Recent from talks
Nothing was collected or created yet.
Microorganism
View on Wikipedia

| Part of a series on |
| Biology |
|---|
 |
A microorganism, or microbe,[a] is an organism of microscopic size, which may exist in its single-celled form or as a colony of cells. The possible existence of unseen microbial life was suspected from antiquity, with an early attestation in Jain literature authored in 6th-century BC India. The scientific study of microorganisms began with their observation under the microscope in the 1670s by Anton van Leeuwenhoek. In the 1850s, Louis Pasteur found that microorganisms caused food spoilage, debunking the theory of spontaneous generation. In the 1880s, Robert Koch discovered that microorganisms caused the diseases tuberculosis, cholera, diphtheria, and anthrax.
Microorganisms are extremely diverse, representing most unicellular organisms in all three domains of life: two of the three domains, Archaea and Bacteria, only contain microorganisms. The third domain, Eukaryota, includes all multicellular organisms as well as many unicellular protists and protozoans that are microbes. Some protists are related to animals and some to green plants. Many multicellular organisms are also microscopic, namely micro-animals, some fungi, and some algae.
Microorganisms can have very different habitats, and live everywhere from the poles to the equator, in deserts, geysers, rocks, and the deep sea. Some are adapted to extremes such as very hot or very cold conditions, others to high pressure, and a few, such as Deinococcus radiodurans, to high radiation environments. Microorganisms also make up the microbiota found in and on all multicellular organisms. There is evidence that 3.45-billion-year-old Australian rocks once contained microorganisms, the earliest direct evidence of life on Earth.[1][2]
Microbes are important in human culture and health in many ways, serving to ferment foods and treat sewage, and to produce fuel, enzymes, and other bioactive compounds. Microbes are essential tools in biology as model organisms and have been put to use in biological warfare and bioterrorism. Microbes are a vital component of fertile soil. In the human body, microorganisms make up the human microbiota, including the essential gut flora. The pathogens responsible for many infectious diseases are microbes and, as such, are the target of hygiene measures.
Discovery
[edit]Ancient precursors
[edit]

The possible existence of microscopic organisms was discussed for many centuries before their discovery in the 17th century. By the 6th century BC, the Jains of present-day India postulated the existence of tiny organisms called nigodas.[3] These nigodas are said to be born in clusters; they live everywhere, including the bodies of plants, animals, and people; and their life lasts only for a fraction of a second.[4] According to Mahavira, the 24th preacher of Jainism, the humans destroy these nigodas on a massive scale, when they eat, breathe, sit, and move.[3] Many modern Jains assert that Mahavira's teachings presage the existence of microorganisms as discovered by science.[5]
The earliest known idea to indicate the possibility of diseases spreading by yet unseen organisms was that of the Roman scholar Marcus Terentius Varro in a 1st-century BC book entitled On Agriculture in which he called the unseen creatures animalia minuta, and warns against locating a homestead near a swamp:[6]
… and because there are bred certain minute creatures that cannot be seen by the eyes, which float in the air and enter the body through the mouth and nose and they cause serious diseases.[6]
In The Canon of Medicine (1020), Avicenna suggested that tuberculosis and other diseases might be contagious.[7][8]
Early modern
[edit]In the 15th-century, Turkish scientist Akshamsaddin speculated about microbial life relating to disease in his work Maddat ul-Hayat (The Material of Life):
It is a mistake to assume that diseases appear in individuals one by one. Diseases are transmitted from person to person. This transmission takes place through small seeds that are invisible to the eye, but are still alive.[9][10]
In 1546, Girolamo Fracastoro proposed that epidemic diseases were caused by transferable seedlike entities that could transmit infection by direct or indirect contact, or even without contact over long distances.[11]
Antonie van Leeuwenhoek is considered to be one of the fathers of microbiology. He was the first in 1673 to discover and conduct scientific experiments with microorganisms, using simple single-lensed microscopes of his own design.[12][13][14][15] Robert Hooke, a contemporary of Leeuwenhoek, also used microscopy to observe microbial life in the form of the fruiting bodies of moulds. In his 1665 book Micrographia, he made drawings of studies, and he coined the term cell.[16]
19th century
[edit]
Louis Pasteur (1822–1895) exposed boiled broths to the air, in vessels that contained a filter to prevent particles from passing through to the growth medium, and also in vessels without a filter, but with air allowed in via a curved tube so dust particles would settle and not come in contact with the broth. By boiling the broth beforehand, Pasteur ensured that no microorganisms survived within the broths at the beginning of his experiment. Nothing grew in the broths in the course of Pasteur's experiment. This meant that the living organisms that grew in such broths came from outside, as spores on dust, rather than spontaneously generated within the broth. Thus, Pasteur refuted the theory of spontaneous generation and supported the germ theory of disease.[17]

In 1876, Robert Koch (1843–1910) established that microorganisms can cause disease. He found that the blood of cattle that were infected with anthrax always had large numbers of Bacillus anthracis. Koch found that he could transmit anthrax from one animal to another by taking a small sample of blood from the infected animal and injecting it into a healthy one, and this caused the healthy animal to become sick. He also found that he could grow the bacteria in a nutrient broth, then inject it into a healthy animal, and cause illness. Based on these experiments, he devised criteria for establishing a causal link between a microorganism and a disease and these are now known as Koch's postulates.[18] Although these postulates cannot be applied in all cases, they do retain historical importance to the development of scientific thought and are still being used today.[19]
The discovery of microorganisms such as Euglena that did not fit into either the animal or plant kingdoms, since they were photosynthetic like plants, but motile like animals, led to the naming of a third kingdom in the 1860s. In 1860 John Hogg called this the Protoctista, and in 1866 Ernst Haeckel named it the Protista.[20][21][22]
The work of Pasteur and Koch did not accurately reflect the true diversity of the microbial world because of their exclusive focus on microorganisms having direct medical relevance. It was not until the work of Martinus Beijerinck and Sergei Winogradsky in the late 19th century that the true breadth of microbiology was revealed.[23] Beijerinck made two major contributions to microbiology: the discovery of viruses and the development of enrichment culture techniques.[24] While his work on the tobacco mosaic virus established the basic principles of virology, it was his development of enrichment culturing that had the most immediate impact on microbiology by allowing for the cultivation of a wide range of microbes with wildly different physiologies. Winogradsky was the first to develop the concept of chemolithotrophy and to thereby reveal the essential role played by microorganisms in geochemical processes.[25] He was responsible for the first isolation and description of both nitrifying and nitrogen-fixing bacteria.[23] French-Canadian microbiologist Félix d'Hérelle co-discovered bacteriophages and was one of the earliest applied microbiologists.[26]
Classification and structure
[edit]Microorganisms can be found almost anywhere on Earth. Bacteria and archaea are almost always microscopic, while a number of eukaryotes are also microscopic, including most protists, some fungi, as well as some micro-animals and plants. Viruses are generally regarded as not living and therefore not considered to be microorganisms, although a subfield of microbiology is virology, the study of viruses.[27][28][29]
Evolution
[edit]Single-celled microorganisms were the first forms of life to develop on Earth, approximately 3.5 billion years ago.[30][31][32] Further evolution was slow,[33] and for about 3 billion years in the Precambrian eon, (much of the history of life on Earth), all organisms were microorganisms.[34][35] Bacteria, algae and fungi have been identified in amber that is 220 million years old, which shows that the morphology of microorganisms has changed little since at least the Triassic period.[36] The newly discovered biological role played by nickel, however – especially that brought about by volcanic eruptions from the Siberian Traps – may have accelerated the evolution of methanogens towards the end of the Permian–Triassic extinction event.[37]
Microorganisms tend to have a relatively fast rate of evolution. Most microorganisms can reproduce rapidly, and bacteria are also able to freely exchange genes through conjugation, transformation and transduction, even between widely divergent species.[38] This horizontal gene transfer, coupled with a high mutation rate and other means of transformation, allows microorganisms to swiftly evolve (via natural selection) to survive in new environments and respond to environmental stresses. This rapid evolution is important in medicine, as it has led to the development of multidrug resistant pathogenic bacteria, superbugs, that are resistant to antibiotics.[39]
A possible transitional form of microorganism between a prokaryote and a eukaryote was discovered in 2012 by Japanese scientists. Parakaryon myojinensis is a unique microorganism larger than a typical prokaryote, but with nuclear material enclosed in a membrane as in a eukaryote, and the presence of endosymbionts. This is seen to be the first plausible evolutionary form of microorganism, showing a stage of development from the prokaryote to the eukaryote.[40][41]
Archaea
[edit]Archaea are prokaryotic unicellular organisms, and form the first domain of life in Carl Woese's three-domain system. A prokaryote is defined as having no cell nucleus or other membrane bound-organelle. Archaea share this defining feature with the bacteria with which they were once grouped. In 1990 the microbiologist Woese proposed the three-domain system that divided living things into bacteria, archaea and eukaryotes,[42] and thereby split the prokaryote domain.
