Recent from talks
Nothing was collected or created yet.
Protein production
View on Wikipedia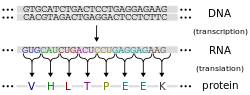
Protein production is the biotechnological process of generating a specific protein. It is typically achieved by the manipulation of gene expression in an organism such that it expresses large amounts of a recombinant gene. This includes the transcription of the recombinant DNA to messenger RNA (mRNA), the translation of mRNA into polypeptide chains, which are ultimately folded into functional proteins and may be targeted to specific subcellular or extracellular locations.[1]
Protein production systems (also known as expression systems) are used in the life sciences, biotechnology, and medicine. Molecular biology research uses numerous proteins and enzymes, many of which are from expression systems; particularly DNA polymerase for PCR, reverse transcriptase for RNA analysis, restriction endonucleases for cloning, and to make proteins that are screened in drug discovery as biological targets or as potential drugs themselves. There are also significant applications for expression systems in industrial fermentation, notably the production of biopharmaceuticals such as human insulin to treat diabetes, and to manufacture enzymes.
Protein production systems
[edit]Commonly used protein production systems include those derived from bacteria,[2][3] yeast,[4][5] baculovirus/insect,[6] mammalian cells,[7][8] and more recently filamentous fungi such as Myceliophthora thermophila.[9] When biopharmaceuticals are produced with one of these systems, process-related impurities termed host cell proteins also arrive in the final product in trace amounts.[10]
Cell-based systems
[edit]This article includes a list of general references, but it lacks sufficient corresponding inline citations. (January 2024) |
The oldest and most widely used expression systems are cell-based and may be defined as the "combination of an expression vector, its cloned DNA, and the host for the vector that provide a context to allow foreign gene function in a host cell, that is, produce proteins at a high level".[11][12] Overexpression is an abnormally and excessively high level of gene expression which produces a pronounced gene-related phenotype.[13][14][clarification needed]
There are many ways to introduce foreign DNA to a cell for expression, and many different host cells may be used for expression — each expression system has distinct advantages and liabilities. Expression systems are normally referred to by the host and the DNA source or the delivery mechanism for the genetic material. For example, common hosts are bacteria (such as E. coli, B. subtilis), yeast (such as S. cerevisiae[5]) or eukaryotic cell lines. Common DNA sources and delivery mechanisms are viruses (such as baculovirus, retrovirus, adenovirus), plasmids, artificial chromosomes and bacteriophage (such as lambda). The best expression system depends on the gene involved, for example the Saccharomyces cerevisiae is often preferred for proteins that require significant posttranslational modification. Insect or mammal cell lines are used when human-like splicing of mRNA is required. Nonetheless, bacterial expression has the advantage of easily producing large amounts of protein, which is required for X-ray crystallography or nuclear magnetic resonance experiments for structure determination.
Because bacteria are prokaryotes, they are not equipped with the full enzymatic machinery to accomplish the required post-translational modifications or molecular folding. Hence, multi-domain eukaryotic proteins expressed in bacteria often are non-functional. Also, many proteins become insoluble as inclusion bodies that are difficult to recover without harsh denaturants and subsequent cumbersome protein-refolding.
To address these concerns, expressions systems using multiple eukaryotic cells were developed for applications requiring the proteins be conformed as in, or closer to eukaryotic organisms: cells of plants (i.e. tobacco), of insects or mammalians (i.e. bovines) are transfected with genes and cultured in suspension and even as tissues or whole organisms, to produce fully folded proteins. Mammalian in vivo expression systems have however low yield and other limitations (time-consuming, toxicity to host cells,..). To combine the high yield/productivity and scalable protein features of bacteria and yeast, and advanced epigenetic features of plants, insects and mammalians systems, other protein production systems are developed using unicellular eukaryotes (i.e. non-pathogenic 'Leishmania' cells).
Bacterial systems
[edit]Escherichia coli
[edit]
E. coli is one of the most widely used expression hosts, and DNA is normally introduced in a plasmid expression vector. The techniques for overexpression in E. coli are well developed and work by increasing the number of copies of the gene or increasing the binding strength of the promoter region so assisting transcription.[3]
For example, a DNA sequence for a protein of interest could be cloned or subcloned into a high copy-number plasmid containing the lac (often LacUV5) promoter, which is then transformed into the bacterium E. coli. Addition of IPTG (a lactose analog) activates the lac promoter and causes the bacteria to express the protein of interest.[2]
E. coli strain BL21 and BL21(DE3) are two strains commonly used for protein production. As members of the B lineage, they lack lon and OmpT proteases, protecting the produced proteins from degradation. The DE3 prophage found in BL21(DE3) provides T7 RNA polymerase (driven by the LacUV5 promoter), allowing for vectors with the T7 promoter to be used instead.[15]
Corynebacterium
[edit]Non-pathogenic species of the gram-positive Corynebacterium are used for the commercial production of various amino acids. The C. glutamicum species is widely used for producing glutamate and lysine,[16] components of human food, animal feed and pharmaceutical products.
