Recent from talks
Nothing was collected or created yet.
Cytochrome b
View on Wikipedia| Cytochrome b, C-terminal domain | |||||||||
|---|---|---|---|---|---|---|---|---|---|
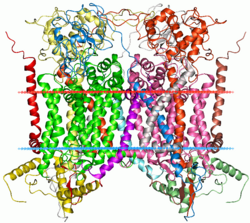 Mitochondrial cytochrome bc1 complex | |||||||||
| Identifiers | |||||||||
| Symbol | Cytochrom_B_C | ||||||||
| Pfam | PF00032 | ||||||||
| InterPro | IPR005798 | ||||||||
| |||||||||
Cytochrome b is a protein found in the membranes of aerobic cells. In eukaryotic mitochondria (inner membrane) and in aerobic prokaryotes, cytochrome b is a component of respiratory chain complex III (EC 1.10.2.2) — also known as the bc1 complex or ubiquinol-cytochrome c reductase. In plant chloroplasts and cyanobacteria, there is a homologous protein, cytochrome b6, a component of the plastoquinone-plastocyanin reductase (EC 1.10.99.1), also known as the b6f complex. These complexes are involved in electron transport, the pumping of protons to create a proton-motive force (PMF). This proton gradient is used for the generation of ATP. These complexes play a vital role in cells.[1][2][3]
Structure and function
[edit]Cytochrome b/b6 is an integral membrane protein of approximately 400 amino acid residues that probably has 8 transmembrane segments. In plants and cyanobacteria, cytochrome b6 consists of two protein subunits encoded by the petB and petD genes. Cytochrome b/b6 non-covalently binds two heme groups, known as b562 and b566. Four conserved histidine residues are postulated to be the ligands of the iron atoms of these two heme groups.[2][3]
The heme groups are key parts of the internal electron transfer pathway and indispensable to the functioning of the two quinol oxidizing complexes. Two units of b/b6 also form a quinol entry pathway.[4]
Use in phylogenetics
[edit]Cytochrome b is commonly used as a region of mitochondrial DNA for determining phylogenetic relationships between organisms, due to its sequence variability. It is considered to be most useful in determining relationships within families and genera. Comparative studies involving cytochrome b have resulted in new classification schemes and have been used to assign newly described species to a genus as well as to deepen the understanding of evolutionary relationships.[5]
Clinical significance
[edit]Mutations in cytochrome b primarily result in exercise intolerance in human patients; though more rare, severe multi-system pathologies have also been reported.[6]
Single-point mutations in cytochrome b of Plasmodium falciparum and P. berghei are associated with resistance to the anti-malarial drug atovaquone.[7]
Human genes
[edit]Human genes encoding cytochrome b proteins include:
Fungicide target
[edit]Cyt b is targeted by the QoI class of fungicides, Fungicide Resistance Action Committee group 11. The cyt b mutations G143A and F129L provide resistance against the main body of group 11, although G143A does not work against metyltetraprole (11A).[8] G143A is significant in Botrytis cinerea in California strawberry production.[9]
References
[edit]- ^ Blankenship, Robert (2009). Molecular Mechanisms of Photosynthesis. Blackwell Publishing. pp. 124–132.
- ^ a b Howell N (August 1989). "Evolutionary conservation of protein regions in the proton motive cytochrome b and their possible roles in redox catalysis". J. Mol. Evol. 29 (2): 157–69. Bibcode:1989JMolE..29..157H. doi:10.1007/BF02100114. PMID 2509716. S2CID 7298013.
- ^ a b Esposti MD, De Vries S, Crimi M, Ghelli A, Patarnello T, Meyer A (July 1993). "Mitochondrial cytochrome b: evolution and structure of the protein" (PDF). Biochim. Biophys. Acta. 1143 (3): 243–71. doi:10.1016/0005-2728(93)90197-N. PMID 8329437.
- ^ Sarewicz, M; Pintscher, S; Pietras, R; Borek, A; Bujnowicz, Ł; Hanke, G; Cramer, WA; Finazzi, G; Osyczka, A (24 February 2021). "Catalytic Reactions and Energy Conservation in the Cytochrome bc(1) and b(6)f Complexes of Energy-Transducing Membranes". Chemical Reviews. 121 (4): 2020–2108. doi:10.1021/acs.chemrev.0c00712. PMC 7908018. PMID 33464892.
- ^ Castresana, J. (2001). "Cytochrome b Phylogeny and the Taxonomy of Great Apes and Mammals". Molecular Biology and Evolution. 18 (4): 465–471. doi:10.1093/oxfordjournals.molbev.a003825. PMID 11264397.
- ^ Blakely EL, Mitchell AL, Fisher N, Meunier B, Nijtmans LG, Schaefer AM, Jackson MJ, Turnbull DM, Taylor RW (July 2005). "A mitochondrial cytochrome b mutation causing severe respiratory chain enzyme deficiency in humans and yeast". FEBS J. 272 (14): 3583–92. doi:10.1111/j.1742-4658.2005.04779.x. PMID 16008558. S2CID 13938075.
- ^ Siregar JE, Syafruddin D, Matsuoka H, Kita K, Marzuki S (June 2008). "Mutation underlying resistance of Plasmodium berghei to atovaquone in the quinone binding domain 2 (Qo(2)) of the cytochrome b gene". Parasitology International. 57 (2): 229–32. doi:10.1016/j.parint.2007.12.002. PMID 18248769.
- ^ FRAC (Fungicide Resistance Action Committee) (March 2021). "FRAC Code List ©*2021: Fungal control agents sorted by cross resistance pattern and mode of action (including coding for FRAC Groups on product labels)" (PDF). Archived from the original (PDF) on 2021-11-05. Retrieved 2021-06-16.
- ^
- Petrasch, Stefan; Knapp, Steven J.; van Kan, Jan A. L.; Blanco-Ulate, Barbara (2019-04-04). "Grey mould of strawberry, a devastating disease caused by the ubiquitous necrotrophic fungal pathogen Botrytis cinerea". Molecular Plant Pathology. 20 (6). British Society for Plant Pathology (W-B): 877–892. Bibcode:2019MolPP..20..877P. doi:10.1111/mpp.12794. ISSN 1464-6722. PMC 6637890. PMID 30945788. S2CID 93002697.
- Cosseboom, Scott D.; Ivors, Kelly L.; Schnabel, Guido; Bryson, Patricia K.; Holmes, Gerald J. (2019). "Within-Season Shift in Fungicide Resistance Profiles of Botrytis cinerea in California Strawberry Fields". Plant Disease. 103 (1). American Phytopathological Society: 59–64. Bibcode:2019PlDis.103...59C. doi:10.1094/pdis-03-18-0406-re. ISSN 0191-2917. PMID 30422743. S2CID 205345358.
External links
[edit]- Cytochromes+b at the U.S. National Library of Medicine Medical Subject Headings (MeSH)

