Recent from talks
Nothing was collected or created yet.
Mucin
View on Wikipedia
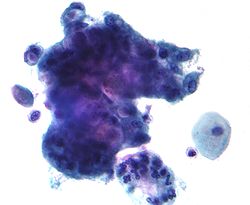 | |
| Identifiers | |
|---|---|
| Symbol | Mucin |
| Membranome | 111 |
Mucins (/ˈmjuːsɪn/) are a family of high molecular weight, heavily glycosylated proteins (glycoconjugates) produced by epithelial tissues in most animals.[1] Mucins' key characteristic is their ability to form gels; therefore they are a key component in most gel-like secretions, serving functions from lubrication to cell signalling to forming chemical barriers.[1] They often take an inhibitory role.[1] Some mucins are associated with controlling mineralization, including nacre formation in mollusks,[2] calcification in echinoderms[3] and bone formation in vertebrates.[4] They bind to pathogens as part of the immune system. Overexpression of the mucin proteins, especially MUC1, is associated with many types of cancer.[5][6]
Although some mucins are membrane-bound due to the presence of a hydrophobic membrane-spanning domain that favors retention in the plasma membrane, most mucins are secreted as principal components of mucus by mucous membranes or are secreted to become a component of saliva.
Genes and proteins
[edit]Human mucins include genes with the HUGO symbol MUC 1 through 22. Of these mucins, the following classes have been defined by localization:[7][8][9][10]
- Secreted mucins in humans, with their chromosomal location, repeat size in amino acids (aa), whether they are gel-forming (Y) or not (N), and their tissue expression.[11]
| Mucin | gel | chromosome | repeat size (aa) | tissue expression |
|---|---|---|---|---|
| MUC2 | Y | 11p15.5 | 23 | Jejunum, ileum, colon, endometrium |
| MUC5A | Y | 11p15.5 | 8 | Respiratory tract, stomach, conjunctiva, endocervix, endometrium |
| MUC5B | Y | 11p15.5 | 29 | Respiratory tract, submandibular glands, endocervix |
| MUC6 | Y | 11p15.5 | 169 | Stomach, ileum, gall bladder, endocervix, endometrium |
| MUC19 | Y | 12q12 | 19 | corneal and conjunctival epithelia; lacrimal gland[12] |
| MUC7 | N | 4q13–q21 | 23 | Sublingual and submandibular glands |
| MUC8 | N | 12q24.3 | 13/41 | Respiratory tract, uterus, endocervix, endometrium |
| MUC9 | N | 1p13 | 15 | Fallopian tubes |
| MUC20 | N | 3 | 19 | kidney (high), moderately in placenta, lung, prostate, liver, digestive system |
- Membrane-bound (transmembrane) mucins: MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC21 (formerly C6orf205), MUC22 (highly polymorphic[13])
The major secreted airway mucins are MUC5AC and MUC5B, while MUC2 is secreted mostly in the intestine but also in the airway. MUC7 is the major salivary protein.[10]
Protein structure
[edit]Mature mammalian mucins are composed of two distinct regions:[7]
- The amino- and carboxy-terminal regions are very lightly glycosylated, but rich in cysteines. The cysteine residues participate in establishing disulfide linkages within and among mucin monomers.
- A large central region ("PTS domain") formed of multiple tandem repeats of 10 to 80 residue sequences in which up to half of the amino acids are serine or threonine. This area becomes saturated with hundreds of O-linked oligosaccharides. N-linked oligosaccharides are also found on mucins, but in less abundance than O-linked sugars.
Evolutionary classification
[edit]The functional classification does not correspond to an exact evolutionary relationship, which is still incomplete and ongoing.[10] Known-related groups include:
- The gel-forming mucins (2, 5AC, 5B, 6, 19) are related both to each other and to otogelin and von Willebrand Factor (PTHR11339).[14] Four of these occur in a well-conserved gene cluster (at 11p15.5 in humans).[15]
- The EGF-like domain containing mucins. These include MUC3(A,B), MUC4, MUC12, MUC13, and MUC17.[16]
- Some EGF-like mucins, plus MUC1 and MUC16, carry SEA domains, a vertebrate invention. It is unclear whether this points to a common origin among these transmembrane mucins.[14]
- MUC21 and MUC22 are related to each other by sharing a C-terminal domain (PF14654). They also occur in a human gene cluster on 6p21.33.
