localize to the nucleoplasm must undergo processing in the endoplasmic reticulum and golgi apparatus before being delivered to the nucleoplasm as part of the secretory pathway. These proteins also differ in function, as proteins that localize to the nucleoplasm are largely involved in DNA-dependent processes including cell division and gene regulation, while cytosolic proteins are mainly involved in protein modification, mRNA degradation, metabolic processes, signal transduction, and cell death.[20]
The cytoplasm and the nucleoplasm are both highly gelatinous structures enclosed by membranous structures- the plasma membrane and the nuclear envelope, respectively. However, while the cytoplasm is contained by a single lipid bilayer membrane, the nuclear envelope that compartmentalizes the nucleoplasm consists of two separate lipid bilayers- an outer membrane and an inner membrane.[21] The cytoplasm is also found in all known cells while nucleoplasm is only found in eukaryotic cells, as prokaryotic cells lack a well-defined nucleus and membrane-bound organelles. Additionally, during cell division, the cytoplasm divides during cytokinesis, while the nucleoplasm is released with the dissolution of the nuclear envelope, refilling only after the nuclear envelope reforms.
The organelles and other structures within the cytoplasm and nucleoplasm are organized by protein filaments within their respective compartments. The cytoplasm contains the cytoskeleton, a network of protein filaments found in all cells, while the nucleoplasm is believed to contain the nuclear matrix, a hypothetically analogous network of filaments that organizes the organelles and genetic information within the nucleus. While the structure and function of the cytoskeleton have been well documented, the exact function, and even the presence, of the nuclear matrix is disputed.[22] While the exact composition of the nuclear matrix has not been confirmed, type V intermediate filaments, known as nuclear lamins, have been documented in the nucleoplasm, functioning in the structural support of the nucleus as well as the regulation of DNA replication, transcription, and chromatin organization.[23] Cytoplasmic streaming, the circular flow of cytoplasm driven by the cytoskeleton, has been well documented in the cytoplasm, aiding in intracellular transport, but this process has not been documented in the nucleoplasm.
References
[edit]- ^ "karyoplasm". Collins English Dictionary. Retrieved 2 December 2022.
- ^ a b c d e f g h "The human cell in nucleoplasm". Human Protein Atlas. Archived from the original on 2024-09-15. Retrieved 2025-01-08.
- ^ "karyolymph". Collins English Dictionary. Retrieved 2 December 2022.
- ^ Kühn, T; Ihalainen, TO; Hyväluoma, J; Dross, N; Willman, SF; Langowski, J; Vihinen-Ranta, M; Timonen, J (2011). "Protein diffusion in mammalian cell cytoplasm". PLOS ONE. 6 (8) e22962. doi:10.1371/journal.pone.0022962. PMC 3158749. PMID 21886771.
- ^ Harris H (1999). The Birth of the Cell. New Haven: Yale University Press. ISBN 978-0-300-07384-3.
- ^ Brown R (1866). "On the Organs and Mode of Fecundation of Orchidex and Asclepiadea". Miscellaneous Botanical Works I: 511–514.
- ^ "Strasburger, Eduard Adolf". Universalium. 2010. Retrieved October 31, 2022.
- ^ a b Kalverda, Bernike; Pickersgill, Helen; Shloma, Victor V.; Fornerod, Maarten (2010). "Nucleoporins Directly Stimulate Expression of Developmental and Cell-Cycle Genes Inside the Nucleoplasm". Cell. 140 (3): 306–383. doi:10.1016/j.cell.2010.01.011. PMID 20144760. S2CID 17260209.
- ^ a b c d Khan, Asmat Ullah; Qu, Rongmei; Ouyang, Jun; Dai, Jingxing (2020-04-03). "Role of Nucleoporins and Transport Receptors in Cell Differentiation". Frontiers in Physiology. 11: 239. doi:10.3389/fphys.2020.00239. PMC 7145948. PMID 32308628.
- ^ Hed, Greer (October 6, 2022). "What is Nucleoplasm?". wisegeek. Retrieved October 28, 2022.
- ^ "Gene". www.genome.gov. Retrieved 7 January 2025.
- ^ Casem, Merri Lynn (2016). Case Studies in Cell Biology. Elsevier. pp. 73–103. ISBN 978-0-12-801394-6.
- ^ Stein; Thrall, C.L. (1973). "Evidence for the presence of nonhistone chromosomal proteins in the nucleoplasm of HeLa S3 cells". FEBS Letters. 32 (1): 41–45. doi:10.1016/0014-5793(73)80732-X. PMID 4715686. S2CID 20285491.
- ^ Houtkooper; Cantó, C.; Wanders, R.J.; Auwerx, J. (2010). "The Secret Life of NAD+: An Old Metabolite Controlling New Metabolic Signaling Pathways". Endocrine Reviews. 31 (2): 194–223. doi:10.1210/er.2009-0026. PMC 2852209. PMID 20007326.
- ^ Israelsen; Vander Heiden, M.G. (2015). "Pyruvate kinase: Function, regulation and role in cancer". Seminars in Cell & Developmental Biology. 43: 43–51. doi:10.1016/j.semcdb.2015.08.004. PMC 4662905. PMID 26277545.
- ^ Falcón; Chen, S.; Wood, M.S.; Aris, J.P. (2010). "Acetyl-coenzyme A synthetase 2 is a nuclear protein required for replicative longevity in Saccharomyces cerevisiae". Molecular and Cellular Biochemistry. 333 (1–2): 99–108. doi:10.1007/s11010-009-0209-z. PMC 3618671. PMID 19618123.
- ^ Goodsell, David (October 2009). "Molecule of the Month: Sodium-Potassium Pump". PDB-101. doi:10.2210/rcsb_pdb/mom_2009_10. Retrieved October 30, 2022.
- ^ Galva, Charitha; Artigas, Pablo; Gatto, Craig (December 2012). "Nuclear Na+/K+-ATPase plays an active role in nucleoplasmic Ca2+ homeostasis". Journal of Cell Science. 125 (24): 6137–6147. doi:10.1242/jcs.114959. PMC 3585523. PMID 23077175.
- ^ Wu, Yufei; Pegoraro, Adrian; Weitz, David; Janmey, Paul; Sun, Sean (February 2022). "The correlation between cell and nucleus size is explained by an eukaryotic cell growth model". PLOS Computational Biology. 18 (2) e1009400. doi:10.1371/journal.pcbi.1009400. PMC 8893647. PMID 35180215.
- ^ "The human cell in cytoplasm". Human Protein Atlas.
- ^ "Nuclear Membrane". National Human Genome Research Institute.
- ^ Pederson, Thoru (March 2000). "Half a Century of "The Nuclear Matrix"". Molecular Biology of the Cell. 11 (3): 799–805. doi:10.1091/mbc.11.3.799. PMC 14811. PMID 10712500.
- ^ Dechat, Thomas; Adam, Stephen A.; Taimen, Pekka; Shimi, Takeshi; Goldman, Robert D. (November 2010). "Nuclear Lamins". Cold Spring Harbor Perspectives in Biology. 2 (11) a000547. doi:10.1101/cshperspect.a000547. PMC 2964183. PMID 20826548.