Archaea differ from bacteria in both their genetics and biochemistry. For example, while bacterial cell membranes are made from phosphoglycerides with ester bonds, Achaean membranes are made of ether lipids.[43] Archaea were originally described as extremophiles living in extreme environments, such as hot springs, but have since been found in all types of habitats.[44] Only now are scientists beginning to realize how common archaea are in the environment, with Thermoproteota (formerly Crenarchaeota) being the most common form of life in the ocean, dominating ecosystems below 150 metres (490 ft) in depth.[45][46] These organisms are also common in soil and play a vital role in ammonia oxidation.[47]
The combined domains of archaea and bacteria make up the most diverse and abundant group of organisms on Earth and inhabit practically all environments where the temperature is below +140 °C (284 °F). They are found in water, soil, air, as the microbiome of an organism, hot springs and even deep beneath the Earth's crust in rocks.[48] The number of prokaryotes is estimated to be around five nonillion, or 5 × 1030, accounting for at least half the biomass on Earth.[49]
The biodiversity of the prokaryotes is unknown, but may be very large. A May 2016 estimate, based on laws of scaling from known numbers of species against the size of organism, gives an estimate of perhaps 1 trillion species on the planet, of which most would be microorganisms. Currently, only one-thousandth of one percent of that total have been described.[50] Archael cells of some species aggregate and transfer DNA from one cell to another through direct contact, particularly under stressful environmental conditions that cause DNA damage.[51][52]
Bacteria
[edit]
Like archaea, bacteria are prokaryotic – unicellular, and having no cell nucleus or other membrane-bound organelle. Bacteria are microscopic, with a few extremely rare exceptions, such as Thiomargarita namibiensis.[53] Bacteria function and reproduce as individual cells, but they can often aggregate in multicellular colonies.[54] Some species such as myxobacteria can aggregate into complex swarming structures, operating as multicellular groups as part of their life cycle,[55] or form clusters in bacterial colonies such as E. coli.
Their genome is usually a circular bacterial chromosome – a single loop of DNA, although they can also harbor small pieces of DNA called plasmids. These plasmids can be transferred between cells through bacterial conjugation. Bacteria have an enclosing cell wall, which provides strength and rigidity to their cells. They reproduce by binary fission or sometimes by budding, but do not undergo meiotic sexual reproduction. However, many bacterial species can transfer DNA between individual cells by a horizontal gene transfer process referred to as natural transformation.[56] Some species form extraordinarily resilient spores, but for bacteria this is a mechanism for survival, not reproduction. Under optimal conditions bacteria can grow extremely rapidly and their numbers can double as quickly as every 20 minutes.[57]
Eukaryotes
[edit]Most living things that are visible to the naked eye in their adult form are eukaryotes, including humans. However, many eukaryotes are also microorganisms. Unlike bacteria and archaea, eukaryotes contain organelles such as the cell nucleus, the Golgi apparatus and mitochondria in their cells. The nucleus is an organelle that houses the DNA that makes up a cell's genome. DNA (Deoxyribonucleic acid) itself is arranged in complex chromosomes.[58] Mitochondria are organelles vital in metabolism as they are the site of the citric acid cycle and oxidative phosphorylation. They evolved from symbiotic bacteria and retain a remnant genome.[59] Like bacteria, plant cells have cell walls, and contain organelles such as chloroplasts in addition to the organelles in other eukaryotes. Chloroplasts produce energy from light by photosynthesis, and were also originally symbiotic bacteria.[59]
Unicellular eukaryotes consist of a single cell throughout their life cycle. This qualification is significant since most multicellular eukaryotes consist of a single cell called a zygote only at the beginning of their life cycles. Microbial eukaryotes can be either haploid or diploid, and some organisms have multiple cell nuclei.