- MUC7 is a recent invention in placental mammals. It started as a copy in the secretory calcium-binding phosphoprotein (SCPP) gene cluster and rapidly gained PTS repeats.[17]
Function in humans
[edit]Mucins have been found to have important functions in defense against bacterial and fungal infections. MUC5B, the predominant mucin in the mouth and female genital tract, has been shown to significantly reduce attachment and biofilm formation of Streptococcus mutans, a bacterium with the potential to form cavities.[18] Unusually, MUC5B does not kill the bacteria but rather maintains it in the planktonic (non-biofilm) phase, thus maintaining a diverse and healthy oral microbiome.[18] Similar effects of MUC5B and other mucins have been demonstrated with other pathogens, such as Candida albicans, Helicobacter pylori, and even HIV.[19][20] In the mouth, mucins can also recruit anti-microbial proteins such as statherins and histatine 1, which further reduces risk of infection.[20]
Eleven mucins are expressed by the eye surface epithelia, goblet cells and associated glands, even though most of them are expressed at very low levels. They maintain wetness, lubricate the blink, stabilize the tear film, and create a physical barrier to the outside world.[12]
Glycosylation and aggregation
[edit]Mucin genes encode mucin monomers that are synthesized as rod-shaped apomucin cores that are post-translationally modified by exceptionally abundant glycosylation.
The dense "sugar coating" of mucins gives them considerable water-holding capacity and also makes them resistant to proteolysis, which may be important in maintaining mucosal barriers.
Mucins are secreted as massive aggregates of proteins with molecular masses of roughly 1 to 10 million Da. Within these aggregates, monomers are linked to one another mostly by non-covalent interactions, although intermolecular disulfide bonds may also play a role in this process.
Secretion
[edit]Upon stimulation, MARCKS (myristylated alanine-rich C kinase substrate) protein coordinates the secretion of mucin from mucin-filled vesicles within the specialized epithelial cells.[21] Fusion of the vesicles to the plasma membrane causes release of the mucin, which as it exchanges Ca2+ for Na+ expands up to 600 fold. The result is a viscoelastic product of interwoven molecules which, combined with other secretions (e.g., from the airway epithelium and the submucosal glands in the respiratory system), is called mucus.[22][23]
Clinical significance
[edit]Increased mucin production occurs in many adenocarcinomas, including cancers of the pancreas, lung, breast, ovary, colon and other tissues. Mucins are also overexpressed in lung diseases such as asthma, bronchitis, chronic obstructive pulmonary disease (COPD) or cystic fibrosis.[24] Two membrane mucins, MUC1 and MUC4 have been extensively studied in relation to their pathological implication in the disease process.[25][26][27] Mucins are under investigation as possible diagnostic markers for malignancies and other disease processes in which they are most commonly over- or mis-expressed.
Abnormal deposits of mucin are responsible for the non-pitting facial edema seen in untreated hypothyroidism. This edema is seen in the pretibial area as well.[28][page needed]
Non-vertebrate mucins
[edit]Beyond the better-studied vertebrate mucins, other animals also express (not necessarily related) proteins with similar properties. These include:
- Drosophila is known to express mucin proteins containing PTS-rich repeats.[29]
- Trypanosoma cruzi express cell-surface mucins (Pfam PF01456).[30]
Some other organisms produce mucilage that does not have a protein component, only polysacchides.
Cosmetic use
[edit]Misuse of skincare products containing snail secretions of mucin have resulted in pain, swelling, and oozing.[31][32] Counterfeit versions of a Korean snail mucin product called COSRX have been selling online, putting users at risk.[33]
See also
[edit]References
[edit]- ^ a b c Lang T, Klasson S, Larsson E, Johansson ME, Hansson GC, Samuelsson T (2016). "Searching the Evolutionary Origin of Epithelial Mucus Protein Components—Mucins and FCGBP". Molecular Biology and Evolution. 33 (8). Oxford Academic: 1921–1936. doi:10.1093/molbev/msw066. PMC 4948705. PMID 27189557.
- ^ Marin F, Corstjens P, de Gaulejac B, de Vrind-De Jong E, Westbroek P (July 2000). "Mucins and molluscan calcification. Molecular characterization of mucoperlin, a novel mucin-like protein from the nacreous shell layer of the fan mussel Pinna nobilis (Bivalvia, pteriomorphia)". The Journal of Biological Chemistry. 275 (27): 20667–20675. doi:10.1074/jbc.M003006200. hdl:1887/50061. PMID 10770949.
- ^ Boskey AL (2003). "Biomineralization: an overview". Connective Tissue Research. 44 Suppl 1 (1): 5–9. doi:10.1080/713713622. PMID 12952166.
- ^ Midura RJ, Hascall VC (October 1996). "Bone sialoprotein--a mucin in disguise?". Glycobiology. 6 (7): 677–681. doi:10.1093/glycob/6.7.677. PMID 8953277.
- ^ Niv Y (April 2008). "MUC1 and colorectal cancer pathophysiology considerations". World Journal of Gastroenterology. 14 (14): 2139–2141. doi:10.3748/wjg.14.2139. PMC 2703837. PMID 18407586.