Expression of functionally active human epidermal growth factor has been done in C. glutamicum,[17] thus demonstrating a potential for industrial-scale production of human proteins. Expressed proteins can be targeted for secretion through either the general, secretory pathway (Sec) or the twin-arginine translocation pathway (Tat).[18]
Unlike gram-negative bacteria, the gram-positive Corynebacterium lack lipopolysaccharides that function as antigenic endotoxins in humans.[citation needed]
Pseudomonas fluorescens
[edit]The non-pathogenic and gram-negative bacteria, Pseudomonas fluorescens, is used for high level production of recombinant proteins; commonly for the development bio-therapeutics and vaccines. P. fluorescens is a metabolically versatile organism, allowing for high throughput screening and rapid development of complex proteins. P. fluorescens is most well known for its ability to rapid and successfully produce high titers of active, soluble protein.[19]
Eukaryotic systems
[edit]Yeasts
[edit]Expression systems using either S. cerevisiae or Pichia pastoris allow stable and lasting production of proteins that are processed similarly to mammalian cells, at high yield, in chemically defined media of proteins.[4][5]
Filamentous fungi
[edit]Filamentous fungi, especially Aspergillus and Trichoderma, have long been used to produce diverse industrial enzymes from their own genomes ("native", "homologous") and from recombinant DNA ("heterologous").[9]
More recently, Myceliophthora thermophila C1 has been developed into an expression platform for screening and production of native and heterologous proteins.The expression system C1 shows a low viscosity morphology in submerged culture, enabling the use of complex growth and production media. C1 also does not "hyperglycosylate" heterologous proteins, as Aspergillus and Trichoderma tend to do.[9]
Baculovirus-infected cells
[edit]Baculovirus-infected insect cells[20] (Sf9, Sf21, High Five strains) or mammalian cells[21] (HeLa, HEK 293) allow production of glycosylated or membrane proteins that cannot be produced using fungal or bacterial systems.[20][6] It is useful for production of proteins in high quantity. Genes are not expressed continuously because infected host cells eventually lyse and die during each infection cycle.[22]
Non-lytic insect cell expression
[edit]Non-lytic insect cell expression is an alternative to the lytic baculovirus expression system. In non-lytic expression, vectors are transiently or stably transfected into the chromosomal DNA of insect cells for subsequent gene expression.[23][24] This is followed by selection and screening of recombinant clones.[25] The non-lytic system has been used to give higher protein yield and quicker expression of recombinant genes compared to baculovirus-infected cell expression.[24] Cell lines used for this system include: Sf9, Sf21 from Spodoptera frugiperda cells, Hi-5 from Trichoplusia ni cells, and Schneider 2 cells and Schneider 3 cells from Drosophila melanogaster cells.[23][25] With this system, cells do not lyse and several cultivation modes can be used.[23] Additionally, protein production runs are reproducible.[23][24] This system gives a homogeneous product.[24] A drawback of this system is the requirement of an additional screening step for selecting viable clones.[25]
Leishmania tarentolae (cannot infect mammals) expression systems allow stable and lasting production of proteins at high yield, in chemically defined media. Produced proteins exhibit fully eukaryotic post-translational modifications, including glycosylation and disulfide bond formation.[citation needed]
Mammalian systems
[edit]The most common mammalian expression systems are Chinese Hamster ovary (CHO) and Human embryonic kidney (HEK) cells.[26][27][28]
- Chinese hamster ovary cell[27]
- Mouse myeloma lymphoblstoid (e.g. NS0 cell)[26]
- Fully Human
- Human embryonic kidney cells (HEK-293)[27]
- Human embryonic retinal cells (Crucell's Per.C6)[27]
- Human amniocyte cells (Glycotope and CEVEC)[citation needed]
Cell-free systems
[edit]Cell-free production of proteins is performed in vitro using purified RNA polymerase, ribosomes, tRNA and ribonucleotides. These reagents may be produced by extraction from cells or from a cell-based expression system. Due to the low expression levels and high cost of cell-free systems, cell-based systems are more widely used.[29]
See also
[edit]References
[edit]- ^ Gräslund S, Nordlund P, Weigelt J, Hallberg BM, Bray J, Gileadi O, et al. (February 2008). "Protein production and purification". Nature Methods. 5 (2): 135–46. doi:10.1038/nmeth.f.202. PMC 3178102. PMID 18235434.
- ^ a b Baneyx F (October 1999). "Recombinant protein expression in Escherichia coli". Current Opinion in Biotechnology. 10 (5): 411–21. doi:10.1016/s0958-1669(99)00003-8. PMID 10508629.
- ^ a b Rosano, Germán; Ceccarelli, Eduardo (2014-04-17). "Recombinant protein expression in Escherichia coli: advances and challenges". Frontiers in Microbiology. 5: 172. doi:10.3389/fmicb.2014.00172. PMC 4029002. PMID 24860555.
- ^ a b Cregg JM, Cereghino JL, Shi J, Higgins DR (September 2000). "Recombinant protein expression in Pichia pastoris". Molecular Biotechnology. 16 (1): 23–52. doi:10.1385/MB:16:1:23. PMID 11098467. S2CID 35874864.
- ^ a b c Malys N, Wishart JA, Oliver SG, McCarthy JE (2011). "Protein production in Saccharomyces cerevisiae for systems biology studies". Methods in Systems Biology. Methods in Enzymology. Vol. 500. pp. 197–212. doi:10.1016/B978-0-12-385118-5.00011-6. ISBN 9780123851185. PMID 21943899.