Unicellular eukaryotes usually reproduce asexually by mitosis under favorable conditions. However, under stressful conditions such as nutrient limitations and other conditions associated with DNA damage, they tend to reproduce sexually by meiosis and syngamy.[60]
Protists
[edit]
Of eukaryotic groups, the protists are most commonly unicellular and microscopic. This is a highly diverse group of organisms that are not easy to classify.[61][62] Several algae species are multicellular protists, and slime molds have unique life cycles that involve switching between unicellular, colonial, and multicellular forms.[63] The number of species of protists is unknown since only a small proportion has been identified. Protist diversity is high in oceans, deep sea-vents, river sediment and an acidic river, suggesting that many eukaryotic microbial communities may yet be discovered.[64][65]
Fungi
[edit]The fungi have several unicellular species, such as baker's yeast (Saccharomyces cerevisiae) and fission yeast (Schizosaccharomyces pombe). Some fungi, such as the pathogenic yeast Candida albicans, can undergo phenotypic switching and grow as single cells in some environments, and filamentous hyphae in others.[66]
Plants
[edit]The green algae are a large group of photosynthetic eukaryotes that include many microscopic organisms. Although some green algae are classified as protists, others such as charophyta are classified with embryophyte plants, which are the most familiar group of land plants. Algae can grow as single cells, or in long chains of cells. The green algae include unicellular and colonial flagellates, usually but not always with two flagella per cell, as well as various colonial, coccoid, and filamentous forms. In the Charales, which are the algae most closely related to higher plants, cells differentiate into several distinct tissues within the organism. There are about 6000 species of green algae.[67]
Ecology
[edit]Microorganisms are found in almost every habitat present in nature, including hostile environments such as the North and South poles, deserts, geysers, and rocks. They also include all the marine microorganisms of the oceans and deep sea. Some types of microorganisms have adapted to extreme environments and sustained colonies; these organisms are known as extremophiles. Extremophiles have been isolated from rocks as much as 7 kilometres below the Earth's surface,[68] and it has been suggested that the amount of organisms living below the Earth's surface is comparable with the amount of life on or above the surface.[48] Extremophiles have been known to survive for a prolonged time in a vacuum, and can be highly resistant to radiation, which may even allow them to survive in space.[69] Many types of microorganisms have intimate symbiotic relationships with other larger organisms; some of which are mutually beneficial (mutualism), while others can be damaging to the host organism (parasitism). If microorganisms can cause disease in a host they are known as pathogens. Microorganisms play critical roles in Earth's biogeochemical cycles as they are responsible for decomposition and nitrogen fixation.[70]
Bacteria use regulatory networks that allow them to adapt to almost every environmental niche on earth.[71][72] A network of interactions among diverse types of molecules including DNA, RNA, proteins and metabolites, is utilised by the bacteria to achieve regulation of gene expression. In bacteria, the principal function of regulatory networks is to control the response to environmental changes, for example nutritional status and environmental stress.[73] A complex organization of networks permits the microorganism to coordinate and integrate multiple environmental signals.[71]
Extremophiles
[edit]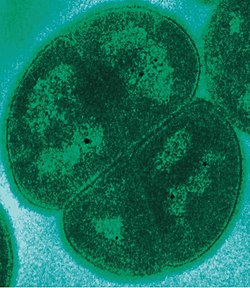
Extremophiles are microorganisms that have adapted so that they can survive and even thrive in extreme environments that are normally fatal to most life-forms. Thermophiles and hyperthermophiles thrive in high temperatures. Psychrophiles thrive in extremely low temperatures. – Temperatures as high as 130 °C (266 °F),[74] as low as −17 °C (1 °F)[75] Halophiles such as Halobacterium salinarum (an archaean) thrive in high salt conditions, up to saturation.[76] Alkaliphiles thrive in an alkaline pH of about 8.5–11.[77] Acidophiles can thrive in a pH of 2.0 or less.[78] Piezophiles thrive at very high pressures: up to 1,000–2,000 atm, down to 0 atm as in a vacuum of space.[b] A few extremophiles such as Deinococcus radiodurans are radioresistant,[80] resisting radiation exposure of up to 5k Gy. Extremophiles are significant in different ways. They extend terrestrial life into much of the Earth's hydrosphere, crust and atmosphere, their specific evolutionary adaptation mechanisms to their extreme environment can be exploited in biotechnology, and their very existence under such extreme conditions increases the potential for extraterrestrial life.[81]
Plants and soil
[edit]The nitrogen cycle in soils depends on the fixation of atmospheric nitrogen. This is achieved by a number of diazotrophs. One way this can occur is in the root nodules of legumes that contain symbiotic bacteria of the genera Rhizobium, Mesorhizobium, Sinorhizobium, Bradyrhizobium, and Azorhizobium.[82]
The roots of plants create a narrow region known as the rhizosphere that supports many microorganisms known as the root microbiome.[83]
These microorganisms in the root microbiome are able to interact with each other and surrounding plants through signals and cues. For example, mycorrhizal fungi are able to communicate with the root systems of many plants through chemical signals between both the plant and fungi. This results in a mutualistic symbiosis between the two. However, these signals can be eavesdropped by other microorganisms, such as the soil bacteria, Myxococcus xanthus, which preys on other bacteria. Eavesdropping, or the interception of signals from unintended receivers, such as plants and microorganisms, can lead to large-scale, evolutionary consequences. For example, signaler-receiver pairs, like plant-microorganism pairs, may lose the ability to communicate with neighboring populations because of variability in eavesdroppers. In adapting to avoid local eavesdroppers, signal divergence could occur and thus, lead to the isolation of plants and microorganisms from the inability to communicate with other populations.[84]
Symbiosis
[edit]
A lichen is a symbiosis of a macroscopic fungus with photosynthetic microbial algae or cyanobacteria.[85][86]
Applications
[edit]Microorganisms are useful in producing foods, treating waste water, creating biofuels and a wide range of chemicals and enzymes. They are invaluable in research as model organisms. They have been weaponised and sometimes used in warfare and bioterrorism. They are vital to agriculture through their roles in maintaining soil fertility and in decomposing organic matter. They also have applications in aquaculture, such as in biofloc technology.
Food production
[edit]Microorganisms are used in a fermentation process to make yoghurt, cheese, curd, kefir, ayran, xynogala, and other types of food. Fermentation cultures provide flavour and aroma, and inhibit undesirable organisms.[87] They are used to leaven bread, and to convert sugars to alcohol in wine and beer. Microorganisms are used in brewing, wine making, baking, pickling and other food-making processes.[88]
| Product | Contribution of microorganisms |
|---|---|
| Cheese | Growth of microorganisms contributes to ripening and flavor. The flavor and appearance of a particular cheese is due in large part to the microorganisms associated with it. Lactobacillus Bulgaricus is one of the microbes used in production of dairy products |
| Alcoholic beverages | Yeast is used to convert sugar, grape juice, or malt-treated grain into alcohol. Other microorganisms may also be used; a mold converts starch into sugar to make the Japanese rice wine, sake. Acetobacter Aceti a kind of bacterium is used in production of alcoholic beverages |