- ^ Brockhausen I, Melamed J (August 2021). "Mucins as anti-cancer targets: perspectives of the glycobiologist". Glycoconjugate Journal. 38 (4): 459–474. doi:10.1007/s10719-021-09986-8. PMID 33704667. S2CID 232191632.
- ^ a b Moniaux N, Escande F, Porchet N, Aubert JP, Batra SK (October 2001). "Structural organization and classification of the human mucin genes". Frontiers in Bioscience. 6: D1192 – D1206. doi:10.2741/moniaux. PMID 11578969.
- ^ Perez-Vilar J, Hill RL (2004). "Mucin Family of Glycoproteins". In Lennarz WJ, Lane MD (eds.). Encyclopedia of Biological Chemistry. Vol. 2. Oxford: Academic Press/Elsevier. pp. 758–764. doi:10.1016/B0-12-443710-9/00411-7. ISBN 978-0-12-443710-4.
- ^ Hoorens PR, Rinaldi M, Li RW, Goddeeris B, Claerebout E, Vercruysse J, et al. (March 2011). "Genome wide analysis of the bovine mucin genes and their gastrointestinal transcription profile". BMC Genomics. 12 140. doi:10.1186/1471-2164-12-140. PMC 3056801. PMID 21385362.
- ^ a b c Kasprzak A, Adamek A (March 2019). "Mucins: the Old, the New and the Promising Factors in Hepatobiliary Carcinogenesis". International Journal of Molecular Sciences. 20 (6): 1288. doi:10.3390/ijms20061288. PMC 6471604. PMID 30875782.
- ^ Corfield AP (1 January 2015). "Mucins: A biologically relevant glycan barrier in mucosal protection". Biochimica et Biophysica Acta (BBA) - General Subjects. 1850 (1): 236–252. doi:10.1016/j.bbagen.2014.05.003. ISSN 0304-4165. PMID 24821013.
- ^ a b Martinez-Carrasco R, Argüeso P, Fini ME (1 July 2021). "Membrane-associated mucins of the human ocular surface in health and disease". The Ocular Surface. 21: 313–330. doi:10.1016/j.jtos.2021.03.003. ISSN 1542-0124. PMC 8328898. PMID 33775913.
- ^ Norman PJ, Norberg SJ, Guethlein LA, Nemat-Gorgani N, Royce T, Wroblewski EE, et al. (May 2017). "Sequences of 95 human MHC haplotypes reveal extreme coding variation in genes other than highly polymorphic HLA class I and II". Genome Research. 27 (5): 813–823. doi:10.1101/gr.213538.116. PMC 5411776. PMID 28360230.
{{cite journal}}: CS1 maint: overridden setting (link) - ^ a b Lang T, Hansson GC, Samuelsson T (October 2007). "Gel-forming mucins appeared early in metazoan evolution". Proceedings of the National Academy of Sciences of the United States of America. 104 (41): 16209–16214. Bibcode:2007PNAS..10416209L. doi:10.1073/pnas.0705984104. PMC 2042186. PMID 17911254.
- ^ Lang T, Klasson S, Larsson E, Johansson ME, Hansson GC, Samuelsson T (August 2016). "Searching the Evolutionary Origin of Epithelial Mucus Protein Components-Mucins and FCGBP". Molecular Biology and Evolution. 33 (8): 1921–1936. doi:10.1093/molbev/msw066. PMC 4948705. PMID 27189557.
- ^ Liberelle M, Jonckheere N, Melnyk P, Van Seuningen I, Lebègue N (May 2020). "EGF-Containing Membrane-Bound Mucins: A Hidden ErbB2 Targeting Pathway?". Journal of Medicinal Chemistry. 63 (10): 5074–5088. doi:10.1021/acs.jmedchem.9b02001. PMID 32027502. S2CID 211044898.
- ^ Xu D, Pavlidis P, Thamadilok S, Redwood E, Fox S, Blekhman R, et al. (August 2016). "Recent evolution of the salivary mucin MUC7". Scientific Reports. 6 (1) 31791. Bibcode:2016NatSR...631791X. doi:10.1038/srep31791. PMC 4997351. PMID 27558399.
{{cite journal}}: CS1 maint: overridden setting (link) - ^ a b Frenkel ES, Ribbeck K (January 2015). "Salivary mucins protect surfaces from colonization by cariogenic bacteria". Applied and Environmental Microbiology. 81 (1): 332–338. Bibcode:2015ApEnM..81..332F. doi:10.1128/aem.02573-14. PMC 4272720. PMID 25344244.