- ^ a b Kost TA, Condreay JP, Jarvis DL (May 2005). "Baculovirus as versatile vectors for protein expression in insect and mammalian cells". Nature Biotechnology. 23 (5): 567–75. doi:10.1038/nbt1095. PMC 3610534. PMID 15877075.
- ^ Rosser MP, Xia W, Hartsell S, McCaman M, Zhu Y, Wang S, Harvey S, Bringmann P, Cobb RR (April 2005). "Transient transfection of CHO-K1-S using serum-free medium in suspension: a rapid mammalian protein expression system". Protein Expression and Purification. 40 (2): 237–43. doi:10.1016/j.pep.2004.07.015. PMID 15766864.
- ^ Lackner A, Genta K, Koppensteiner H, Herbacek I, Holzmann K, Spiegl-Kreinecker S, Berger W, Grusch M (September 2008). "A bicistronic baculovirus vector for transient and stable protein expression in mammalian cells". Analytical Biochemistry. 380 (1): 146–8. doi:10.1016/j.ab.2008.05.020. PMID 18541133.
- ^ a b c Visser H, Joosten V, Punt PJ, Gusakov AV, Olson PT, Joosten R, et al. (June 2011). "Development of a mature fungal technology and production platform for industrial enzymes based on a Myceliophthora thermophila isolate, previously known as Chrysosporium lucknowense C1". Industrial Biotechnology. 7 (3): 214–223. doi:10.1089/ind.2011.7.214.
Aspergillus and Trichoderma are currently the main fungal genera used to produce industrial enzymes.
- ^ Wang, Xing; Hunter, Alan K.; Mozier, Ned M. (2009-06-15). "Host cell proteins in biologics development: Identification, quantitation and risk assessment". Biotechnology and Bioengineering. 103 (3): 446–458. doi:10.1002/bit.22304. ISSN 0006-3592. PMID 19388135. S2CID 22707536.
- ^ "Definition: expression system". Online Medical Dictionary. Centre for Cancer Education, University of Newcastle upon Tyne: Cancerweb. 1997-11-13. Retrieved 2008-06-10.
- ^ "Expression system - definition". Biology Online. Biology-Online.org. 2005-10-03. Retrieved 2008-06-10.
- ^ "overexpression". Oxford Living Dictionary. Oxford University Press. 2017. Archived from the original on February 10, 2018. Retrieved 18 May 2017.
The production of abnormally large amounts of a substance which is coded for by a particular gene or group of genes; the appearance in the phenotype to an abnormally high degree of a character or effect attributed to a particular gene.
- ^ "overexpress". NCI Dictionary of Cancer Terms. National Cancer Institute at the National Institutes of Health. 2011-02-02. Retrieved 18 May 2017.
overexpress
In biology, to make too many copies of a protein or other substance. Overexpression of certain proteins or other substances may play a role in cancer development. - ^ Jeong, H; Barbe, V; Lee, CH; Vallenet, D; Yu, DS; Choi, SH; Couloux, A; Lee, SW; Yoon, SH; Cattolico, L; Hur, CG; Park, HS; Ségurens, B; Kim, SC; Oh, TK; Lenski, RE; Studier, FW; Daegelen, P; Kim, JF (11 December 2009). "Genome sequences of Escherichia coli B strains REL606 and BL21(DE3)". Journal of Molecular Biology. 394 (4): 644–52. doi:10.1016/j.jmb.2009.09.052. PMID 19786035.
- ^ Brinkrolf K, Schröder J, Pühler A, Tauch A (September 2010). "The transcriptional regulatory repertoire of Corynebacterium glutamicum: reconstruction of the network controlling pathways involved in lysine and glutamate production". Journal of Biotechnology. 149 (3): 173–82. doi:10.1016/j.jbiotec.2009.12.004. PMID 19963020.
- ^ Date M, Itaya H, Matsui H, Kikuchi Y (January 2006). "Secretion of human epidermal growth factor by Corynebacterium glutamicum". Letters in Applied Microbiology. 42 (1): 66–70. doi:10.1111/j.1472-765x.2005.01802.x. PMID 16411922.
- ^ Meissner D, Vollstedt A, van Dijl JM, Freudl R (September 2007). "Comparative analysis of twin-arginine (Tat)-dependent protein secretion of a heterologous model protein (GFP) in three different Gram-positive bacteria". Applied Microbiology and Biotechnology. 76 (3): 633–42. doi:10.1007/s00253-007-0934-8. PMID 17453196. S2CID 6238466.
- ^ Retallack DM, Jin H, Chew L (February 2012). "Reliable protein production in a Pseudomonas fluorescens expression system". Protein Expression and Purification. 81 (2): 157–65. doi:10.1016/j.pep.2011.09.010. PMID 21968453.
- ^ a b Altmann F, Staudacher E, Wilson IB, März L (February 1999). "Insect cells as hosts for the expression of recombinant glycoproteins". Glycoconjugate Journal. 16 (2): 109–23. doi:10.1023/A:1026488408951. PMID 10612411. S2CID 34863069.
- ^ Kost TA, Condreay JP (October 1999). "Recombinant baculoviruses as expression vectors for insect and mammalian cells". Current Opinion in Biotechnology. 10 (5): 428–33. doi:10.1016/S0958-1669(99)00005-1. PMID 10508635.