| Vinegar | Certain bacteria are used to convert alcohol into acetic acid, which gives vinegar its acid taste. Acetobacter Aceti is used on production of vinegar, which gives vinegar odor of alcohol and alcoholic taste |
| Citric acid | Certain fungi are used to make citric acid, a common ingredient of soft drinks and other foods. |
| Vitamins | Microorganisms are used to make vitamins, including C, B2 , B12. |
| Antibiotics | With only a few exceptions, microorganisms are used to make antibiotics. Penicillin, Amoxicillin, Tetracycline, and Erythromycin |
Water treatment
[edit]
These depend for their ability to clean up water contaminated with organic material on microorganisms that can respire dissolved substances. Respiration may be aerobic, with a well-oxygenated filter bed such as a slow sand filter.[89] Anaerobic digestion by methanogens generate useful methane gas as a by-product.[90]
Energy
[edit]Microorganisms are used in fermentation to produce ethanol,[91] and in biogas reactors to produce methane.[92] Scientists are researching the use of algae to produce liquid fuels,[93] and bacteria to convert various forms of agricultural and urban waste into usable fuels.[94]
Chemicals, enzymes
[edit]Microorganisms are used to produce many commercial and industrial chemicals, enzymes and other bioactive molecules. Organic acids produced on a large industrial scale by microbial fermentation include acetic acid produced by acetic acid bacteria such as Acetobacter aceti, butyric acid made by the bacterium Clostridium butyricum, lactic acid made by Lactobacillus and other lactic acid bacteria,[95] and citric acid produced by the mould fungus Aspergillus niger.[95]
Microorganisms are used to prepare bioactive molecules such as Streptokinase from the bacterium Streptococcus,[96] Cyclosporin A from the ascomycete fungus Tolypocladium inflatum,[97] and statins produced by the yeast Monascus purpureus.[98]
Science
[edit]
Microorganisms are essential tools in biotechnology, biochemistry, genetics, and molecular biology. The yeasts Saccharomyces cerevisiae and Schizosaccharomyces pombe are important model organisms in science, since they are simple eukaryotes that can be grown rapidly in large numbers and are easily manipulated.[99] They are particularly valuable in genetics, genomics and proteomics.[100][101] Microorganisms can be harnessed for uses such as creating steroids and treating skin diseases. Scientists are also considering using microorganisms for living fuel cells,[102] and as a solution for pollution.[103]
Warfare
[edit]In the Middle Ages, as an early example of biological warfare, diseased corpses were thrown into castles during sieges using catapults or other siege engines. Individuals near the corpses were exposed to the pathogen and were likely to spread that pathogen to others.[104]
In modern times, bioterrorism has included the 1984 Rajneeshee bioterror attack[105] and the 1993 release of anthrax by Aum Shinrikyo in Tokyo.[106]
Soil
[edit]Microbes can make nutrients and minerals in the soil available to plants, produce hormones that spur growth, stimulate the plant immune system and trigger or dampen stress responses. In general a more diverse set of soil microbes results in fewer plant diseases and higher yield.[107]
Human health
[edit]Human gut flora
[edit]Microorganisms can form an endosymbiotic relationship with other, larger organisms. For example, microbial symbiosis plays a crucial role in the immune system. The microorganisms that make up the gut flora in the gastrointestinal tract contribute to gut immunity, synthesize vitamins such as folic acid and biotin, and ferment complex indigestible carbohydrates.[108] Some microorganisms that are seen to be beneficial to health are termed probiotics and are available as dietary supplements, or food additives.[109]
Disease
[edit]
Microorganisms are the causative agents (pathogens) in many infectious diseases. The organisms involved include pathogenic bacteria, causing diseases such as plague, tuberculosis and anthrax; protozoan parasites, causing diseases such as malaria, sleeping sickness, dysentery and toxoplasmosis; and also fungi causing diseases such as ringworm, candidiasis or histoplasmosis. However, other diseases such as influenza, yellow fever or AIDS are caused by pathogenic viruses, which are not usually classified as living organisms and are not, therefore, microorganisms by the strict definition. No clear examples of archaean pathogens are known,[110] although a relationship has been proposed between the presence of some archaean methanogens and human periodontal disease.[111] Numerous microbial pathogens are capable of sexual processes that appear to facilitate their survival in their infected host.[112]
Hygiene
[edit]Hygiene is a set of practices to avoid infection or food spoilage by eliminating microorganisms from the surroundings. As microorganisms, in particular bacteria, are found virtually everywhere, harmful microorganisms may be reduced to acceptable levels rather than actually eliminated. In food preparation, microorganisms are reduced by preservation methods such as cooking, cleanliness of utensils, short storage periods, or by low temperatures. If complete sterility is needed, as with surgical equipment, an autoclave is used to kill microorganisms with heat and pressure.[113][114]
In fiction
[edit]- Osmosis Jones, a 2001 film, and its show Ozzy & Drix, set in a stylized version of the human body, featured anthropomorphic microorganisms.
- War of the Worlds (2005 film), when alien lifeforms attempt to conquer Earth, they are ultimately defeated by a common microbe to which humans are immune.
See also
[edit]- Catalogue of Life
- Impedance microbiology
- Microbial biogeography
- Microbial intelligence
- Microbiological culture
- Microbivory, an eating behavior of some animals feeding on living microbes
- Nanobacterium
- Nylon-eating bacteria
- Petri dish
- Staining
- Budapest Treaty (Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purposes of Patent Procedure)
Notes
[edit]- ^ The word microorganism (/ˌmaɪkroʊˈɔːrɡənɪzəm/) uses combining forms of micro- (from the Greek: μικρός, mikros, 'small') and organism from ὀργανισμός, organismós, 'organism'). It is usually written as a single word but is sometimes hyphenated (micro-organism), especially in older texts. The informal synonym microbe (/ˈmaɪkroʊb/) comes from mikros and βίος, bíos, 'life'.
- ^ The piezophilic bacteria Halomonas salaria requires a pressure of 1,000 atm; nanobes, a speculative organism, have been reportedly found in the earth's crust at 2,000 atm.[79]
References
[edit]- ^ Tyrell, Kelly April (18 December 2017). "Oldest fossils ever found show life on Earth began before 3.5 billion years ago". University of Wisconsin–Madison. Retrieved 18 December 2017.
- ^ Schopf, J. William; Kitajima, Kouki; Spicuzza, Michael J.; Kudryavtsev, Anatolly B.; Valley, John W. (2017). "SIMS analyses of the oldest known assemblage of microfossils document their taxon-correlated carbon isotope compositions". PNAS. 115 (1): 53–58. Bibcode:2018PNAS..115...53S. doi:10.1073/pnas.1718063115. PMC 5776830. PMID 29255053.
- ^ a b Long, Jeffery D. (2013). Jainism: An Introduction. I.B.Tauris. p. 100. ISBN 978-0-85771-392-6.
- ^ Singh, Upinder (2008). A History of Ancient and Early Medieval India: From the Stone Age to the 12th Century. Pearson Education India. p. 315. ISBN 978-81-317-1677-9.
- ^ Dundas, Paul (2003). The Jains. Routledge. p. 106. ISBN 978-1-134-50165-6.
- ^ a b Varro on Agriculture 1, xii Loeb
- ^ Tschanz, David W. "Arab Roots of European Medicine". Heart Views. 4 (2). Archived from the original on 3 May 2011.
- ^ Colgan, Richard (2009). Advice to the Young Physician: On the Art of Medicine. Springer. p. 33. ISBN 978-1-4419-1033-2.
- ^ Taşköprülüzâde: Shaqaiq-e Numaniya, v. 1, p. 48
- ^ Osman Şevki Uludağ: Beş Buçuk Asırlık Türk Tabâbet Tarihi (Five and a Half Centuries of Turkish Medical History). Istanbul, 1969, pp. 35–36
- ^ Nutton, Vivian (1990). "The Reception of Fracastoro's Theory of Contagion: The Seed That Fell among Thorns?". Osiris. 2nd Series, Vol. 6, Renaissance Medical Learning: Evolution of a Tradition: 196–234. doi:10.1086/368701. JSTOR 301787. PMID 11612689. S2CID 37260514.