- ^ Kavanaugh NL, Zhang AQ, Nobile CJ, Johnson AD, Ribbeck K (November 2014). Berman J (ed.). "Mucins suppress virulence traits of Candida albicans". mBio. 5 (6) e01911-14: e01911. doi:10.1128/mBio.01911-14. PMC 4235211. PMID 25389175.
- ^ a b Frenkel ES, Ribbeck K (January 2015). "Salivary mucins in host defense and disease prevention". Journal of Oral Microbiology. 7 (1) 29759. doi:10.3402/jom.v7.29759. PMC 4689954. PMID 26701274.
- ^ Li Y, Martin LD, Spizz G, Adler KB (November 2001). "MARCKS protein is a key molecule regulating mucin secretion by human airway epithelial cells in vitro". The Journal of Biological Chemistry. 276 (44): 40982–40990. doi:10.1074/jbc.M105614200. PMID 11533058.
- ^ Rogers DF (September 2007). "Physiology of airway mucus secretion and pathophysiology of hypersecretion". Respiratory Care. 52 (9): 1134–46, discussion 1146–9. PMID 17716382.
- ^ Perez-Vilar J (February 2007). "Mucin granule intraluminal organization". American Journal of Respiratory Cell and Molecular Biology. 36 (2): 183–190. doi:10.1165/rcmb.2006-0291TR. PMC 2176109. PMID 16960124.
- ^ Morrison CB, Markovetz MR, Ehre C (November 2019). "Mucus, mucins, and cystic fibrosis". Pediatric Pulmonology. 54 (Suppl 3): S84 – S96. doi:10.1002/ppul.24530. PMC 6853602. PMID 31715083.
- ^ Singh AP, Moniaux N, Chauhan SC, Meza JL, Batra SK (January 2004). "Inhibition of MUC4 expression suppresses pancreatic tumor cell growth and metastasis". Cancer Research. 64 (2): 622–630. doi:10.1158/0008-5472.CAN-03-2636. PMID 14744777.
- ^ Singh AP, Chauhan SC, Bafna S, Johansson SL, Smith LM, Moniaux N, et al. (March 2006). "Aberrant expression of transmembrane mucins, MUC1 and MUC4, in human prostate carcinomas". The Prostate. 66 (4): 421–429. doi:10.1002/pros.20372. PMID 16302265. S2CID 21904013.
{{cite journal}}: CS1 maint: overridden setting (link) - ^ Singh AP, Chaturvedi P, Batra SK (January 2007). "Emerging roles of MUC4 in cancer: a novel target for diagnosis and therapy". Cancer Research. 67 (2): 433–436. doi:10.1158/0008-5472.CAN-06-3114. PMID 17234748.
- ^ Black JM, Hawk JH (2009). Medical Surgical Nursing: clinical management for positive outcomes. Elsevier India. ISBN 978-81-312-2982-8.
- ^ Syed ZA, Härd T, Uv A, van Dijk-Härd IF (August 2008). "A potential role for Drosophila mucins in development and physiology". PLOS ONE. 3 (8) e3041. Bibcode:2008PLoSO...3.3041S. doi:10.1371/journal.pone.0003041. PMC 2515642. PMID 18725942.
- ^ Cámara ML, Balouz V, Centeno Cameán C, Cori CR, Kashiwagi GA, Gil SA, et al. (May 2019). "Trypanosoma cruzi surface mucins are involved in the attachment to the Triatoma infestans rectal ampoule". PLOS Neglected Tropical Diseases. 13 (5) e0007418. doi:10.1371/journal.pntd.0007418. PMC 6544316. PMID 31107901.
{{cite journal}}: CS1 maint: overridden setting (link) - ^ McCoy K, Class MM, Ricles V, Wagoner G, Cross D, Trautz A, et al. (2024). "Kids These Days: Social Media's Influence on Adolescent Behaviors". The Journal of Clinical and Aesthetic Dermatology. 17 (5): 40–42. PMC 11107899. PMID 38779370.
- ^ Singh N, Brown AN, Gold MH (2024). "Snail extract for skin: A review of uses, projections, and limitations". Journal of Cosmetic Dermatology. 23 (4): 1113–1121. doi:10.1111/jocd.16269. PMID 38429932.
- ^ Mull A (17 June 2024). "Online Shopping Has Become a Giant Fake Product Machine". Businessweek. Retrieved 28 June 2024.
Further reading
[edit]- Ali MS, Hutton DA, Wilson JA, Pearson JP (September 2005). "Major secretory mucin expression in chronic sinusitis". Otolaryngology–Head and Neck Surgery. 133 (3): 423–428. doi:10.1016/j.otohns.2005.06.005. PMID 16143194. S2CID 42482788.