- ^ Yin J, Li G, Ren X, Herrler G (January 2007). "Select what you need: a comparative evaluation of the advantages and limitations of frequently used expression systems for foreign genes". Journal of Biotechnology. 127 (3): 335–47. doi:10.1016/j.jbiotec.2006.07.012. PMID 16959350.
- ^ a b c d Dyring, Charlotte (2011). "Optimising the drosophila S2 expression system for production of therapeutic vaccines". BioProcessing Journal. 10 (2): 28–35. doi:10.12665/j102.dyring.
- ^ a b c d Olczak M, Olczak T (December 2006). "Comparison of different signal peptides for protein secretion in nonlytic insect cell system". Analytical Biochemistry. 359 (1): 45–53. doi:10.1016/j.ab.2006.09.003. PMID 17046707.
- ^ a b c McCarroll L, King LA (October 1997). "Stable insect cell cultures for recombinant protein production". Current Opinion in Biotechnology. 8 (5): 590–4. doi:10.1016/s0958-1669(97)80034-1. PMID 9353223.
- ^ a b Zhu J (2012-09-01). "Mammalian cell protein expression for biopharmaceutical production". Biotechnology Advances. 30 (5): 1158–70. doi:10.1016/j.biotechadv.2011.08.022. PMID 21968146.
- ^ a b c d Almo SC, Love JD (June 2014). "Better and faster: improvements and optimization for mammalian recombinant protein production". Current Opinion in Structural Biology. New constructs and expression of proteins / Sequences and topology. 26: 39–43. doi:10.1016/j.sbi.2014.03.006. PMC 4766836. PMID 24721463.
- ^ Hacker DL, Balasubramanian S (June 2016). "Recombinant protein production from stable mammalian cell lines and pools". Current Opinion in Structural Biology. New constructs and expression of proteins • Sequences and topology. 38: 129–36. doi:10.1016/j.sbi.2016.06.005. PMID 27322762.
- ^ Rosenblum G, Cooperman BS (January 2014). "Engine out of the chassis: cell-free protein synthesis and its uses". FEBS Letters. 588 (2): 261–8. Bibcode:2014FEBSL.588..261R. doi:10.1016/j.febslet.2013.10.016. PMC 4133780. PMID 24161673.
Further reading
[edit]- Higgins SJ, Hames BD (1999). Protein Expression: A Practical Approach. Oxford University Press. ISBN 978-0-19-963623-5.
- Baneyx, François (2004). Protein Expression Technologies: Current Status and Future Trends. Garland Science. ISBN 978-0-9545232-5-1.
External links
[edit]Protein production
View on GrokipediaNatural Protein Production
Prokaryotic Protein Synthesis
Prokaryotic protein synthesis is a rapid and efficient process that couples transcription and translation directly in the cytoplasm, allowing ribosomes to begin translating nascent mRNA while it is still being synthesized by RNA polymerase, without the compartmentalization imposed by a nucleus.[6] This coupling enhances gene expression speed and coordination primarily in bacteria, with archaea also lacking a nucleus and exhibiting coupled transcription-translation but featuring initiation mechanisms more similar to eukaryotes, including the use of unformylated Met-tRNA^Met^ and archaeal initiation factors (aIF1, aIF1A, aIF2) homologous to eukaryotic eIFs.[7] In bacteria, this coupling enables quick responses to environmental changes.[8] Transcription in prokaryotes begins with the binding of RNA polymerase holoenzyme, which includes a core enzyme and a sigma (σ) factor, to the promoter region of the DNA. The sigma factor recognizes specific consensus sequences, such as the -10 box (TATAAT) and -35 box (TTGACA), facilitating unwinding of the DNA double helix at the promoter to form an open complex. Initiation occurs when the first nucleotide is incorporated, followed by elongation where RNA polymerase moves along the template strand at a rate of about 50 nucleotides per second, synthesizing a complementary mRNA strand. Termination happens at specific signals, either through rho-independent hairpin loops in the mRNA or rho-dependent helicase activity that dissociates the polymerase. Translation commences shortly after transcription initiation due to the coupling, with the 70S ribosome assembling on the mRNA. First, tRNAs are charged by aminoacyl-tRNA synthetases, which attach specific amino acids to their cognate tRNAs using ATP.[9] In bacteria, initiation involves the 30S ribosomal subunit binding to the Shine-Dalgarno sequence upstream of the start codon (AUG), followed by recruitment of fMet-tRNA^fMet^ (N-formylmethionine-tRNA) in the P site and the 50S subunit to form the complete 70S ribosome, aided by initiation factors IF1, IF2, and IF3.[9] During elongation, aminoacyl-tRNA enters the A site via elongation factor Tu (EF-Tu), and peptidyl transferase, a ribozyme within the 23S rRNA of the 50S subunit, catalyzes peptide bond formation by transferring the growing polypeptide from the P-site tRNA to the A-site amino acid; translocation then shifts the mRNA and tRNAs via EF-G, exposing the next codon.[9] Termination occurs when a stop codon (UAA, UAG, or UGA) enters the A site, triggering release factors RF1 or RF2 to hydrolyze the peptidyl-tRNA bond, followed by ribosome recycling with EF-G and RRF.[9] A hallmark of prokaryotic mRNA is its polycistronic nature, where a single transcript can encode multiple proteins from consecutive genes, often organized into operons for coordinated expression.[2] Unlike eukaryotic mRNA, prokaryotic transcripts lack introns and require no splicing, streamlining the process.[10] The 70S ribosomes facilitate high synthesis rates, incorporating up to 20 amino acids per second per ribosome.[2] For instance, the lac operon in Escherichia coli exemplifies operon structure, with structural genes (lacZ, lacY, lacA) regulated by a promoter, operator, and the lacI repressor gene; in the presence of lactose, allolactose binds the repressor, derepressing transcription to produce β-galactosidase, lactose permease, and transacetylase for lactose metabolism. Sigma factors, such as σ^70^ in E. coli, play a crucial role in promoter recognition, with alternative sigmas enabling responses to stress or development.Eukaryotic Protein Synthesis