- ^ Leeuwenhoek, A. (1753). "Part of a Letter from Mr Antony van Leeuwenhoek, concerning the Worms in Sheeps Livers, Gnats, and Animalcula in the Excrements of Frogs". Philosophical Transactions. 22 (260–276): 509–518. Bibcode:1700RSPT...22..509V. doi:10.1098/rstl.1700.0013.
- ^ Leeuwenhoek, A. (1753). "Part of a Letter from Mr Antony van Leeuwenhoek, F. R. S. concerning Green Weeds Growing in Water, and Some Animalcula Found about Them". Philosophical Transactions. 23 (277–288): 1304–1311. Bibcode:1702RSPT...23.1304V. doi:10.1098/rstl.1702.0042. S2CID 186209549.
- ^ Lane, Nick (2015). "The Unseen World: Reflections on Leeuwenhoek (1677) 'Concerning Little Animal'". Philos Trans R Soc Lond B Biol Sci. 370 (1666) 20140344. doi:10.1098/rstb.2014.0344. PMC 4360124. PMID 25750239.
- ^ Payne, A. S. The Cleere Observer: A Biography of Antoni Van Leeuwenhoek, p. 13, Macmillan, 1970
- ^ Gest, H. (2005). "The remarkable vision of Robert Hooke (1635–1703): first observer of the microbial world". Perspect. Biol. Med. 48 (2): 266–272. doi:10.1353/pbm.2005.0053. PMID 15834198. S2CID 23998841.
- ^ Bordenave, G. (2003). "Louis Pasteur (1822–1895)". Microbes Infect. 5 (6): 553–560. doi:10.1016/S1286-4579(03)00075-3. PMID 12758285.
- ^ The Nobel Prize in Physiology or Medicine 1905 Nobelprize.org Accessed 22 November 2006.
- ^ O'Brien, S.; Goedert, J. (1996). "HIV causes AIDS: Koch's postulates fulfilled". Curr Opin Immunol. 8 (5): 613–618. doi:10.1016/S0952-7915(96)80075-6. PMID 8902385.
- ^ Scamardella, J. M. (1999). "Not plants or animals: a brief history of the origin of Kingdoms Protozoa, Protista and Protoctista" (PDF). International Microbiology. 2 (4): 207–221. PMID 10943416. Archived from the original (PDF) on 14 June 2011. Retrieved 1 October 2017.
- ^ Rothschild, L. J. (1989). "Protozoa, Protista, Protoctista: what's in a name?". J Hist Biol. 22 (2): 277–305. doi:10.1007/BF00139515. PMID 11542176. S2CID 32462158.
- ^ Solomon, Eldra Pearl; Berg, Linda R.; Martin, Diana W., eds. (2005). "Kingdoms or Domains?". Biology (7th ed.). Brooks/Cole Thompson Learning. pp. 421–7. ISBN 978-0-534-49276-2.
- ^ a b Madigan, M.; Martinko, J., eds. (2006). Brock Biology of Microorganisms (13th ed.). Pearson Education. p. 1096. ISBN 978-0-321-73551-5.
- ^ Johnson, J. (2001) [1998]. "Martinus Willem Beijerinck". APSnet. American Phytopathological Society. Archived from the original on 20 June 2010. Retrieved 2 May 2010. Retrieved from Internet Archive 12 January 2014.
- ^ Paustian, T.; Roberts, G. (2009). "Beijerinck and Winogradsky Initiate the Field of Environmental Microbiology". Through the Microscope: A Look at All Things Small (3rd ed.). Textbook Consortia. § 1–14. Archived from the original on 14 September 2008. Retrieved 3 October 2017.
- ^ Keen, E. C. (2012). "Felix d'Herelle and Our Microbial Future". Future Microbiology. 7 (12): 1337–1339. doi:10.2217/fmb.12.115. PMID 23231482.
- ^ Lim, Daniel V. (2001). "Microbiology". eLS. John Wiley. doi:10.1038/npg.els.0000459. ISBN 978-0-470-01590-2.
- ^ "What is Microbiology?". highveld.com. Archived from the original on 15 February 2015. Retrieved 2 June 2017.
- ^ Cann, Alan (2011). Principles of Molecular Virology (5th ed.). Academic Press. ISBN 978-0-12-384939-7.
- ^ Schopf, J. (2006). "Fossil evidence of Archaean life". Philos Trans R Soc Lond B Biol Sci. 361 (1470): 869–885. doi:10.1098/rstb.2006.1834. PMC 1578735. PMID 16754604.
- ^ Altermann, W.; Kazmierczak, J. (2003). "Archean microfossils: a reappraisal of early life on Earth". Res Microbiol. 154 (9): 611–617. doi:10.1016/j.resmic.2003.08.006. PMID 14596897.
- ^ Cavalier-Smith, T. (2006). "Cell evolution and Earth history: stasis and revolution". Philos Trans R Soc Lond B Biol Sci. 361 (1470): 969–1006. doi:10.1098/rstb.2006.1842. PMC 1578732. PMID 16754610.
- ^ Schopf, J. (1994). "Disparate rates, differing fates: tempo and mode of evolution changed from the Precambrian to the Phanerozoic". PNAS. 91 (15): 6735–6742. Bibcode:1994PNAS...91.6735S. doi:10.1073/pnas.91.15.6735. PMC 44277. PMID 8041691.
- ^ Stanley, S. (May 1973). "An Ecological Theory for the Sudden Origin of Multicellular Life in the Late Precambrian". PNAS. 70 (5): 1486–1489. Bibcode:1973PNAS...70.1486S. doi:10.1073/pnas.70.5.1486. PMC 433525. PMID 16592084.
- ^ DeLong, E.; Pace, N. (2001). "Environmental diversity of bacteria and archaea". Syst Biol. 50 (4): 470–478. CiteSeerX 10.1.1.321.8828. doi:10.1080/106351501750435040. PMID 12116647.
- ^ Schmidt, A.; Ragazzi, E.; Coppellotti, O.; Roghi, G. (2006). "A microworld in Triassic amber". Nature. 444 (7121): 835. Bibcode:2006Natur.444..835S. doi:10.1038/444835a. PMID 17167469. S2CID 4401723.
- ^ Schirber, Michael (27 July 2014). "Microbe's Innovation May Have Started Largest Extinction Event on Earth". Space.com. Astrobiology Magazine.
That spike in nickel allowed methanogens to take off.
- ^ Wolska, K. (2003). "Horizontal DNA transfer between bacteria in the environment". Acta Microbiol Pol. 52 (3): 233–243. PMID 14743976.
- ^ Enright, M.; Robinson, D.; Randle, G.; Feil, E.; Grundmann, H.; Spratt, B. (May 2002). "The evolutionary history of methicillin-resistant Staphylococcus aureus (MRSA)". Proc Natl Acad Sci USA. 99 (11): 7687–7692. Bibcode:2002PNAS...99.7687E. doi:10.1073/pnas.122108599. PMC 124322. PMID 12032344.
- ^ "Deep sea microorganisms and the origin of the eukaryotic cell" (PDF). Retrieved 24 October 2017.
