Recent from talks
Nothing was collected or created yet.
Eukaryogenesis
View on Wikipedia

Eukaryogenesis, the process which created the eukaryotic cell and lineage, is a milestone in the evolution of life, since eukaryotes include all complex cells and almost all multicellular organisms. The process is widely agreed to have involved symbiogenesis, in which an archaeon and one or more bacteria came together to create the first eukaryotic common ancestor (FECA). This cell had a new level of complexity and capability, with a nucleus, at least one centriole and cilium, facultatively aerobic mitochondria, sex (meiosis and syngamy), a dormant cyst with a cell wall of chitin and/or cellulose and peroxisomes. It evolved into a population of single-celled organisms that included the last eukaryotic common ancestor (LECA), gaining capabilities along the way, though the sequence of steps involved has been disputed, and may not have started with symbiogenesis. In turn, the LECA gave rise to the eukaryotes' crown group, containing the ancestors of animals, fungi, plants, and a diverse range of single-celled organisms.
Context
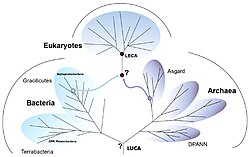
[edit]Life arose on Earth once it had cooled enough for oceans to form. That developed into the last universal common ancestor (LUCA), an organism which had ribosomes and the genetic code, some 4 billion years ago. It gave rise to two main branches of prokaryotic life, the Bacteria and the Archaea. From among these small-celled, rapidly-dividing ancestors arose the Eukaryotes, with much larger cells, nuclei, and distinctive biochemistry.[1][2] The eukaryotes form a domain that contains all complex cells and most types of multicellular organism, including the animals, plants, and fungi.[3][4]
Symbiogenesis
[edit]
According to the theory of symbiogenesis (the endosymbiotic theory) championed by Lynn Margulis, a member of the archaea gained a bacterial cell as a component. The archaeal cell was a member of the Promethearchaeati kingdom. The bacterium was one of the alphaproteobacteria, which had the ability to use oxygen in its respiration. This enabled it – and the archaeal cells that included it – to survive in the presence of oxygen, which was poisonous to other organisms adapted to reducing conditions. The endosymbiotic bacteria became the eukaryotic cell's mitochondria, providing most of the energy of the cell.[1][5] Lynn Margulis and colleagues have suggested that the cell also acquired a Spirochaete bacterium as a symbiont, providing the cell skeleton of microtubules and the ability to move, including the ability to pull chromosomes into two sets during mitosis, cell division.[6] More recently, the archaean has been identified as belonging to the unranked taxon Heimdallarchaeia of the phylum Promethearchaeota.[7]
Last eukaryotic common ancestor (LECA)
[edit]The last eukaryotic common ancestor (LECA) is the hypothetical last common ancestor of all living eukaryotes, around 2 billion years ago,[3][4] and was most likely a biological population.[8] It is believed to have been a protist with a nucleus, at least one centriole and cilium, facultatively aerobic mitochondria, sex (meiosis and syngamy), a dormant cyst with a cell wall of chitin and/or cellulose, and peroxisomes.[9][10]
It had been proposed that the LECA fed by phagocytosis, engulfing other organisms.[9][10] However, in 2022, Nico Bremer and colleagues confirmed that the LECA had mitochondria, and stated that it had multiple nuclei, but disputed that it was phagotrophic. This would mean that the ability found in many eukaryotes to engulf materials developed later, rather than being acquired first and then used to engulf the alphaproteobacteria that became mitochondria.[11]
The LECA has been described as having "spectacular cellular complexity".[12] Its cell was divided into compartments.[12] It appears to have inherited a set of endosomal sorting complex proteins that enable membranes to be remodelled, including pinching off vesicles to form endosomes.[13] Its apparatuses for transcribing DNA into RNA, and then for translating the RNA into proteins, were separated, permitting extensive RNA processing and allowing the expression of genes to become more complex.[14] It had mechanisms for reshuffling its genetic material, and possibly for manipulating its own evolvability. All of these gave the LECA "a compelling cohort of selective advantages".[12]
Eukaryotic sex
[edit]Sex in eukaryotes is a composite process, consisting of meiosis and fertilisation, which can be coupled to reproduction.[15] Dacks and Roger[16] proposed on the basis of a phylogenetic analysis that facultative sex was likely present in the common ancestor of all eukaryotes. Early in eukaryotic evolution, about 2 billion years ago, organisms needed a solution to the major problem that oxidative metabolism releases reactive oxygen species that damage the genetic material, DNA.[15] Eukaryotic sex provides a process, homologous recombination during meiosis, for using informational redundancy to repair such DNA damage.[15]