- Ramsey KA, Rushton ZL, Ehre C (June 2016). "Mucin Agarose Gel Electrophoresis: Western Blotting for High-molecular-weight Glycoproteins". Journal of Visualized Experiments. 112 (112) 54153. doi:10.3791/54153. PMC 4927784. PMID 27341489.
External links
[edit]- Mucins at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- "Mucin" at Dorland's Medical Dictionary
Mucin
View on GrokipediaOverview and Classification
Definition and Properties
Mucins are high-molecular-weight glycoproteins that serve as the primary structural components of mucus, forming protective barriers on epithelial surfaces throughout the body. They are produced by specialized goblet cells and other epithelial cells, and are encoded by a family of at least 21 MUC genes recognized by the Human Genome Organization. Mucins can be classified into two main categories: secreted gel-forming mucins, such as MUC2 and MUC5AC, which contribute to the viscoelastic mucus gel, and cell-surface or transmembrane mucins, such as MUC1 and MUC4, which extend from the apical surface of epithelial cells to form the glycocalyx.[1][3][4] Structurally, mucins feature a central protein backbone characterized by tandemly repeated amino acid sequences rich in proline, threonine, and serine (PTS domains), which provide sites for dense O-linked glycosylation. These O-glycans, initiated by the attachment of N-acetylgalactosamine (GalNAc) to serine or threonine residues, constitute 50-80% of the molecule's mass and form branched oligosaccharide chains, resulting in a characteristic "bottlebrush" or extended rod-like conformation. The protein core also includes cysteine-rich domains at the N- and C-termini that facilitate polymerization through disulfide bond formation in secreted mucins, leading to large multimers with molecular weights often exceeding several million Daltons.[5][1][3] The biochemical properties of mucins arise primarily from their extensive glycosylation, which imparts high hydrophilicity, water-binding capacity, and resistance to proteolysis, enabling the formation of hydrated gels. These gels exhibit viscoelastic behavior, balancing elasticity and viscosity to provide lubrication, hydration, and mechanical protection against shear forces and pathogens. Glycan diversity, including cores such as Core 1 (Galβ1-3GalNAc) and Core 2, often capped with sialic acid or fucose, further modulates charge, solubility, and interactions with microbes or signaling molecules.[4][5][3]Types and Nomenclature
Mucins are high-molecular-weight glycoproteins classified primarily into two major categories based on their structural features and cellular localization: secreted mucins and membrane-bound (transmembrane) mucins.[6] This classification reflects their distinct roles in forming protective mucus layers or anchoring to cell surfaces, respectively. Secreted mucins are further subdivided into gel-forming (oligomeric) types, which polymerize to create viscous barriers, and non-gel-forming (soluble or low-molecular-weight) types, which contribute monomeric or smaller oligomeric structures.[7] Membrane-bound mucins, in contrast, possess a transmembrane domain that tethers them to the plasma membrane, often extending heavily glycosylated ectodomains into the extracellular space.[6] The gel-forming secreted mucins include MUC2, MUC5AC, MUC5B, MUC6, and MUC19, which are characterized by cysteine-rich domains facilitating dimerization and subsequent multimerization via disulfide bonds and linker regions. Four of these (MUC2, MUC5AC, MUC5B, MUC6) cluster genetically on chromosome 11p15.5 (with MUC19 on 12q12), underscoring their evolutionary relatedness.[7][8] For instance, MUC2 predominates in the intestinal tract, forming the primary component of the colonic mucus layer, while MUC5AC and MUC5B are major constituents of airway and gastric mucus, respectively.[4] Non-gel-forming secreted mucins, such as MUC7 and MUC8, lack extensive polymerization domains and are typically monomeric or form small oligomers; MUC7, for example, is prominent in salivary secretions.[7] Membrane-bound mucins encompass MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC20, and MUC21, each featuring a single-pass transmembrane helix and variable cytoplasmic tails for signaling functions.[9] These are distributed across multiple chromosomes, with notable clustering on 7q22 (MUC3A/B, MUC12, MUC17) and 3q29 (MUC4, MUC20). MUC1, the archetypal member, is ubiquitously expressed on epithelial surfaces and plays roles in cell adhesion and signaling, whereas MUC16 (also known as CA-125) is highly expressed in ovarian tissues.[6] Some genes, like MUC14 (now EMCN, endomucin) and MUC9 (OVGP1, oviductal glycoprotein 1), exhibit atypical mucin features and are sometimes considered peripheral to the core family, while MUC22 remains poorly characterized.[9]| Category | Examples | Key Structural Features | Chromosomal Locations |