Eukaryotic protein synthesis is a compartmentalized process separated between the nucleus and cytoplasm, enabling extensive regulation and quality control that contrasts with the coupled transcription-translation in prokaryotes. Unlike prokaryotes, where these steps occur simultaneously in the cytoplasm, the nuclear envelope in eukaryotes isolates transcription from translation, allowing for intricate mRNA processing before export. This separation supports the production of diverse, functional proteins in multicellular organisms through slower but highly regulated mechanisms, typically synthesizing proteins at rates of about 5 amino acids per second. Eukaryotic mRNAs are monocistronic, encoding a single protein, which facilitates precise control over individual gene expression. Transcription in eukaryotes is primarily mediated by RNA polymerase II (Pol II), which initiates at promoter regions enhanced by upstream enhancers and transcription factors. The process begins with the assembly of the pre-initiation complex at the TATA box or other core promoters, followed by Pol II elongation. Co-transcriptionally, the nascent pre-mRNA undergoes 5' capping by the addition of a 7-methylguanosine cap shortly after initiation, which protects the mRNA and aids in later export and translation. At the 3' end, polyadenylation involves cleavage and addition of a poly(A) tail by the cleavage and polyadenylation specificity factor (CPSF) complex, stabilizing the mRNA. Introns are removed via splicing by the spliceosome, a large ribonucleoprotein complex that recognizes splice sites and joins exons, ensuring mature mRNA formation. Processed mRNAs are exported from the nucleus through nuclear pore complexes via the NXF1/NXT1 (TAP/p15) heterodimer, which binds the mRNA and interacts with nucleoporins for translocation. Quality control mechanisms, such as nonsense-mediated decay (NMD), degrade mRNAs with premature termination codons during or after export, preventing production of truncated proteins; NMD involves factors like UPF1, UPF2, and UPF3 that recognize aberrant stop codons during pioneer translation rounds. In the cytoplasm, translation occurs on 80S ribosomes composed of 40S and 60S subunits. Initiation requires eukaryotic initiation factors (eIFs), including eIF4F for cap recognition, eIF2 for Met-tRNA delivery, and eIF1/eIF1A for accuracy; the 43S pre-initiation complex scans from the 5' cap to the start AUG codon in a 5'-to-3' direction. Elongation proceeds with eEF1A delivering aminoacyl-tRNAs and eEF2 facilitating translocation, while termination involves eRF1/eRF3 recognizing stop codons and releasing the polypeptide. Post-translational modifications (PTMs) in eukaryotes occur primarily in the endoplasmic reticulum (ER), Golgi apparatus, and cytoplasm, diversifying protein function and localization. Glycosylation, a key ER/Golgi process, includes N-linked addition of oligosaccharides to asparagine residues in the ER lumen and O-linked to serines/threonines in the Golgi, influencing protein folding, stability, and trafficking. Phosphorylation by kinases on serines, threonines, or tyrosines regulates activity, as seen in signaling pathways. Ubiquitination tags proteins with ubiquitin chains via E1-E2-E3 enzymes, directing them to proteasomal degradation or altering interactions. Protein folding is assisted by chaperones like Hsp70, which bind nascent chains to prevent aggregation, and in the ER, protein disulfide isomerase (PDI) forms disulfide bonds. The Golgi apparatus further modifies and sorts proteins into vesicles for secretion or membrane insertion, ensuring proper cellular distribution.Recombinant Protein Production Systems
Bacterial Expression Systems
Bacterial expression systems, particularly those utilizing Escherichia coli, serve as cost-effective and rapid platforms for producing recombinant proteins, leveraging the bacterium's fast growth rate and simple genetics to achieve high yields of non-glycosylated proteins that do not require complex post-translational modifications (PTMs).[11] These systems emerged in the 1970s with the advent of recombinant DNA technology, exemplified by the 1978 synthesis of human insulin chains in E. coli by Genentech scientists, marking the first commercial recombinant protein and demonstrating the feasibility of bacterial hosts for therapeutic production.[12] Today, they remain dominant for industrial-scale expression due to their scalability and low media costs, often yielding up to 30% of total cellular protein for optimized constructs.[13] Key host strains include E. coli BL21(DE3), which harbors a chromosomal copy of the T7 RNA polymerase gene under lacUV5 control for robust expression of T7-promoter-driven genes, making it ideal for high-level protein production.[14] In contrast, DH5α is primarily used for cloning and plasmid propagation due to its endonuclease-deficient genotype (recA1, endA1), which minimizes DNA rearrangements and improves plasmid stability during initial construct assembly.[15] Other variants, such as those engineered for reduced protease activity, further enhance solubility and yield for challenging proteins. Expression vectors in these systems are typically circular plasmids derived from origins like pMB1 (a ColE1 derivative), enabling high-copy replication in E. coli (20–50 copies per cell).[11] They incorporate selectable markers, such as ampicillin resistance genes, to ensure maintenance in antibiotic-supplemented media, and strong promoters including the IPTG-inducible lac promoter for moderate control, the hybrid tac promoter for enhanced strength, and the T7 promoter for maximal transcription rates when paired with T7 polymerase.