- ^ Yamaguchi, Masashi; et al. (1 December 2012). "Prokaryote or eukaryote? A unique microorganism from the deep sea". Journal of Electron Microscopy. 61 (6): 423–431. doi:10.1093/jmicro/dfs062. PMID 23024290.
- ^ Woese, C.; Kandler, O.; Wheelis, M. (1990). "Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya". Proc Natl Acad Sci USA. 87 (12): 4576–4579. Bibcode:1990PNAS...87.4576W. doi:10.1073/pnas.87.12.4576. PMC 54159. PMID 2112744.
- ^ De Rosa, M.; Gambacorta, A.; Gliozzi, A. (1 March 1986). "Structure, biosynthesis, and physicochemical properties of archaebacterial lipids". Microbiol. Rev. 50 (1): 70–80. doi:10.1128/mmbr.50.1.70-80.1986. PMC 373054. PMID 3083222.
- ^ Robertson, C.; Harris, J.; Spear, J.; Pace, N. (2005). "Phylogenetic diversity and ecology of environmental Archaea". Curr Opin Microbiol. 8 (6): 638–642. doi:10.1016/j.mib.2005.10.003. PMID 16236543.
- ^ Karner, M. B.; DeLong, E. F.; Karl, D. M. (2001). "Archaeal dominance in the mesopelagic zone of the Pacific Ocean". Nature. 409 (6819): 507–510. Bibcode:2001Natur.409..507K. doi:10.1038/35054051. PMID 11206545. S2CID 6789859.
- ^ Sinninghe Damsté, J. S.; Rijpstra, W. I.; Hopmans, E. C.; Prahl, F. G.; Wakeham, S. G.; Schouten, S. (June 2002). "Distribution of Membrane Lipids of Planktonic Crenarchaeota in the Arabian Sea". Appl. Environ. Microbiol. 68 (6): 2997–3002. Bibcode:2002ApEnM..68.2997S. doi:10.1128/AEM.68.6.2997-3002.2002. PMC 123986. PMID 12039760.
- ^ Leininger, S.; Urich, T.; Schloter, M.; Schwark, L.; Qi, J.; Nicol, G. W.; Prosser, J. I.; Schuster, S. C.; Schleper, C. (2006). "Archaea predominate among ammonia-oxidizing prokaryotes in soils". Nature. 442 (7104): 806–809. Bibcode:2006Natur.442..806L. doi:10.1038/nature04983. PMID 16915287. S2CID 4380804.
- ^ a b Gold, T. (1992). "The deep, hot biosphere". Proc. Natl. Acad. Sci. USA. 89 (13): 6045–6049. Bibcode:1992PNAS...89.6045G. doi:10.1073/pnas.89.13.6045. PMC 49434. PMID 1631089.
- ^ Whitman, W.; Coleman, D.; Wiebe, W. (1998). "Prokaryotes: The unseen majority". PNAS. 95 (12): 6578–6583. Bibcode:1998PNAS...95.6578W. doi:10.1073/pnas.95.12.6578. PMC 33863. PMID 9618454.
- ^ "Researchers find that Earth may be home to 1 trillion species". National Science Foundation. 2 May 2016. Retrieved 6 May 2016.
- ^ van Wolferen, M.; Wagner, A; van der Does, C.; Albers, S. V. (2016). "The archaeal Ced system imports DNA". Proc Natl Acad Sci USA. 113 (9): 2496–501. Bibcode:2016PNAS..113.2496V. doi:10.1073/pnas.1513740113. PMC 4780597. PMID 26884154.
- ^ Bernstein H, Bernstein C. Sexual communication in archaea, the precursor to meiosis. pp. 103–117 in Biocommunication of Archaea (Guenther Witzany, ed.) 2017. Springer International Publishing ISBN 978-3-319-65535-2 DOI 10.1007/978-3-319-65536-9
- ^ Schulz, H.; Jorgensen, B. (2001). "Big bacteria". Annu Rev Microbiol. 55: 105–137. doi:10.1146/annurev.micro.55.1.105. PMID 11544351.
- ^ Shapiro, J. A. (1998). "Thinking about bacterial populations as multicellular organisms" (PDF). Annu. Rev. Microbiol. 52: 81–104. doi:10.1146/annurev.micro.52.1.81. PMID 9891794. Archived from the original (PDF) on 17 July 2011.
- ^ Muñoz-Dorado, J.; Marcos-Torres, F. J.; García-Bravo, E.; Moraleda-Muñoz, A.; Pérez, J. (2016). "Myxobacteria: Moving, Killing, Feeding, and Surviving Together". Frontiers in Microbiology. 7: 781. doi:10.3389/fmicb.2016.00781. PMC 4880591. PMID 27303375.
- ^ Johnsbor, O.; Eldholm, V.; Håvarstein, L. S. (December 2007). "Natural genetic transformation: prevalence, mechanisms and function". Res. Microbiol. 158 (10): 767–778. doi:10.1016/j.resmic.2007.09.004. PMID 17997281.
- ^ Eagon, R. (1962). "Pseudomonas Natriegens, a Marine Bacterium With a Generation Time of Less Than 10 Minutes". J Bacteriol. 83 (4): 736–737. doi:10.1128/JB.83.4.736-737.1962. PMC 279347. PMID 13888946.
- ^ Eukaryota: More on Morphology. (Retrieved 10 October 2006)
- ^ a b Dyall, S.; Brown, M.; Johnson, P. (2004). "Ancient invasions: from endosymbionts to organelles". Science. 304 (5668): 253–257. Bibcode:2004Sci...304..253D. doi:10.1126/science.1094884. PMID 15073369. S2CID 19424594.
- ^ Bernstein, H.; Bernstein, C.; Michod, R. E. (2012). "Chapter 1". In Kimura, Sakura; Shimizu, Sora (eds.). DNA repair as the primary adaptive function of sex in bacteria and eukaryotes. DNA Repair: New Research. Nova Sci. Publ. pp. 1–49. ISBN 978-1-62100-808-8. Archived from the original on 22 July 2018.
- ^ Cavalier-Smith T (1 December 1993). "Kingdom protozoa and its 18 phyla". Microbiol. Rev. 57 (4): 953–994. doi:10.1128/mmbr.57.4.953-994.1993. PMC 372943. PMID 8302218.
- ^ Corliss, J. O. (1992). "Should there be a separate code of nomenclature for the protists?". BioSystems. 28 (1–3): 1–14. Bibcode:1992BiSys..28....1C. doi:10.1016/0303-2647(92)90003-H. PMID 1292654.
- ^ Devreotes, P. (1989). "Dictyostelium discoideum: a model system for cell-cell interactions in development". Science. 245 (4922): 1054–1058. Bibcode:1989Sci...245.1054D. doi:10.1126/science.2672337. PMID 2672337.
- ^ Slapeta, J.; Moreira, D.; López-García, P. (2005). "The extent of protist diversity: insights from molecular ecology of freshwater eukaryotes". Proc. Biol. Sci. 272 (1576): 2073–2081. doi:10.1098/rspb.2005.3195. PMC 1559898. PMID 16191619.