|---|---|---|---|
| Gel-forming secreted | MUC2, MUC5AC, MUC5B, MUC6, MUC19 | Cysteine-rich subdomains for polymerization; von Willebrand factor-like domains | 11p15.5 (most) |
| Non-gel-forming secreted | MUC7, MUC8 | Histatins or smaller repeat regions; no extensive oligomers | Varied (e.g., 4q13.3 for MUC7) |
| Membrane-bound | MUC1, MUC3A/B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC20, MUC21 | Transmembrane domain; SEA module (in most); EGF-like motifs (in some) | Multiple (e.g., 1q22 for MUC1, 19p13.2 for MUC16) |
Molecular Biology
Encoding Genes
Mucins are encoded by a family of genes designated as MUC genes in humans, with 21 members identified by the HUGO Gene Nomenclature Committee.[9] These genes were numbered sequentially based on the chronological order of their discovery, starting from MUC1 in the late 1980s.[7] The MUC gene family exhibits significant structural diversity, but a common feature across most members is the presence of one or more large central exons containing variable number tandem repeats (VNTRs). These VNTRs encode proline-, serine-, and threonine-rich peptide domains that serve as scaffolds for extensive O-linked glycosylation, which constitutes the hallmark of mucin proteins.[7] The length and sequence variability in these tandem repeat regions contribute to polymorphism and functional diversity among mucins.[10] The MUC genes are broadly classified into two functional categories based on the structure of their encoded proteins: secreted mucins and membrane-bound (transmembrane) mucins. Secreted mucins, which form gel-like protective barriers on mucosal surfaces, include the gel-forming subtypes MUC2, MUC5AC, MUC5B, MUC6, and MUC19, as well as the non-gel-forming MUC7 and MUC8. Membrane-bound mucins, such as MUC1, MUC3A/B, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC20, and MUC21, feature a hydrophobic transmembrane domain and a short cytoplasmic tail, enabling cell surface association and roles in signaling and adhesion.[9] Additional genes like MUC9 (OVGP1), MUC14 (EMCN), and MUC22 have less defined classifications but share mucin-like domains. This dichotomy reflects evolutionary adaptations, with secreted forms emphasizing lubrication and pathogen trapping, while membrane forms facilitate cell-cell interactions.[7] Chromosomally, the MUC genes are dispersed across multiple loci, with notable clustering observed for the major gel-forming secreted mucins. MUC2, MUC5AC, MUC5B, and MUC6 are tightly linked within a 400-kb gene cluster on chromosome 11p15.5, oriented in the order MUC6-MUC2-MUC5AC-MUC5B. This genomic organization suggests coordinated regulation and shared evolutionary origins from ancient tandem duplications. Other membrane-bound genes are scattered, such as MUC1 on 1q22, MUC4 and MUC20 on 3q29, and MUC16 on 19p13.2, reflecting independent evolutionary histories.[9] The cluster on 11p15.5 is particularly significant, as polymorphisms in these genes, including VNTR length variations, influence mucin production and susceptibility to respiratory and gastrointestinal diseases.| Gene | Type | Chromosomal Location | Key Features |
|---|---|---|---|
| MUC1 | Membrane-bound | 1q22 | VNTR of 20-125 repeats encoding 60-bp units; involved in cell signaling.[7] |
| MUC2 | Secreted (gel-forming) | 11p15.5 | Large central exon with two VNTR regions (48-bp and 69-bp repeats); predominant in intestinal mucus.[11] |
| MUC3A/B | Membrane-bound | 7q22.1 | Paired genes with large tandem repeats; expressed in gastrointestinal epithelia.[9] |
| MUC4 | Membrane-bound | 3q29 | No VNTR but extensive EGF-like domains; largest mucin gene (~25 kb).[7] |
| MUC5AC | Secreted (gel-forming) | 11p15.5 | VNTR of 24-123 repeats (525-bp units); key in airway and gastric mucus.[7] |
| MUC5B | Secreted (gel-forming) | 11p15.5 | VNTR of 21-38 repeats (507-bp units); primary mucin in airway secretions. |
| MUC6 | Secreted (gel-forming) | 11p15.5 | VNTR of ~12 repeats (507-bp units); stomach-specific protective role. |
| MUC7 | Secreted (non-gel) | 4q13.3 | Short VNTR of 6-9 repeats (69-bp units); salivary mucin.[7] |
| MUC16 | Membrane-bound | 19p13.2 | Extremely large (~14,000 aa) with ~156 SEA modules; ovarian cancer marker.[9] |
| MUC20 | Membrane-bound | 3q29 | Smaller size; kidney and colon expression.[9] |
Protein Architecture