[11] Fusion tags like the polyhistidine (His-tag) are commonly appended to facilitate affinity purification via immobilized metal ion chromatography, often yielding proteins >95% pure after a single step.[11] Induction strategies optimize expression timing and level; for lac- or tac-based systems, isopropyl β-D-1-thiogalactopyranoside (IPTG) is added at 0.1–1 mM to derepress the promoter after initial cell growth, typically at mid-log phase (OD600 ~0.6).[16] Auto-induction media, containing glucose to repress early expression followed by lactose as both carbon source and inducer, enable growth to high densities (OD600 >10) without manual addition, simplifying screening and scaling while boosting yields by 2–5 fold.[17] Codon optimization aligns the gene sequence with E. coli's tRNA pool, reducing rare codon usage that causes ribosomal stalling and improving expression efficiency by up to 10-fold for heterologous genes.[11] The production process begins with transformation of competent E. coli cells via heat shock or electroporation with the recombinant plasmid, followed by selection on agar plates.[18] Cultures are then scaled from shake flasks (small-volume screening) to fermenters for high-density fed-batch cultivation, maintaining pH, oxygen, and nutrients to reach biomass >50 g/L dry cell weight.[13] Harvest involves centrifugation, followed by cell lysis using sonication, French press, or chemical methods to release cytoplasmic proteins; if insoluble inclusion bodies form (common for overexpressed hydrophobic proteins), they are solubilized in urea or guanidine hydrochloride and refolded via dilution or dialysis to recover active forms.[18] Despite their advantages, bacterial systems lack eukaryotic PTMs such as glycosylation, limiting their use for proteins requiring such modifications for activity or stability.[11] Additionally, Gram-negative hosts like E. coli produce lipopolysaccharides (LPS), potent endotoxins that contaminate extracts and necessitate rigorous purification (e.g., polymyxin B columns) to achieve <0.1 EU/mg for biomedical applications.[19]Eukaryotic Expression Systems
Eukaryotic expression systems are essential for the recombinant production of proteins that necessitate complex post-translational modifications (PTMs), such as proper folding, glycosylation, and secretion, which are often absent or inaccurate in prokaryotic hosts. These systems utilize yeast, insect, or mammalian cells to mimic native eukaryotic processing machinery, enabling the production of functional therapeutics like monoclonal antibodies and hormones that require specific glycan structures for activity, stability, and immunogenicity. Unlike bacterial systems, which may produce endotoxins unsuitable for clinical applications, eukaryotic hosts provide a more biologically relevant environment for such proteins.[20][21] Yeast-based systems, such as Saccharomyces cerevisiae and Pichia pastoris, offer cost-effective scalability and inducible promoters for high-level expression of secreted enzymes and industrial proteins. In S. cerevisiae, the GAL promoter enables galactose-inducible expression, facilitating controlled production of recombinant proteins in fermenters. P. pastoris, a methylotrophic yeast, employs the AOX1 promoter for methanol-inducible expression, achieving yields up to 10 g/L for secreted proteins like phytases and glucose oxidase due to its efficient secretory pathway and protease-deficient strains. However, yeast systems often encounter hyperglycosylation issues, where excessive mannose addition results in hypermannosylated proteins that may alter pharmacokinetics and reduce therapeutic efficacy.[22][23][24] Insect cell systems, primarily using the baculovirus expression vector system (BEVS) in Spodoptera frugiperda cell lines like Sf9 and Sf21, support transient expression with PTMs resembling those in higher eukaryotes, including phosphorylation, acetylation, and N-linked glycosylation. Baculovirus infection drives rapid, high-level protein production without stable integration, making it ideal for structural studies and vaccine antigens; Sf9 cells are favored for virus propagation, while Sf21 cells enhance yields for complex assemblies like virus-like particles. This transient approach avoids genomic disruption but requires virus amplification, limiting it to smaller-scale applications compared to stable mammalian lines.[25][26] Mammalian systems, particularly Chinese hamster ovary (CHO) and human embryonic kidney (HEK293) cells, dominate biopharmaceutical production for their ability to perform human-like PTMs, especially sialylated glycosylation critical for serum half-life. CHO cells are used for stable integration via dihydrofolate reductase (DHFR) selection and amplification, yielding up to 10 g/L of biologics like monoclonal antibodies in fed-batch bioreactors.[27] HEK293 cells excel in transient transfection for rapid prototyping, achieving comparable titers for antibodies and viral vectors through optimized protocols. These systems support secretion into culture media, simplifying purification.[28][29][30] Key vectors and methods in eukaryotic systems include shuttle vectors that replicate in both bacterial and eukaryotic hosts for cloning and expression, such as pPICZ for P. pastoris or pcDNA for mammalian cells. Viral delivery, exemplified by adeno-associated virus (AAV) vectors, facilitates transient or integrative expression in mammalian and insect cells, while CRISPR-Cas9 enables precise genomic integration to enhance stability and yield. The overall process involves DNA transfection or transduction, antibiotic or metabolic selection for stable lines, clonal isolation, and scale-up in shake flasks to bioreactors for optimized fed-batch or perfusion cultures.