Scenarios
[edit]Competing sequences of mitochondria, membranes, and nucleus
[edit]Biologists have proposed multiple scenarios for the creation of the eukaryotes. While there is broad agreement that the LECA must have had a nucleus, mitochondria, and internal membranes, the order in which these were acquired has been disputed.[12] In the syntrophic model, the first eukaryotic common ancestor (FECA, around 2.2 gya) gained mitochondria, then membranes, then a nucleus.[12] In the phagotrophic model, it gained a nucleus, then membranes, then mitochondria.[12] In a more complex process, it gained all three in short order, then other capabilities. Other models have been proposed. Whatever happened, many lineages must have been created, but the LECA either out-competed or came together with the other lineages to form a single point of origin for the eukaryotes.[12]
Nick Lane and William Martin have argued that mitochondria came first, on the grounds that energy had been the limiting factor on the size of the prokaryotic cell.[17] Enrique M. Muro et al. have argued, however, that the genetic system needed to reach a critical point that led to a new regulatory system (with introns and the spliceosome), which enabled coordination between genetic networks.[18] The phagotrophic model presupposes the ability to engulf food, enabling the cell to engulf the aerobic bacterium that became the mitochondrion.[12]
Eugene Koonin and others, noting that the archaea share many features with eukaryotes, argue that rudimentary eukaryotic traits such as membrane-lined compartments were acquired before endosymbiosis added mitochondria to the early eukaryotic cell, while the cell wall was lost. In the same way, mitochondrial acquisition must not be regarded as the end of the process, for still new complex families of genes had to be developed after or during the endosymbiotic exchange. In this way, from FECA to LECA, the organisms can be considered as proto-eukaryotes. At the end of the process, LECA was already a complex organism with protein families involved in cellular compartmentalization.[19][20]
Viral eukaryogenesis
[edit]Another scenario is viral eukaryogenesis, which proposes that the eukaryotes arose as an emergent superorganism, with the nucleus deriving from a "viral factory" alongside the alphaproteobacterium mitochondrion, hosted by an archaeal cell. In this scenario, eukaryogenesis began when a virus colonised an archaeal cell, making it support the production of viruses. The virus may later have assisted the bacterium's entry into the reprogrammed cell.[21] Eukaryotes share genes for several DNA synthesis and transcription enzymes with DNA viruses (Nucleocytoviricota). Those viruses may thus be older than the LECA and may have exchanged DNA with proto-eukaryotes.[22]
Diversification: crown eukaryotes
[edit]In turn, the LECA gave rise to the eukaryotes' crown group, containing the ancestors of animals, fungi, plants, and a diverse range of single-celled organisms with the new capabilities and complexity of the eukaryotic cell.[23][24] Single cells without cell walls are fragile and have a low probability of being fossilised. If fossilised, they have few features to distinguish them clearly from prokaryotes: size, morphological complexity, and (eventually) multicellularity. Early eukaryote fossils, from the late Paleoproterozoic, include acritarch microfossils with relatively robust ornate carbonaceous vesicles of Tappania from 1.63 gya and Shuiyousphaeridium from 1.8 gya.[24]
The position of the LECA on the eukaryotic tree of life remains controversial. Some studies believe that the first split after the LECA happened between the Unikonta and the Bikonta (Stechmann and Cavalier-Smith 2003), or between Amorphea and all other eukaryotes (Adl et al. 2012; Derelle and Lang 2012). Some believe that the first split happened within Excavata (al Jewari and Baldauf 2023).[25] Yet others believe in a first split between the Opisthokonta and all others (Cerón-Romero et al. 2024).[26]
References
[edit]- ^ a b c McGrath, Casey (31 May 2022). "Highlight: Unraveling the Origins of LUCA and LECA on the Tree of Life". Genome Biology and Evolution. 14 (6) evac072. doi:10.1093/gbe/evac072. PMC 9168435.
- ^ Weiss, Madeline C.; Sousa, F. L.; Mrnjavac, N.; et al. (2016). "The physiology and habitat of the last universal common ancestor" (PDF). Nature Microbiology. 1 (9): 16116. doi:10.1038/nmicrobiol.2016.116. PMID 27562259. S2CID 2997255.