Mucins are high-molecular-weight glycoproteins defined by their modular protein backbones, which feature densely O-glycosylated regions that confer extended, rigid structures essential for mucus formation and cellular protection. The core architecture consists of a signal peptide for secretion or membrane targeting, flanked by N- and C-terminal domains that mediate oligomerization and interactions, with a central proline-, serine-, and threonine-rich (PST) domain serving as the primary site of O-linked glycosylation. This glycosylation, often comprising 70-90% of the molecule's mass, creates a bottle-brush-like conformation that extends the protein up to 100-1000 nm in length, providing steric hindrance and lubrication.[12][13] Secreted, gel-forming mucins such as MUC2, MUC5AC, MUC5B, MUC6, and MUC19 exhibit a conserved domain organization optimized for polymerization into viscoelastic networks. The N-terminus includes multiple von Willebrand factor D (VWD) domains (e.g., VWD1-4) and cysteine-knot (CK) motifs that facilitate N-terminal trimerization and C-terminal dimerization through disulfide bonds, enabling the formation of linear multimers that cross-link via transglutaminase activity. The central mucin domain comprises variable number tandem repeats (VNTRs) of 10-30 amino acids rich in serines and threonines, where dense O-glycosylation with short, sialylated or sulfated glycans imparts rigidity and negative charge, crucial for gel expansion upon secretion. C-terminal cysteine-rich domains (CysD) further stabilize non-covalent interactions, while the overall architecture results in proteins exceeding 2 MDa, as seen in MUC2's ~5,000-residue backbone. Seminal studies on recombinant domain expression have elucidated these assembly mechanisms, confirming VWD domains' role in initial oligomerization.[12][14] In contrast, membrane-bound mucins like MUC1, MUC4, MUC12, MUC13, and MUC16 integrate into the plasma membrane, forming a glycocalyx that modulates cell signaling and adhesion. These proteins share an extracellular N-terminal region with a SEA (sea urchin sperm protein, enterokinase, agrin) domain, which undergoes autocatalytic cleavage to generate α and β subunits held by non-covalent bonds, and a VNTR-rich mucin domain analogous to secreted forms but shorter (e.g., 20-100 repeats in MUC1). The C-terminus features a single transmembrane helix anchoring the protein, followed by a short cytoplasmic tail (10-70 residues) containing phosphorylation sites for intracellular signaling via kinases like PKC. MUC4 uniquely incorporates three EGF-like domains in its extracellular region, enabling interactions with receptor tyrosine kinases such as ErbB2 to promote anti-adhesive and proliferative signals. O-glycosylation in these mucins is sparser and more variable, often truncated in cancers, altering the extended structure to expose protein epitopes. Structural analyses, including those of recombinant SEA domains, highlight the cleavage site's conservation across species, underscoring its role in subunit maturation.[15][16]Evolutionary Perspectives
Mucins, particularly the gel-forming subtypes, originated early in metazoan evolution, with proteins exhibiting characteristic von Willebrand factor D (VWD), VWD C8 (VWE), and TIL domains identified in the cnidarian starlet sea anemone Nematostella vectensis.[17] These structural modules suggest that ancestral gel-forming mucins evolved as protective adaptations in basal animals, predating the divergence of major metazoan lineages. Gel-forming mucins are also present in non-vertebrate chordates such as the sea squirt Ciona intestinalis and the lancelet Branchiostoma floridae, indicating broad conservation across invertebrates and early vertebrates.[17] In vertebrates, the number of gel-forming mucin genes varies significantly, reflecting lineage-specific expansions. Humans possess five such genes (MUC2, MUC5AC, MUC5B, MUC6, and MUC19), while teleost fishes like zebrafish (Danio rerio) and pufferfish (Takifugu rubripes) have only one identifiable MUC2 ortholog each.[17] A notable expansion occurs in amphibians, where the frog Xenopus tropicalis encodes at least 25 gel-forming mucins, including 16 MUC2 homologs and nine MUC5-type proteins, likely driven by gene duplication events that enhanced mucus production in moist environments.[17] Membrane-bound mucins, in contrast, show distinct evolutionary trajectories; for instance, human MUC1 is mammal-specific and derived from a heparin sulfate proteoglycan ancestor via acquisition of a SEA domain, while MUC4 and MUC16 evolved from separate progenitors involving NIDO, AMOP, VWD, and multiple SEA domains, respectively, with no close sequence similarity beyond shared motifs.