[31][32][33] CHO cells produce approximately 70% of therapeutic monoclonal antibodies approved for clinical use, underscoring their prevalence in biomanufacturing. A pivotal historical shift occurred in the 1980s from bacterial systems to eukaryotic hosts for glycosylated proteins, driven by the need for authentic PTMs; this was exemplified by recombinant erythropoietin (EPO), approved in 1989 and produced in CHO cells, marking the first major eukaryotic-derived biologic for anemia treatment.[34][35] The primary advantages of eukaryotic systems include authentic, human-like glycosylation that ensures protein functionality and reduces immunogenicity risks for therapeutics. Challenges encompass slower cell growth rates (doubling times of 12-24 hours versus 20-30 minutes in bacteria), higher media and facility costs, and potential for heterogeneous glycosylation in non-mammalian hosts, necessitating engineering for consistency.[36][37]Cell-Free Expression Systems
Cell-free expression systems enable the synthesis of proteins in vitro using cellular extracts or reconstituted components, without the constraints of intact living cells. These acellular platforms typically incorporate ribosomes, transfer RNAs (tRNAs), and translation enzymes derived from lysed cells, such as wheat germ extracts or rabbit reticulocyte lysates, or through purified recombinant elements as in the PURE system.[38] The foundational work on these systems dates to the 1960s, when Marshall Nirenberg and colleagues developed E. coli-based extracts to decipher the genetic code, demonstrating that exogenous mRNA could direct protein synthesis in a cell-free environment.[39] A modern milestone is the PURE (protein synthesis using recombinant elements) system, introduced in 2001, which reconstitutes translation from highly purified E. coli components for precise control; commercial variants like PURExpress further optimize this for routine use.[40] Essential components of these systems include energy sources such as adenosine triphosphate (ATP) and guanosine triphosphate (GTP), free amino acids, nucleoside triphosphates (NTPs) for transcription, and genetic templates in the form of DNA or mRNA. Many setups employ coupled transcription-translation, where T7 RNA polymerase generates mRNA from a DNA template, which is then immediately translated by the ribosomal machinery. Common formats include batch reactions, where all substrates are added upfront and depleted within hours, yielding approximately 0.1–1 mg/mL of protein; continuous-flow configurations using dialysis membranes to replenish substrates and remove byproducts, extending reactions for higher productivity; and microreactor systems, which miniaturize the process for enhanced efficiency in small volumes.[38][41] These methods support rapid prototyping, with reactions completing in 1–4 hours.[42] Key advantages of cell-free systems lie in their flexibility, as the absence of cell walls and membranes allows expression of toxic proteins that would harm living hosts, such as certain antimicrobial peptides or membrane proteins. They also facilitate straightforward incorporation of unnatural amino acids by supplementing the reaction mix, enabling site-specific modifications for probing protein function or creating novel biomaterials. Additionally, the open nature of these systems supports high-throughput screening, where libraries of protein variants can be synthesized and assayed directly in multi-well plates or microfluidic devices for applications like directed evolution. In structural biology, cell-free expression has proven invaluable for producing isotopically labeled proteins for NMR spectroscopy or crystallography, often yielding sufficient quantities for analysis without the need for cellular optimization.[43][44][45][46] Despite these benefits, cell-free systems face limitations, including high reagent costs due to the need for purified components and energy substrates, which restrict scalability compared to cellular methods. Reaction durations are typically short, lasting only hours before instability sets in from factor degradation or byproduct accumulation, and protein yields remain lower than those from optimized in vivo systems without further engineering. These platforms are thus best suited for small-scale, specialized applications rather than bulk production.[38][41]Advances and Applications
Recent Developments
In recent years, synthetic biology has integrated genetic circuits into cell-free systems to enable precise, orthogonal control over protein expression. For instance, toehold switches—riboregulator RNA devices—have been incorporated into cell-free platforms to modulate translation in response to specific triggers, with 2018 advancements demonstrating their use in low-cost, paper-based biosensors for detecting microbial RNAs without cellular constraints.[47] Complementing this, de novo design of metabolic pathways using CRISPR-Cas systems has facilitated the engineering of novel microbial hosts for enhanced protein production, as highlighted in 2017 reviews on CRISPR-mediated metabolic engineering that enable targeted insertions and optimizations in recombinant strains.[48] Advancements in cell-free protein synthesis have focused on continuous exchange cell-free (CECF) formats, which sustain reactions by replenishing substrates and removing byproducts, achieving yields exceeding 10 mg/mL in optimized setups using glycolytic intermediates as energy sources.[49] These systems have also expanded the genetic code to incorporate over 100 unnatural amino acids site-specifically, leveraging orthogonal tRNA-synthetase pairs in extracts from genomically recoded bacteria, as demonstrated in 2018 studies that broadened chemical diversity in synthesized proteins.