- ^ a b Gabaldón, T. (October 2021). "Origin and Early Evolution of the Eukaryotic Cell". Annual Review of Microbiology. 75 (1): 631–647. doi:10.1146/annurev-micro-090817-062213. PMID 34343017. S2CID 236916203.
- ^ a b Woese, C.R.; Kandler, Otto; Wheelis, Mark L. (June 1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proceedings of the National Academy of Sciences of the United States of America. 87 (12): 4576–4579. Bibcode:1990PNAS...87.4576W. doi:10.1073/pnas.87.12.4576. PMC 54159. PMID 2112744.
- ^ a b Latorre, A.; Durban, A; Moya, A.; Pereto, J. (2011). "The role of symbiosis in eukaryotic evolution". In Gargaud, Muriel; López-Garcìa, Purificacion; Martin H. (eds.). Origins and Evolution of Life: An astrobiological perspective. Cambridge: Cambridge University Press. pp. 326–339. ISBN 978-0-521-76131-4. Archived from the original on 24 March 2019. Retrieved 27 August 2017.
- ^ Margulis, Lynn; Chapman, Michael; Guerrero, Ricardo; Hall, John (29 August 2006). "The last eukaryotic common ancestor (LECA): Acquisition of cytoskeletal motility from aerotolerant spirochetes in the Proterozoic Eon". Proceedings of the National Academy of Sciences. 103 (35): 13080–13085. Bibcode:2006PNAS..10313080M. doi:10.1073/pnas.0604985103. PMC 1559756. PMID 16938841.
- ^ Williams, Tom A.; Cox, Cymon J.; Foster, Peter G.; Szöllősi, Gergely J.; Embley, T. Martin (9 December 2019). "Phylogenomics provides robust support for a two-domains tree of life". Nature Ecology & Evolution. 4 (1): 138–147. Bibcode:2019NatEE...4..138W. doi:10.1038/s41559-019-1040-x. PMC 6942926. PMID 31819234.
- ^ O'Malley, Maureen A.; Leger, Michelle M.; Wideman, Jeremy G.; Ruiz-Trillo, Iñaki (18 February 2019). "Concepts of the last eukaryotic common ancestor". Nature Ecology & Evolution. 3 (3): 338–344. Bibcode:2019NatEE...3..338O. doi:10.1038/s41559-019-0796-3. hdl:10261/201794. PMID 30778187. S2CID 256718457.
- ^ a b Leander, B. S. (May 2020). "Predatory protists". Current Biology. 30 (10): R510 – R516. Bibcode:2020CBio...30.R510L. doi:10.1016/j.cub.2020.03.052. PMID 32428491. S2CID 218710816.
- ^ a b Strassert, Jürgen F. H.; Irisarri, Iker; Williams, Tom A.; Burki, Fabien (25 March 2021). "A molecular timescale for eukaryote evolution with implications for the origin of red algal-derived plastids". Nature Communications. 12 (1): 1879. Bibcode:2021NatCo..12.1879S. doi:10.1038/s41467-021-22044-z. PMC 7994803. PMID 33767194.
- ^ Bremer, Nico; Tria, Fernando D. K.; Skejo, Josip; Garg, Sriram G.; Martin, William F. (31 May 2022). "Ancestral State Reconstructions Trace Mitochondria But Not Phagocytosis to the Last Eukaryotic Common Ancestor". Genome Biology and Evolution. 14 (6) evac079. doi:10.1093/gbe/evac079. PMC 9185374. PMID 35642316.
- ^ a b c d e f g h Koumandou, V. Lila; Wickstead, Bill; Ginger, Michael L.; van der Giezen, Mark; Dacks, Joel B.; Field, Mark C. (2013). "Molecular paleontology and complexity in the last eukaryotic common ancestor". Critical Reviews in Biochemistry and Molecular Biology. 48 (4): 373–396. doi:10.3109/10409238.2013.821444. PMC 3791482. PMID 23895660.
- ^ Makarova, Kira S.; Yutin, Natalya; Bell, Stephen D.; Koonin, Eugene V. (6 September 2010). "Evolution of diverse cell division and vesicle formation systems in Archaea". Nature Reviews Microbiology. 8 (10): 731–741. doi:10.1038/nrmicro2406. PMC 3293450. PMID 20818414.