[18] In mammals, mucin evolution frequently involves the co-option of non-mucin, proline-rich precursor proteins that acquire proline-threonine-serine (PTS)-rich exonic repeats, enabling O-glycosylation and gel-forming properties—a process termed "mucinization."[19] This mechanism accounts for 15 independent, lineage-specific events, explaining the origin of all 28 mucins in the secretory calcium-binding phosphoprotein (SCPP) locus.[19] A representative example is the rodent-specific MUC10, which arose from the proline-rich protein Prol1 through tandem repeat expansions (up to 42 copies in some species) and shifts in expression to salivary glands, adapting to dietary and pathogenic pressures.[19] Such rapid diversification via repeat insertions allows mucins to evolve novel functions without whole-gene duplications, highlighting a key innovation in mammalian mucus adaptation. Recent genomic studies have revealed that the MUC19 gene in some modern human populations carries haplotypes introgressed from Denisovans, suggesting adaptive advantages in mucus properties.[20][19]Biochemistry
Glycosylation Mechanisms
Mucin-type O-glycosylation represents the predominant post-translational modification in mucins, characterized by the dense attachment of O-linked glycans to serine and threonine residues within the protein's proline-, threonine-, and serine-rich (PTS) domains.[21] This process occurs primarily in the Golgi apparatus and is essential for the structural expansion, solubility, and protective functions of mucins, with up to 80-90% of amino acids in these domains being glycosylated.[22] The initiation step involves the transfer of N-acetylgalactosamine (GalNAc) from UDP-GalNAc to the hydroxyl group of serine or threonine, forming the Tn antigen (GalNAcα1-O-Ser/Thr), catalyzed by a family of over 20 polypeptide N-acetylgalactosaminyltransferases (ppGalNAcTs or GALNTs).[21] These enzymes exhibit isoform-specific substrate preferences and tissue distribution, with examples like GALNT1 implicated in ovarian cancer progression and GALNT12 in colorectal carcinoma.[21] Following initiation in the cis-Golgi, the Tn antigen undergoes elongation to form one of several core structures, primarily cores 1 through 8, though cores 1-4 predominate in mucins. Core 1 (T antigen, Galβ1-3GalNAcα1-O-Ser/Thr) is synthesized by the T-synthase enzyme (C1GALT1), which requires the molecular chaperone COSMC to maintain its activity and prevent degradation; mutations or silencing of COSMC lead to persistent Tn expression and are associated with aberrant mucin glycosylation in diseases.[23] Alternative cores include core 2 (GlcNAcβ1-6(Galβ1-3)GalNAcα1-O-Ser/Thr), formed from core 1 by core 2 β1,6-N-acetylglucosaminyltransferases (C2GnT1-3); core 3 (GlcNAcβ1-3GalNAcα1-O-Ser/Thr), generated directly from Tn by β1,3-N-acetylglucosaminyltransferase 6 (C3GnT6); and core 4, an extension of core 3 with an additional GlcNAc branch via C2/4GnT.[21] These core formations occur in the medial Golgi and diversify the glycan repertoire, with core 1 being the most common in secretory mucins like MUC2 and MUC5AC.[23] Subsequent extension and modification in the trans-Golgi and trans-Golgi network involve the addition of monosaccharides such as galactose, N-acetylglucosamine, sialic acid, and fucose, mediated by a cascade of glycosyltransferases. For instance, poly-N-acetyllactosamine (poly-LacNAc) chains are built by alternating β1,4-galactosyltransferases (β4GalTs) and β1,3-N-acetylglucosaminyltransferases (β3GnTs), while sialylation—critical for mucin charge and viscosity—occurs via sialyltransferases like ST6GalNAc-I, which adds α2,6-linked sialic acid to Tn to form sialyl-Tn (STn) antigen.[21] Fucosylation by fucosyltransferases introduces Lewis antigens (e.g., Lewis A or X), enhancing mucin interactions with lectins and pathogens.[22] In mucins, these extensions create heterogeneous, branched glycan trees that contribute to the gel-forming properties, with the degree of sialylation and sulfation influencing mucus rheology and barrier function.[23] The following table summarizes the primary core structures in mucin O-glycosylation, highlighting key enzymes and their linkages:| Core | Structure | Initiating Enzyme | Linkage from Tn or Precursor | Prevalence in Mucins |
|---|---|---|---|---|
| 1 (T) | Galβ1-3GalNAcα1-O-Ser/Thr | C1GALT1 (with COSMC) | β1-3 Gal to Tn | High (e.g., MUC1, MUC5AC)[21] |
| 2 | GlcNAcβ1-6(Galβ1-3)GalNAcα1-O-Ser/Thr | C2GnT1-3 | β1-6 GlcNAc to Core 1 | Moderate (branching in MUC2)[21] |
| 3 | GlcNAcβ1-3GalNAcα1-O-Ser/Thr | C3GnT6 | β1-3 GlcNAc to Tn | Variable (gastric mucins)[23] |
| 4 | GlcNAcβ1-3(GlcNAcβ1-6)GalNAcα1-O-Ser/Thr | C2/4GnT | β1-6 GlcNAc to Core 3 | Low (respiratory mucins)[21] |