[50] In prokaryotic systems, genome-scale engineering has evolved through iterations of multiplex automated genome engineering (MAGE) in Escherichia coli, with enhancements in the 2020s improving recombination efficiency for simultaneous multi-locus edits to boost expression yields and reduce toxicity; recent 2025 studies have further advanced soluble protein production in E. coli by addressing aggregation and disulfide bond challenges.[51][52] Additionally, auxotrophic strains have been refined for cleaner production by limiting endogenous amino acid synthesis, enabling selective metabolic labeling and higher purity in recombinant outputs, as shown in 2021 studies on stable synthetic auxotrophies.[53] Eukaryotic platforms have seen significant glycoengineering, particularly in yeast, where 2016 efforts humanized N-glycosylation pathways by introducing mammalian glycosyltransferases to produce sialylated glycoproteins compatible with therapeutic applications.[54] In plant systems, transient expression via agroinfiltration in Nicotiana benthamiana has scaled up in the 2020s for rapid vaccine antigen production, with 2022 protocols yielding SARS-CoV-2 proteins at multigram levels per plant batch.[55] Specific innovations include cell-free platforms for mRNA vaccine production, exemplified by the 2020 rapid synthesis of COVID-19 mRNA via in vitro transcription, which enabled scalable, cell-free manufacturing without live-cell risks.[56] AI-driven approaches have further optimized codon usage for improved expression, integrating tools like AlphaFold3 (2024) for structure predictions to guide designs that enhance folding efficiency and solubility.[57][58] Emerging hybrid systems combine proto-cells—lipid vesicles encapsulating cell-free extracts—with minimal genomes to mimic cellular compartmentalization, allowing sustained protein production in semi-synthetic environments, as explored in 2020 bottom-up assemblies that integrate genetic and metabolic modules.[59] From 2023 to 2025, cell-free protein synthesis has advanced with AI-optimized workflows for higher productivity and scaled GMP manufacturing up to 4,500 L, as demonstrated by platforms enabling rapid production of therapeutics.[60][61]Industrial and Biomedical Uses
Protein production technologies have revolutionized industrial applications, particularly in the manufacturing of enzymes used in detergents, food processing, and biofuels. Recombinant bacterial and yeast systems are widely employed for producing amylases and lipases, which break down starches and fats, respectively. For instance, Novozymes utilizes the yeast Pichia pastoris to produce lipases for detergent formulations, enabling efficient stain removal in laundry products. The global industrial enzymes market, dominated by such recombinant products, is valued at approximately USD 8.42 billion in 2025 and projected to reach USD 12.01 billion by 2030, driven by demand in the cleaning and food sectors.[62][63] In biomedical therapeutics, recombinant protein production underpins a multibillion-dollar industry, with key examples including insulin, monoclonal antibodies, and growth factors. The first recombinant human insulin was produced in Escherichia coli by Genentech in 1978 and received FDA approval in 1982 as Humulin, marking the advent of genetically engineered pharmaceuticals for diabetes treatment. Monoclonal antibodies, primarily manufactured in Chinese hamster ovary (CHO) cells, dominate the biopharmaceutical market, with sales exceeding USD 200 billion annually by 2025 due to their role in cancer and autoimmune therapies. Erythropoietin (EPO), a growth factor for treating anemia, is produced recombinantly in mammalian cells and was first approved by the FDA in 1989 as Epogen.[64][65][66][67][68] Vaccines and diagnostics also leverage these systems for scalable antigen production. The HPV vaccine Cervarix, approved in 2009, uses baculovirus-infected insect cells to generate virus-like particles from HPV-16 and HPV-18 L1 proteins, providing protection against cervical cancer precursors. Cell-free expression systems enable rapid production of vaccine antigens, such as those derived from mRNA templates, facilitating quick responses to emerging pathogens by bypassing cellular constraints.[63][69][70] Process economics vary significantly by host system, influencing scalability and adoption. Bacterial production, such as in E. coli, achieves costs ranging from USD 0.1 to 1 per gram for industrial applications due to fast growth and simple media, while mammalian systems like CHO cells cost approximately USD 50 to 150 per gram for pharmaceuticals owing to complex glycosylation needs and stringent purification.[71][72] Scaling occurs in bioreactors up to 10,000 liters, with good manufacturing practice (GMP) compliance ensuring product quality through validated sterilization, monitoring, and contamination controls. The FDA's approval of the first recombinant drug in 1982 set precedents for biosafety, requiring containment for genetically modified organisms (GMOs) under NIH guidelines to prevent environmental release during production.[73][74] Unique challenges arise in supply chains, particularly during pandemics, as seen in 2020 when COVID-19 disrupted raw materials and facility access, delaying recombinant protein vaccine scaling despite accelerated approvals. These bottlenecks highlighted vulnerabilities in global dependencies for media components and equipment, prompting investments in resilient, localized manufacturing.[75][76]| Production System | Approximate Cost per Gram (USD) | Typical Scale (Liters) | Key Applications |
|---|---|---|---|
| Bacterial (E. coli) | 0.1–1 (industrial) | 1,000–5,000 | Insulin, industrial enzymes |
| Mammalian (CHO cells) | 50–150 (pharma) | 5,000–20,000 | Monoclonal antibodies, EPO |
| Yeast (P. pastoris) | 1–10 | 1,000–10,000 | Lipases, vaccines |
| Insect cells (baculovirus) | 5–50 | 500–5,000 | HPV VLPs |