- ^ Martin, William; Koonin, Eugene V. (2006). "Introns and the origin of nucleus–cytosol compartmentalization". Nature. 440 (7080): 41–45. Bibcode:2006Natur.440...41M. doi:10.1038/nature04531. ISSN 0028-0836. PMID 16511485.
- ^ a b c Horandl, E.; Speijer, D. (7 February 2018). "How oxygen gave rise to eukaryotic sex". Proceedings of the Royal Society B: Biological Sciences. 285 (1872). The Royal Society. doi:10.1098/rspb.2017.2706. PMC 5829205. PMID 29436502.
- ^ Dacks, J.; Roger, A. J. (1999). "The first sexual lineage and the relevance of facultative sex". Journal of Molecular Evolution. 48 (6): 779–783. Bibcode:1999JMolE..48..779D. doi:10.1007/pl00013156. PMID 10229582. S2CID 9441768.
- ^ Lane, Nick; Martin, William F. (2010). "The energetics of genome complexity". Nature. 467 (7318): 929–934. Bibcode:2010Natur.467..929L. doi:10.1038/nature09486. PMID 20962839. S2CID 17086117.
- ^ Muro, Enrique M.; Ballesteros, Fernando J.; Luque, Bartolo; Bascompte, Jordi (2025). "The emergence of eukaryotes as an evolutionary algorithmic phase transition". PNAS. 122 (13) e2422968122. Bibcode:2025PNAS..12222968M. doi:10.1073/pnas.2422968122. PMC 12002324. PMID 40146859.
- ^ Koonin, Eugene V. (March 2005). "The incredible expanding ancestor of eukaryotes". Cell. 140 (5): 606–608. doi:10.1016/j.cell.2010.02.022. PMC 3293451. PMID 20211127.
- ^ Martijn, J.; Ettema, T.J.G. (February 2013). "From archaeon to eukaryote: the evolutionary dark ages of the eukaryotic cell". Biochem Soc Transactions. 41 (1): 451–7. doi:10.1042/BST20120292. PMID 23356327.
- ^ Bell, Philip J. L. (11 May 2022). "Eukaryogenesis: The Rise of an Emergent Superorganism". Frontiers in Microbiology. 13 858064. doi:10.3389/fmicb.2022.858064. PMC 9130767.
- ^ Karki, Sangita; Barth, Zachary K.; Aylward, Frank O. (2025). "Ancient Host-Virus Gene Transfer Hints at a Diverse Pre-LECA Virosphere". Journal of Molecular Evolution. 93 (3): 295–305. Bibcode:2025JMolE..93..295K. doi:10.1007/s00239-025-10246-8. PMC 12198294. PMID 40298963.
- ^ Van de Peer, Yves; Baldaufrid, Sandra L.; Doolittle, W. Ford; Meyerid, Axel (2000). "An Updated and Comprehensive rRNA Phylogeny of (Crown) Eukaryotes Based on Rate-Calibrated Evolutionary Distances". Journal of Molecular Evolution. 51 (6): 565–576. Bibcode:2000JMolE..51..565V. doi:10.1007/s002390010120. PMID 11116330. S2CID 9400485.
- ^ a b Butterfield, N.J. (2015). "Early evolution of the Eukaryota". Palaeontology. 58 (1): 5–17. Bibcode:2015Palgy..58....5B. doi:10.1111/pala.12139.
- ^ Al Jewari, Caesar; Baldauf, Sandra L. (28 April 2023). "An excavate root for the eukaryote tree of life". Science Advances. 9 (17) eade4973. Bibcode:2023SciA....9E4973A. doi:10.1126/sciadv.ade4973. PMC 10146883. PMID 37115919.
- ^ Cerón-Romero, Mario A.; Fonseca, Miguel M.; de Oliveira Martins, Leonardo; Posada, David; Katz, Laura A. (3 August 2022). "Phylogenomic Analyses of 2,786 Genes in 158 Lineages Support a Root of the Eukaryotic Tree of Life between Opisthokonts and All Other Lineages". Genome Biology and Evolution. 14 (8) evac119. doi:10.1093/gbe/evac119. PMC 9366629. PMID 35880421.
External links
[edit]- Attraction and sex among our microbial Last Eukaryotic Common Ancestors, The Atlantic, 11 November 2020