- ^ Moreira, D.; López-García, P. (2002). "The molecular ecology of microbial eukaryotes unveils a hidden world" (PDF). Trends Microbiol. 10 (1): 31–38. doi:10.1016/S0966-842X(01)02257-0. PMID 11755083.
- ^ Kumamoto, C.A.; Vinces, M. D. (2005). "Contributions of hyphae and hypha-co-regulated genes to Candida albicans virulence". Cell. Microbiol. 7 (11): 1546–1554. doi:10.1111/j.1462-5822.2005.00616.x. PMID 16207242.
- ^ Thomas, David C. (2002). Seaweeds. London: Natural History Museum. ISBN 978-0-565-09175-0.
- ^ Szewzyk, U.; Szewzyk, R.; Stenström, T. (1994). "Thermophilic, anaerobic bacteria isolated from a deep borehole in granite in Sweden". PNAS. 91 (5): 1810–1813. Bibcode:1994PNAS...91.1810S. doi:10.1073/pnas.91.5.1810. PMC 43253. PMID 11607462.
- ^ Horneck, G. (1981). "Survival of microorganisms in space: a review". Adv Space Res. 1 (14): 39–48. doi:10.1016/0273-1177(81)90241-6. PMID 11541716.
- ^ Rousk, Johannes; Bengtson, Per (2014). "Microbial regulation of global biogeochemical cycles". Frontiers in Microbiology. 5 (2): 210–225. doi:10.3389/fmicb.2014.00103. PMC 3954078. PMID 24672519.
- ^ a b Filloux, A. A. M., ed. (2012). Bacterial Regulatory Networks. Caister Academic Press. ISBN 978-1-908230-03-4.
- ^ Gross, R.; Beier, D., eds. (2012). Two-Component Systems in Bacteria. Caister Academic Press. ISBN 978-1-908230-08-9.
- ^ Requena, J. M., ed. (2012). Stress Response in Microbiology. Caister Academic Press. ISBN 978-1-908230-04-1.
- ^ Strain 121, a hyperthermophilic archaea, has been shown to reproduce at 121 °C (250 °F), and survive at 130 °C (266 °F).[1]
- ^ Some Psychrophilic bacteria can grow at −17 °C (1 °F)),[2] and can survive near absolute zero)."Earth microbes on the Moon". Archived from the original on 23 March 2010. Retrieved 20 July 2009.
- ^ Dyall-Smith, Mike, HALOARCHAEA, University of Melbourne. See also Haloarchaea.
- ^ Olsson, Karen; Keis, Stefanie; Morgan, Hugh W.; Dimroth, Peter; Cook, Gregory M. (15 January 2003). "Bacillus alcalophilus can grow at up to pH 11.5" (PDF). Journal of Bacteriology. 185 (2): 461–465. doi:10.1128/JB.185.2.461-465.2003. PMC 145327. PMID 12511491.
- ^ Picrophilus can grow at pH −0.06.[3] Archived 22 June 2010 at the Wayback Machine
- ^ "Microscopy-UK Home (Resources for the microscopy enthusiast and amateur, including free monthly e-zine, Micscape.)".
- ^ Anderson, A. W.; Nordan, H. C.; Cain, R. F.; Parrish, G.; Duggan, D. (1956). "Studies on a radio-resistant micrococcus. I. Isolation, morphology, cultural characteristics, and resistance to gamma radiation". Food Technol. 10 (1): 575–577.
- ^ Cavicchioli, R. (2002). "Extremophiles and the search for extraterrestrial life" (PDF). Astrobiology. 2 (3): 281–292. Bibcode:2002AsBio...2..281C. CiteSeerX 10.1.1.472.3179. doi:10.1089/153110702762027862. PMID 12530238. Archived from the original (PDF) on 26 April 2022. Retrieved 26 October 2017.
- ^ Barea, J.; Pozo, M.; Azcón, R.; Azcón-Aguilar, C. (2005). "Microbial co-operation in the rhizosphere". J Exp Bot. 56 (417): 1761–1778. doi:10.1093/jxb/eri197. PMID 15911555.
- ^ Gottel, Neil R.; Castro, Hector F.; Kerley, Marilyn; Yang, Zamin; Pelletier, Dale A.; Podar, Mircea; Karpinets, Tatiana; Uberbacher, Ed; Tuskan, Gerald A.; Vilgalys, Rytas; Doktycz, Mitchel J.; Schadt, Christopher W. (2011). "Distinct Microbial Communities within the Endosphere and Rhizosphere of Populus deltoides Roots across Contrasting Soil Types". Applied and Environmental Microbiology. 77 (17): 5934–5944. Bibcode:2011ApEnM..77.5934G. doi:10.1128/AEM.05255-11. PMC 3165402. PMID 21764952.
- ^ Rebolleda-Gómez, M.; Wood, C. W. (2019). "Unclear Intentions: Eavesdropping in Microbial and Plant Systems". Frontiers in Ecology and Evolution. 7 (385). doi:10.3389/fevo.2019.00385.
- ^ "What is a lichen?". Australian National Botanic Gardens. Retrieved 30 September 2017.
- ^ "Introduction to Lichens – An Alliance between Kingdoms". University of California Museum of Paleontology. Retrieved 30 September 2017.
- ^ "Dairy Microbiology". University of Guelph. Retrieved 9 October 2006.
- ^ Hui, Y. H.; Meunier-Goddik, L.; Josephsen, J.; Nip, W. K.; Stanfield, P. S. (2004). Handbook of Food and Beverage Fermentation Technology. CRC Press. pp. 27 et passim. ISBN 978-0-8247-5122-7.
- ^ Gray, N. F. (2004). Biology of Wastewater Treatment. Imperial College Press. p. 1164. ISBN 978-1-86094-332-4.
- ^ Tabatabaei, Meisam (2010). "Importance of the methanogenic archaea populations in anaerobic wastewater treatments" (PDF). Process Biochemistry. 45 (8): 1214–1225. doi:10.1016/j.procbio.2010.05.017.
- ^ Kitani, Osumu; Hall, Carl W. (1989). Biomass Handbook. Taylor & Francis US. p. 256. ISBN 978-2-88124-269-4.
- ^ Pimental, David (2007). Food, Energy, and Society. CRC Press. p. 289. ISBN 978-1-4200-4667-0.
- ^ Tickell, Joshua; et al. (2000). From the Fryer to the Fuel Tank: The Complete Guide to Using Vegetable Oil as an Alternative Fuel. Biodiesel America. p. 53. ISBN 978-0-9707227-0-6.
- ^ Inslee, Jay; et al. (2008). Apollo's Fire: Igniting America's Clean Energy Economy. Island Press. p. 157. ISBN 978-1-59726-175-3.
- ^ a b Sauer, Michael; Porro, Danilo; et al. (2008). "Microbial production of organic acids: expanding the markets" (PDF). Trends in Biotechnology. 26 (2): 100–108. doi:10.1016/j.tibtech.2007.11.006. PMID 18191255. Archived from the original (PDF) on 28 September 2017. Retrieved 28 September 2017.
- ^ Babashamsi, Mohammed; et al. (2009). "Production and Purification of Streptokinase by Protected Affinity Chromatography". Avicenna Journal of Medical Biotechnology. 1 (1): 47–51. PMC 3558118. PMID 23407807.
Streptokinase is an extracellular protein, extracted from certain strains of beta hemolytic streptococcus.
- ^ Borel, J. F.; Kis, Z. L.; Beveridge, T. (1995). "The history of the discovery and development of Cyclosporin". In Merluzzi, V. J.; Adams, J. (eds.). The search for anti-inflammatory drugs case histories from concept to clinic. Boston: Birkhäuser. pp. 27–63. ISBN 978-1-4615-9846-6.
- ^ Biology textbook for class XII. National council of educational research and training. 2006. p. 183. ISBN 978-81-7450-639-9.
- ^ Castrillo, J. I.; Oliver, S. G. (2004). "Yeast as a touchstone in post-genomic research: strategies for integrative analysis in functional genomics". J. Biochem. Mol. Biol. 37 (1): 93–106. doi:10.5483/BMBRep.2004.37.1.093. PMID 14761307.
- ^ Suter, B.; Auerbach, D.; Stagljar, I. (2006). "Yeast-based functional genomics and proteomics technologies: the first 15 years and beyond". BioTechniques. 40 (5): 625–644. doi:10.2144/000112151. PMID 16708762.
- ^ Sunnerhagen, P. (2002). "Prospects for functional genomics in Schizosaccharomyces pombe". Curr. Genet. 42 (2): 73–84. doi:10.1007/s00294-002-0335-6. PMID 12478386. S2CID 22067347.
- ^ Soni, S. K. (2007). Microbes: A Source of Energy for 21st Century. New India Publishing. ISBN 978-81-89422-14-1.
- ^ Moses, Vivian; et al. (1999). Biotechnology: The Science and the Business. CRC Press. p. 563. ISBN 978-90-5702-407-8.
- ^ Langford, Roland E. (2004). Introduction to Weapons of Mass Destruction: Radiological, Chemical, and Biological. Wiley-IEEE. p. 140. ISBN 978-0-471-46560-7.
- ^ Novak, Matt (3 November 2016). "The Largest Bioterrorism Attack in US History Was An Attempt To Swing An Election". Gizmodo.
- ^ Takahashi, Hiroshi; Keim, Paul; Kaufmann, Arnold F.; Keys, Christine; Smith, Kimothy L.; Taniguchi, Kiyosu; Inouye, Sakae; Kurata, Takeshi (2004). "Bacillus anthracis Bioterrorism Incident, Kameido, Tokyo, 1993". Emerging Infectious Diseases. 10 (1): 117–120. doi:10.3201/eid1001.030238. PMC 3322761. PMID 15112666.
- ^ Vrieze, Jop de (14 August 2015). "The littlest farmhands". Science. 349 (6249): 680–683. Bibcode:2015Sci...349..680D. doi:10.1126/science.349.6249.680. PMID 26273035.
- ^ O'Hara, A.; Shanahan, F. (2006). "The gut flora as a forgotten organ". EMBO Rep. 7 (7): 688–693. doi:10.1038/sj.embor.7400731. PMC 1500832. PMID 16819463.
- ^ Schlundt, Jorgen. "Health and Nutritional Properties of Probiotics in Food including Powder Milk with Live Lactic Acid Bacteria" (PDF). Report of a Joint FAO/WHO Expert Consultation on Evaluation of Health and Nutritional Properties of Probiotics in Food Including Powder Milk with Live Lactic Acid Bacteria. FAO / WHO. Archived from the original (PDF) on 22 October 2012. Retrieved 17 December 2012.
- ^ Eckburg, P.; Lepp, P.; Relman, D. (2003). "Archaea and Their Potential Role in Human Disease". Infect Immun. 71 (2): 591–596. doi:10.1128/IAI.71.2.591-596.2003. PMC 145348. PMID 12540534.
- ^ Lepp, P.; Brinig, M.; Ouverney, C.; Palm, K.; Armitage, G.; Relman, D. (2004). "Methanogenic Archaea and human periodontal disease". Proc Natl Acad Sci USA. 101 (16): 6176–6181. Bibcode:2004PNAS..101.6176L. doi:10.1073/pnas.0308766101. PMC 395942. PMID 15067114.
- ^ Bernstein H.; Bernstein, C.; Michod, R. E. (January 2018). "Sex in microbial pathogens". Infect Genet Evol. 57: 8–25. Bibcode:2018InfGE..57....8B. doi:10.1016/j.meegid.2017.10.024. PMID 29111273.
- ^ "Hygiene". World Health Organization (WHO). Archived from the original on 23 August 2004. Retrieved 18 May 2017.
- ^ "The Five Keys to Safer Food Programme". World Health Organization. Archived from the original on 7 December 2003. Retrieved 23 May 2021.
External links
[edit]- Microbes.info is a microbiology information portal containing a vast collection of resources including articles, news, frequently asked questions, and links pertaining to the field of microbiology.
- Our Microbial Planet Archived 15 February 2013 at the Wayback Machine A free poster from the National Academy of Sciences about the positive roles of micro-organisms.
- "Uncharted Microbial World: Microbes and Their Activities in the Environment" Report from the American Academy of Microbiology
- Understanding Our Microbial Planet: The New Science of Metagenomics Archived 15 February 2013 at the Wayback Machine A 20-page educational booklet providing a basic overview of metagenomics and our microbial planet.
- Tree of Life Eukaryotes
- Microbe News from Genome News Network
- Medical Microbiology On-line textbook
- Through the microscope: A look at all things small Archived 13 September 2008 at the Wayback Machine On-line microbiology textbook by Timothy Paustian and Gary Roberts, University of Wisconsin–Madison
- Microorganisms in the pond water on YouTube
- Methane-spewing microbe blamed in worst mass extinction. CBCNews
Microorganism
View on GrokipediaA microorganism, or microbe, is a microscopic organism too small to be seen with the naked eye, requiring magnification for observation, and typically includes unicellular or simple multicellular entities such as bacteria, archaea, fungi, protozoa, and algae, while viruses—acellular infectious agents—are frequently grouped with them despite lacking independent metabolism or reproduction.[1][2][3]
Microorganisms exhibit immense diversity, spanning all three domains of life—Bacteria, Archaea, and Eukarya—and dominate Earth's biomass, inhabiting extreme environments from hydrothermal vents to acidic soils and human microbiomes.[3][4]
They drive essential ecological processes, including nutrient cycling through nitrogen fixation, decomposition of organic matter, and symbiotic relationships that enable host survival, without which complex life forms could not persist.[5][6]
Certain pathogenic microbes, however, cause infectious diseases by invading hosts and disrupting physiological functions, contributing to significant morbidity and mortality across species.[7][8]
First observed in the 1670s by Antonie van Leeuwenhoek using self-crafted microscopes, microorganisms revolutionized biology, revealing an invisible world foundational to understanding life, disease, and biotechnology.[9][10]


